UniProt ID: Q9VSL1 (1-133) Uncharacterized protein, isoform A
|
|
|
|
| Sample: |
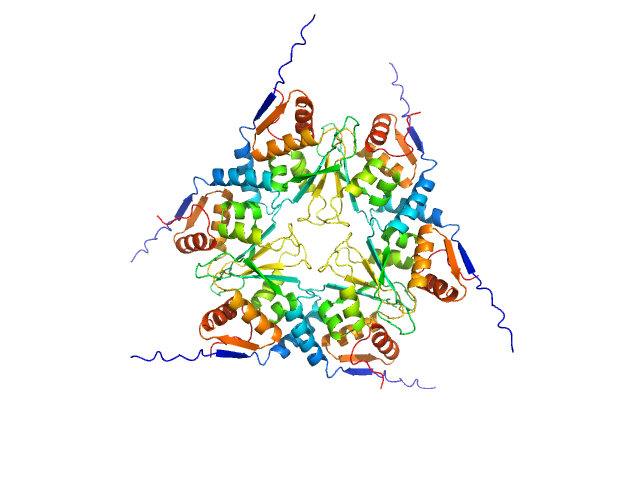
Uncharacterized protein, isoform A hexamer, 92 kDa Drosophila melanogaster protein
|
| Buffer: |
20 mM Tris, pH 7.4, 200 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jul 20
|
The Arthropoda-specific Tramtrack group BTB protein domains use previously unknown interface to form hexamers.
Elife 13 (2024)
Bonchuk AN, Balagurov KI, Baradaran R, Boyko KM, Sluchanko NN, Khrustaleva AM, Burtseva AD, Arkova OV, Khalisova KK, Popov VO, Naschberger A, Georgiev PG
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
166 |
nm3 |
|
|
UniProt ID: A0A0C5PJM1 (452-940) Conjugal transfer mating pair stabilization protein TraG
|
|
|

|
| Sample: |
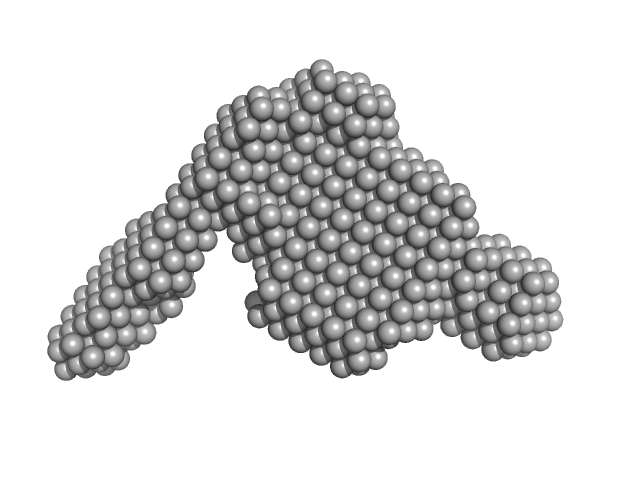
Conjugal transfer mating pair stabilization protein TraG monomer, 57 kDa Shigella flexneri 4c protein
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 5% glycerol, 0.05% NP40, pH: 7 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Feb 23
|
Solution characterization of the dynamic conjugative entry exclusion protein TraG
Structural Dynamics 9(6):064702 (2022)
Bragagnolo N, Audette G
|
| RgGuinier |
5.8 |
nm |
| Dmax |
45.0 |
nm |
|
|
UniProt ID: P11021 (26-405) Endoplasmic reticulum chaperone BiP
UniProt ID: Q49AH0 (127-187) C-terminal of the Cerebral dopamine neurotrophic factor
|
|
|
|
| Sample: |
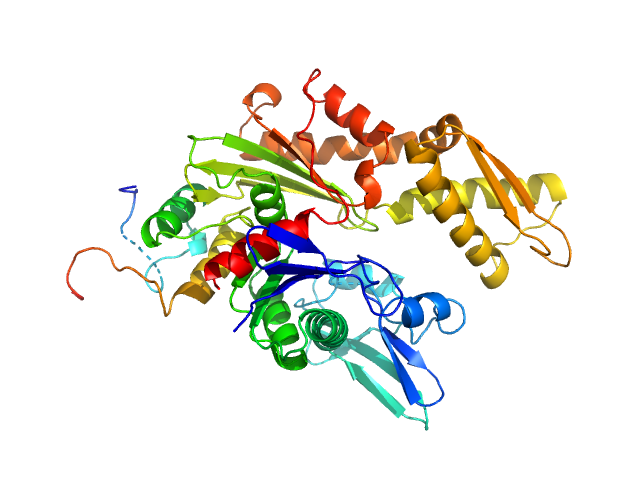
Endoplasmic reticulum chaperone BiP monomer, 42 kDa Homo sapiens protein
C-terminal of the Cerebral dopamine neurotrophic factor monomer, 7 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Mar 23
|
Structural basis of CDNF interaction with the UPR regulator GRP78.
