UniProt ID: P02768 (25-609) FcRn binding optimised human serum albumin V418M, T420A, E505G, V547A
UniProt ID: P61768 (21-119) neonatal Fc receptor ectodomain beta-microglogulin part with C-terminal His6 tag
UniProt ID: None (None-None) Albumin-binding side-chain
UniProt ID: P55899 (24-297) neonatal Fc receptor ectodomain alpha-chain
|
|
|
|
| Sample: |
FcRn binding optimised human serum albumin V418M, T420A, E505G, V547A monomer, 66 kDa Homo sapiens protein
Neonatal Fc receptor ectodomain beta-microglogulin part with C-terminal His6 tag monomer, 13 kDa Homo sapiens protein
Albumin-binding side-chain monomer, 1 kDa
Neonatal Fc receptor ectodomain alpha-chain monomer, 30 kDa Homo sapiens protein
|
| Buffer: |
100 mM MES, 100 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Nov 11
|
Identification of binding sites on human serum albumin for somapacitan - a long-acting growth hormone derivative.
Biochemistry (2020)
Johansson E, Nielsen AD, Demuth H, Wiberg C, Schjødt C, Huang T, Chen J, Jensen S, Petersen J, Thygesen P
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.6 |
nm |
| VolumePorod |
174 |
nm3 |
|
|
UniProt ID: P02768 (25-609) Human serum albumin
UniProt ID: P01241 (27-217) Somapacitan
|
|
|
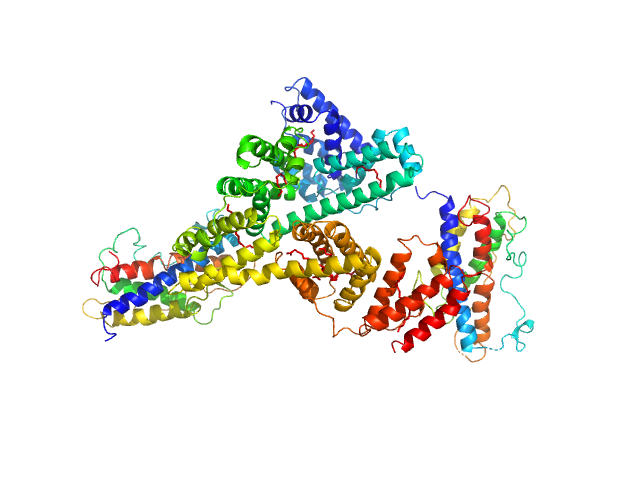
|
| Sample: |
Human serum albumin monomer, 66 kDa Homo sapiens protein
Somapacitan dimer, 44 kDa Homo sapiens protein
|
| Buffer: |
100 mM MES, 140 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, Novo Nordisk A/S on 2015 Sep 4
|
Identification of binding sites on human serum albumin for somapacitan - a long-acting growth hormone derivative.
Biochemistry (2020)
Johansson E, Nielsen AD, Demuth H, Wiberg C, Schjødt C, Huang T, Chen J, Jensen S, Petersen J, Thygesen P
|
| RgGuinier |
4.1 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
202 |
nm3 |
|
|
UniProt ID: P24785 (1-257) Dosage compensation regulator
|
|
|
|
| Sample: |
Dosage compensation regulator monomer, 29 kDa Drosophila melanogaster protein
|
| Buffer: |
20 mM NaPO4, 200 mM NaCl, 1 mM DTT, pH: 6.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Nov 29
|
Structure, dynamics and roX2-lncRNA binding of tandem double-stranded RNA binding domains dsRBD1,2 of Drosophila helicase Maleless.
