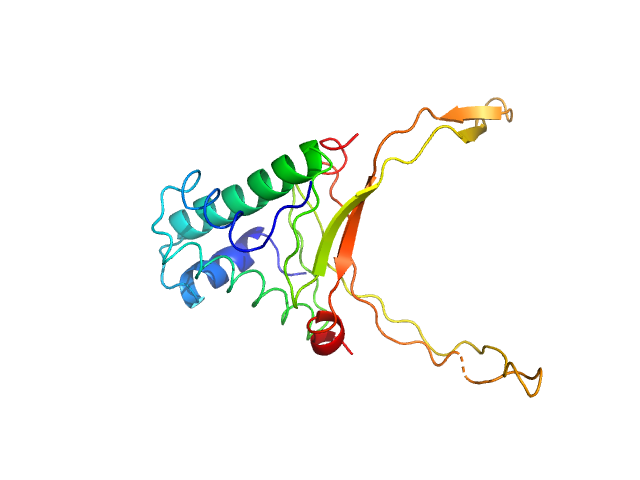
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha
|
|
|
|
| Sample: |
DNA-binding protein HU-alpha octamer, 77 kDa Escherichia coli protein
|
| Buffer: |
10 mM Bis-Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 May 27
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling.
Nat Commun 11(1):2905 (2020)
Remesh SG, Verma SC, Chen JH, Ekman AA, Larabell CA, Adhya S, Hammel M
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.7 |
nm |
|
|
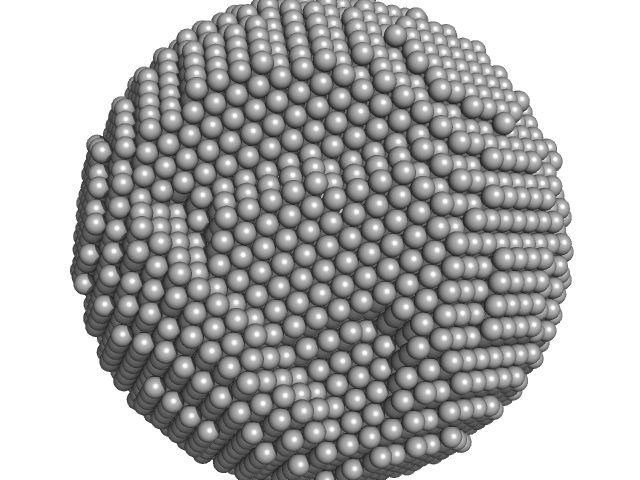
UniProt ID: P02791 (1-175) Apoferritin light chain
|
|
|

|
| Sample: |
Apoferritin light chain 24-mer, 479 kDa Equus caballus protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 2% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Apr 5
|
Adding Size Exclusion Chromatography (SEC) and Light Scattering (LS) Devices to Obtain High-Quality Small Angle X-Ray Scattering (SAXS) Data
Crystals 10(11):975 (2020)
Graewert M, Da Vela S, Gräwert T, Molodenskiy D, Blanchet C, Svergun D, Jeffries C
|
| RgGuinier |
5.4 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
679 |
nm3 |
|
|
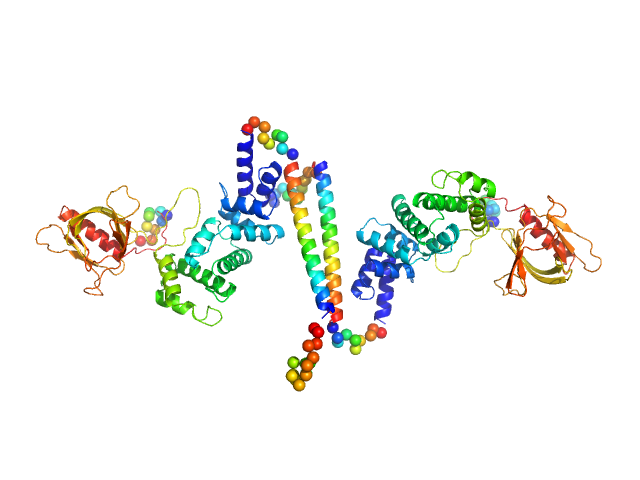
UniProt ID: O08967 (14-390) Cytohesin-3
|
|
|
|
| Sample: |
Cytohesin-3 dimer, 90 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 2 mM MgCl2, 0.1% 2-mercaptoethanol, 5% glycerol, 0.001 mM insitol 1,3,4,5-tetrakis phosphate, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2013 Nov 15
|
Structural Organization and Dynamics of Homodimeric Cytohesin Family Arf GTPase Exchange Factors in Solution and on Membranes.