Nat Commun 15(1):8175 (2024)
Graewert MA, Volkova M, Jonasson K, Määttä JAE, Gräwert T, Mamidi S, Kulesskaya N, Evenäs J, Johnsson RE, Svergun D, Bhattacharjee A, Huttunen HJ
|
| RgGuinier |
2.4 |
nm |
| Dmax |
6.9 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
UniProt ID: A0A509AR09 (2-228) Glutamate--tRNA ligase
|
|
|
|
| Sample: |
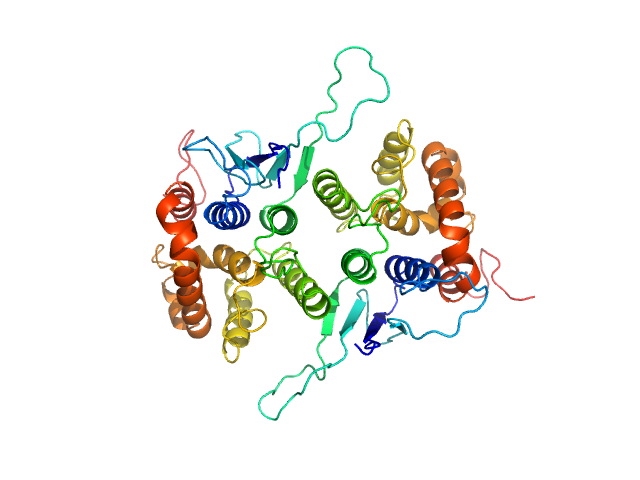
Glutamate--tRNA ligase dimer, 57 kDa Plasmodium berghei (strain … protein
|
| Buffer: |
25 mM HEPES-NaOH, 300 mM NaCl, 5% (v/v) glycerol, 0.005% (m/v) n-dodecyl beta-D-maltoside, 5 mM 2-mercaptoethanol, pH: 7 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2021 Oct 9
|
Solution x-ray scattering highlights discrepancies in Plasmodium multi-aminoacyl-tRNA synthetase complexes.
Protein Sci :e4564 (2023)
Jaramillo Ponce JR, Théobald-Dietrich A, Bénas P, Paulus C, Sauter C, Frugier M
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
96 |
nm3 |
|
|
UniProt ID: A0A3Y0E1G8 (1-375) DGQHR domain-containing protein
|
|
|
|
| Sample: |
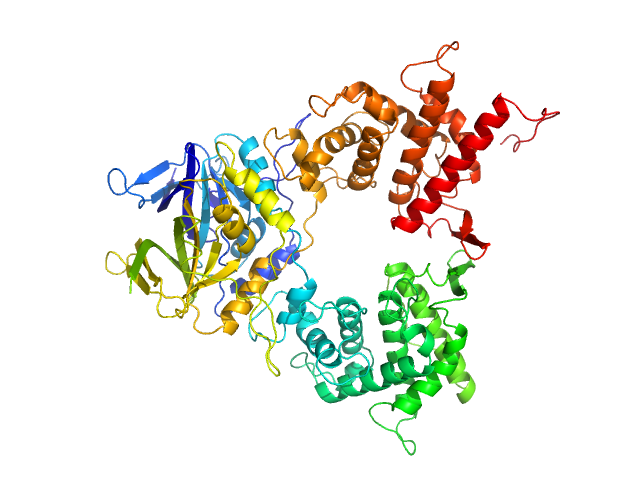
DGQHR domain-containing protein dimer, 88 kDa Salmonella enterica subsp. … protein
|
| Buffer: |
100 mM KCl, 50 mM Tris pH 7.0, 1 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Dec 22
|
7-Deazaguanines in DNA: functional and structural elucidation of a DNA modification system.