Structure (2019)
Das S, Malaby AW, Nawrotek A, Zhang W, Zeghouf M, Maslen S, Skehel M, Chakravarthy S, Irving TC, Bilsel O, Cherfils J, Lambright DG
|
| RgGuinier |
5.1 |
nm |
| Dmax |
25.7 |
nm |
| VolumePorod |
168 |
nm3 |
|
|
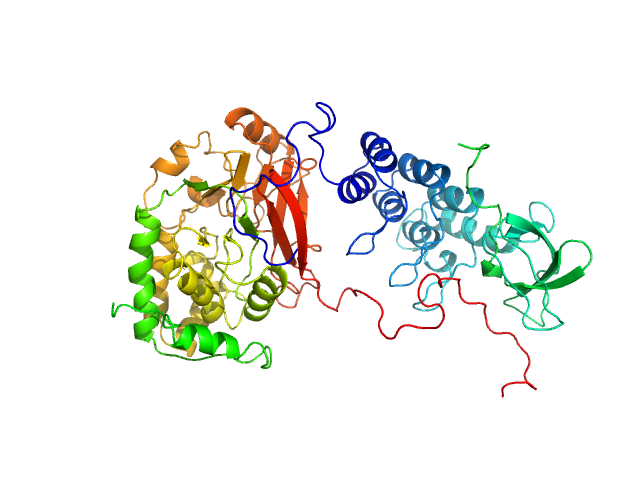
UniProt ID: P62136 (1-330) Serine/threonine-protein phosphatase PP1-alpha catalytic subunit
UniProt ID: Q13625 (905-1128) Apoptosis-stimulating of p53 protein 2
|
|
|
|
| Sample: |
Serine/threonine-protein phosphatase PP1-alpha catalytic subunit monomer, 38 kDa Homo sapiens protein
Apoptosis-stimulating of p53 protein 2 monomer, 26 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 May 31
|
Flexible Tethering of ASPP Proteins Facilitates PP-1c Catalysis.
Structure 27(10):1485-1496.e4 (2019)
Zhou Y, Millott R, Kim HJ, Peng S, Edwards RA, Skene-Arnold T, Hammel M, Lees-Miller SP, Tainer JA, Holmes CFB, Glover JNM
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
144 |
nm3 |
|
|
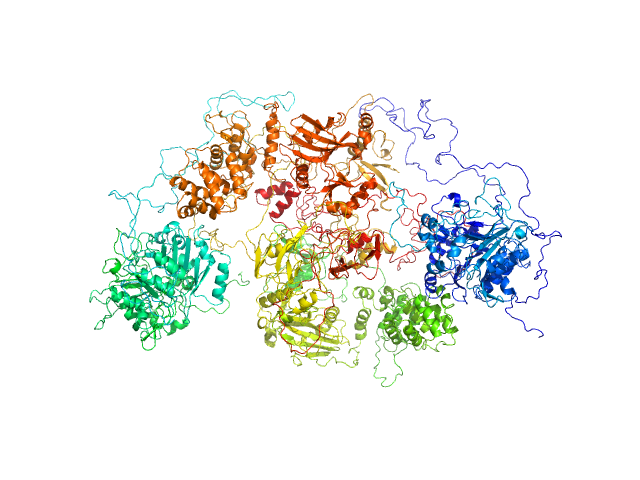
UniProt ID: P49916 (88-1009) DNA ligase 3 (DNA ligase III alpha)
UniProt ID: Q9NUW8 (1-608) Tyrosyl-DNA phosphodiesterase 1
|
|
|
|
| Sample: |
DNA ligase 3 (DNA ligase III alpha), Homo sapiens protein
Tyrosyl-DNA phosphodiesterase 1 monomer, 71 kDa Homo sapiens protein
|
| Buffer: |
200 mM NaCl, 40 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Sep 10
|
Direct interaction of DNA repair protein tyrosyl DNA phosphodiesterase 1 and the DNA ligase III catalytic domain is regulated by phosphorylation of its flexible N-terminus.
J Biol Chem :100921 (2021)
Rashid I, Hammel M, Sverzhinsky A, Tsai MS, Pascal JM, Tainer JA, Tomkinson AE
|
| RgGuinier |
6.5 |
nm |
| Dmax |
25.5 |
nm |
| VolumePorod |
790 |
nm3 |
|
|
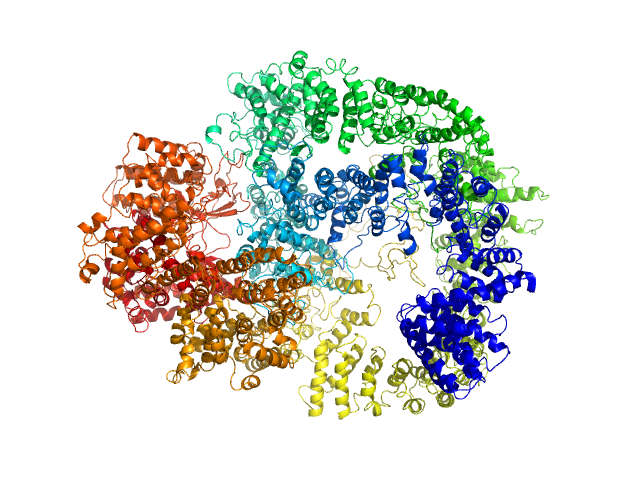
UniProt ID: P78527 (10-4128) DNA-dependent protein kinase catalytic subunit
|
|
|
|
| Sample: |
DNA-dependent protein kinase catalytic subunit monomer, 468 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 100 mM KCl, 5% (v/v) glycerol, 0.2 mM EDTA containing 0.1 mM benzamidine, 0.2 mM PMSF and 0.2 µg/ml pepstatin, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Jan 8
|
Visualizing functional dynamicity in the DNA-dependent protein kinase holoenzyme DNA-PK complex by integrating SAXS with cryo-EM.
Prog Biophys Mol Biol (2020)
Hammel M, Rosenberg DJ, Bierma J, Hura GL, Lees-Miller SP, Tainer JA
|
| RgGuinier |
5.5 |
nm |
| Dmax |
16.1 |
nm |
| VolumePorod |
918 |
nm3 |
|
|
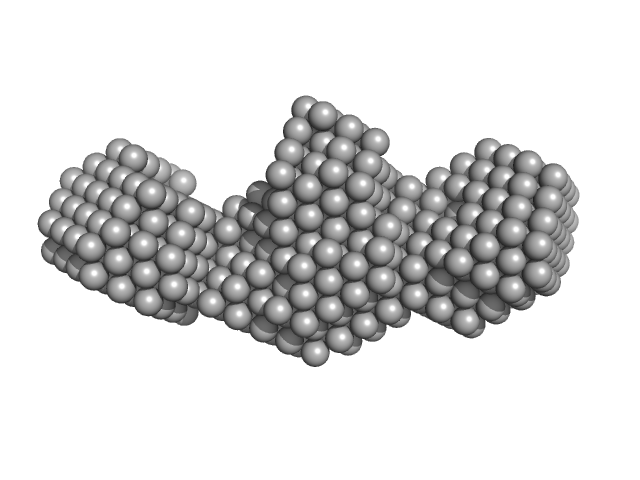
UniProt ID: I6YC99 (39-140) Mce-family protein Mce4A
|
|
|

|
| Sample: |
Mce-family protein Mce4A monomer, 15 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50mM MOPS, 350mM NaCl, 10% Glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 May 10
|
Structural insights into the substrate-binding proteins Mce1A and Mce4A from Mycobacterium tuberculosis
IUCrJ 8(5) (2021)
Asthana P, Singh D, Pedersen J, Hynönen M, Sulu R, Murthy A, Laitaoja M, Jänis J, Riley L, Venkatesan R
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.8 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
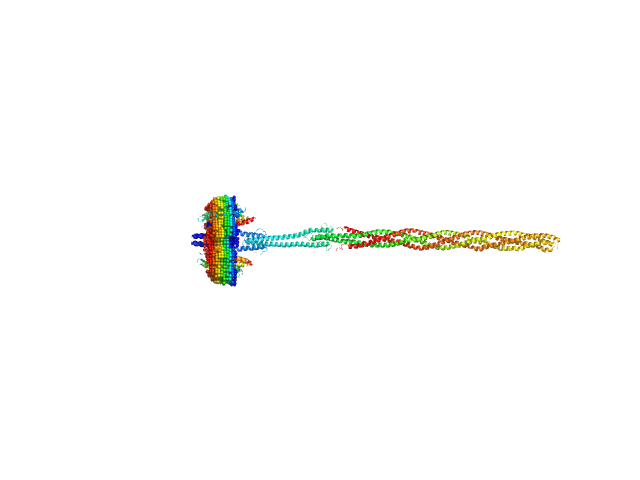
UniProt ID: P42196 (1-239) Sensory rhodopsin II from Natronbacterium pharaonis
UniProt ID: P42259 (3-534) Sensory rhodopsin II transducer from Natronomonas pharaonis
|
|
|

|
| Sample: |