Nucleic Acids Res (2023)
Gedara SH, Wood E, Gustafson A, Liang C, Hung SH, Savage J, Phan P, Luthra A, de Crécy-Lagard V, Dedon P, Swairjo MA, Iwata-Reuyl D
|
| RgGuinier |
3.4 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
126 |
nm3 |
|
|
UniProt ID: P22363-3 (53-297) Phosphoprotein (Isoform P3; C297S)
|
|
|
|
| Sample: |
Phosphoprotein (Isoform P3; C297S) dimer, 56 kDa Rabies virus (strain … protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 1 mM TCEP, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2021 Mar 7
|
Conformational dynamics, RNA binding, and phase separation regulate the multifunctionality of rabies virus P protein.
Nat Commun 16(1):9491 (2025)
Rawlinson SM, Chakraborty S, Sethi A, David CT, Harrison AR, Bird LE, Rozario AM, Uthishtran S, Ardipradja K, Zhao T, Oksayan S, Jans DA, Ang CS, Lu ZH, Yan F, Williamson NA, Arumugam S, Sundaramoorthy V, Bell TDM, Gooley PR, Moseley GW
|
| RgGuinier |
4.2 |
nm |
| Dmax |
17.0 |
nm |
| VolumePorod |
141 |
nm3 |
|
|
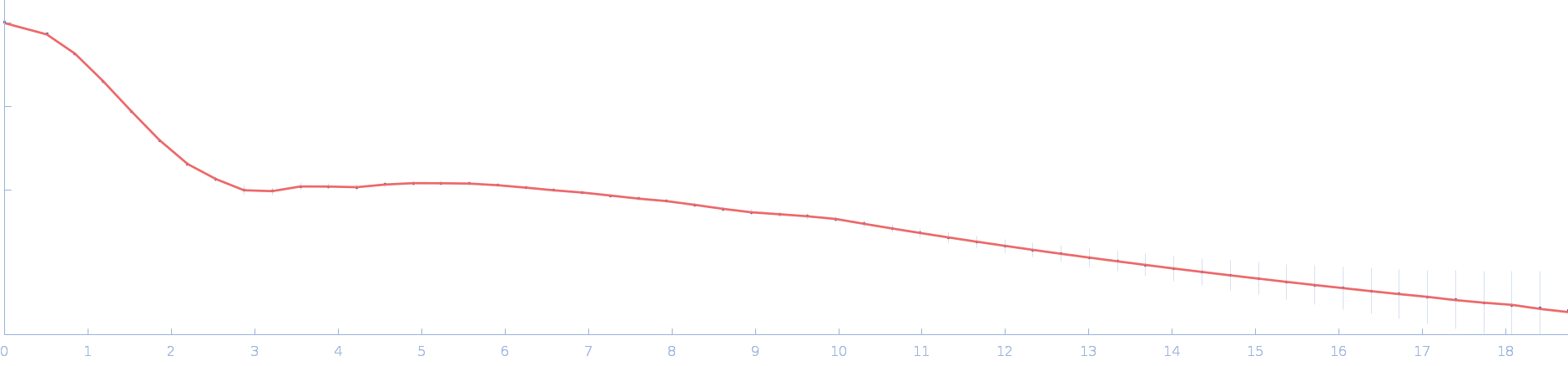
UniProt ID: P73129 (1-96) Ssr1698 protein
|
|
|
|
| Sample: |
Ssr1698 protein dimer, 23 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Hepes, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2022 Jun 13
|
A hemoprotein with a zinc-mirror heme site ties heme availability to carbon metabolism in cyanobacteria.
Nat Commun 15(1):3167 (2024)
Grosjean N, Yee EF, Kumaran D, Chopra K, Abernathy M, Biswas S, Byrnes J, Kreitler DF, Cheng JF, Ghosh A, Almo SC, Iwai M, Niyogi KK, Pakrasi HB, Sarangi R, van Dam H, Yang L, Blaby IK, Blaby-Haas CE
|
| RgGuinier |
1.9 |
nm |
| Dmax |
6.2 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
UniProt ID: P54760 (586-602) Ephrin type-B receptor 4
UniProt ID: P20936 (174-444) Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S)
|
|
|
|
| Sample: |
Ephrin type-B receptor 4 monomer, 2 kDa Homo sapiens protein
Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S) monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 350 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Dec 12
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
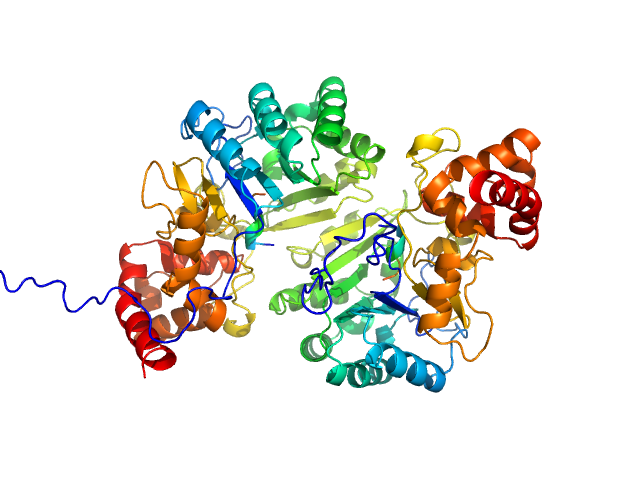
UniProt ID: Q45595 (1-319) Putative peptide biosynthesis protein YydG
|
|
|
|
| Sample: |
Putative peptide biosynthesis protein YydG dimer, 80 kDa Bacillus subtilis (strain … protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2022 Nov 30
|
Structural and mechanistic basis for RiPP epimerization by a radical SAM enzyme.
Nat Chem Biol (2023)
Kubiak X, Polsinelli I, Chavas LMG, Fyfe CD, Guillot A, Fradale L, Brewee C, Grimaldi S, Gerbaud G, Thureau A, Legrand P, Berteau O, Benjdia A
|
| RgGuinier |
2.9 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
113 |
nm3 |
|
|
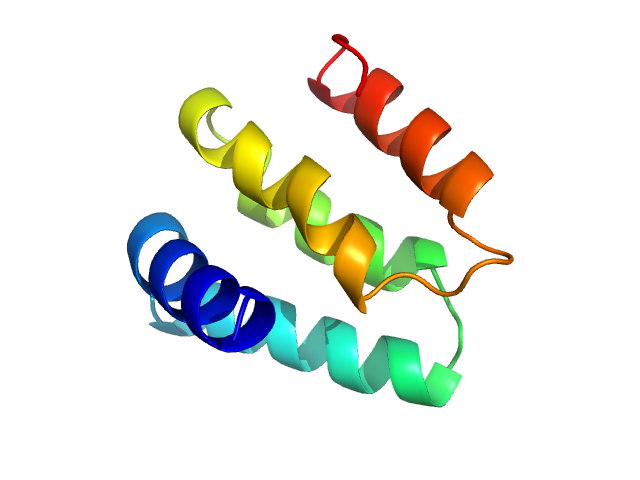
UniProt ID: Q9NZ71-6 (1053-1147) Regulator of telomere elongation helicase 1 (Isoform 6, 1053-1147)
|
|
|
|
| Sample: |
Regulator of telomere elongation helicase 1 (Isoform 6, 1053-1147) monomer, 11 kDa Homo sapiens protein
|
| Buffer: |
50 mM Hepes, 150 mM NaCl,10 (v/v)% glycerol, 1 mM TCEP, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 22
|
Structural and biochemical characterization of the C‐terminal region of the human RTEL
1 helicase
Protein Science 33(9) (2024)
Cortone G, Graewert M, Kanade M, Longo A, Hegde R, González‐Magaña A, Chaves‐Arquero B, Blanco F, Napolitano L, Onesti S
|
| RgGuinier |
1.5 |
nm |
| Dmax |
5.0 |
nm |
| VolumePorod |
24 |
nm3 |
|
|