Sensory rhodopsin II from Natronbacterium pharaonis dimer, 53 kDa Natronomonas pharaonis protein
Sensory rhodopsin II transducer from Natronomonas pharaonis dimer, 116 kDa Natronomonas pharaonis protein
|
| Buffer: |
2800 mM NaCl, 76.6 mM Na/Na-Pi, 1.0 mM EDTA, 0.05% DDM (D2O buffer), pH: 8 |
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 Feb 10
|
Molecular model of a sensor of two-component signaling system
Scientific Reports 11(1) (2021)
Ryzhykau Y, Orekhov P, Rulev M, Vlasov A, Melnikov I, Volkov D, Nikolaev M, Zabelskii D, Murugova T, Chupin V, Rogachev A, Gruzinov A, Svergun D, Brennich M, Gushchin I, Soler-Lopez M, Bothe A, Büldt G, Leonard G, Engelhard M, Kuklin A, Gordeliy V
|
| RgGuinier |
9.3 |
nm |
| Dmax |
37.5 |
nm |
|
|
UniProt ID: Q15059 (25-416) Bromodomain-containing protein 3
|
|
|
|
| Sample: |
Bromodomain-containing protein 3 monomer, 44 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Sep 25
|
Multivalent nucleosome scaffolding by bromodomain and extraterminal domain tandem bromodomains.
J Biol Chem :108289 (2025)
Olp MD, Bursch KL, Wynia-Smith SL, Nuñez R, Goetz CJ, Jackson V, Smith BC
|
| RgGuinier |
5.2 |
nm |
| Dmax |
18.6 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
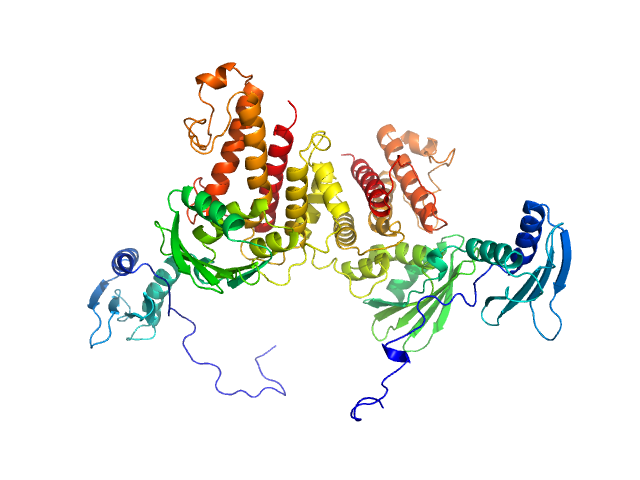
UniProt ID: P52129 (2-357) mRNA endoribonuclease toxin LS (D245R mutant)
|
|
|
|
| Sample: |
MRNA endoribonuclease toxin LS (D245R mutant) dimer, 84 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2020 Jul 18
|
Alternative dimerization is required for activity and inhibition of the HEPN ribonuclease RnlA
Nucleic Acids Research 49(12):7164-7178 (2021)
Garcia-Rodriguez G, Charlier D, Wilmaerts D, Michiels J, Loris R
|
| RgGuinier |
4.3 |
nm |
| Dmax |
15.9 |
nm |
| VolumePorod |
183 |
nm3 |
|
|