Nucleic Acids Res 47(8):4319-4333 (2019)
Ankush Jagtap PK, Müller M, Masiewicz P, von Bülow S, Hollmann NM, Chen PC, Simon B, Thomae AW, Becker PB, Hennig J
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
UniProt ID: P24785 (1-257) Dosage compensation regulator
UniProt ID: None (None-None) roX2 stem-loop 7, 18-mer fragment
|
|
|
|
| Sample: |
Dosage compensation regulator monomer, 29 kDa Drosophila melanogaster protein
RoX2 stem-loop 7, 18-mer fragment monomer, 12 kDa synthetic construct RNA
|
| Buffer: |
20 mM NaPO4, 200 mM NaCl, 1 mM DTT, pH: 6.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Nov 29
|
Structure, dynamics and roX2-lncRNA binding of tandem double-stranded RNA binding domains dsRBD1,2 of Drosophila helicase Maleless.
Nucleic Acids Res 47(8):4319-4333 (2019)
Ankush Jagtap PK, Müller M, Masiewicz P, von Bülow S, Hollmann NM, Chen PC, Simon B, Thomae AW, Becker PB, Hennig J
|
| RgGuinier |
3.1 |
nm |
| Dmax |
13.3 |
nm |
| VolumePorod |
25 |
nm3 |
|
|
UniProt ID: Q03405 (None-None) Urokinase plasminogen activator surface receptor
UniProt ID: P00749 (1-143) Urokinase-type plasminogen activator (Amino Terminal Fragment)
|
|
|

|
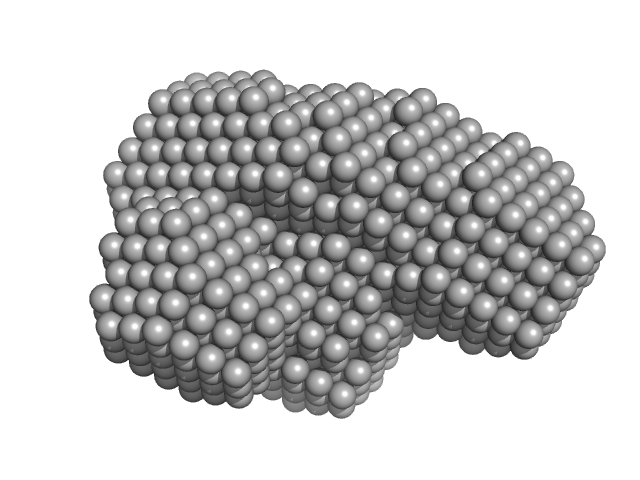
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
Urokinase-type plasminogen activator (Amino Terminal Fragment) monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
20 mM PBS, 5 %(v/v) glycerol, 50 mM NaSO4,, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2011 Jun 18
|
Did evolution create a flexible ligand-binding cavity in the urokinase receptor through deletion of a plesiotypic disulfide bond?
J Biol Chem (2019)
Leth JM, Mertens HDT, Leth-Espensen KZ, Jørgensen TJD, Ploug M
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
UniProt ID: Q03405 (None-None) Urokinase plasminogen activator surface receptor
|
|
|

|
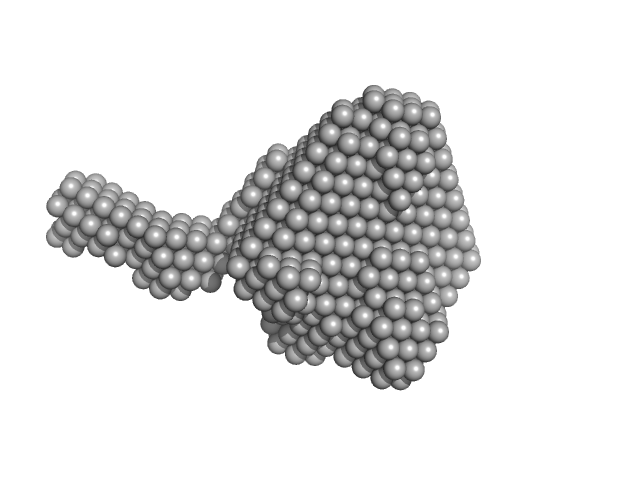
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
|
| Buffer: |
20 mM PBS, 5 %(v/v) glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Dec 1
|
Did evolution create a flexible ligand-binding cavity in the urokinase receptor through deletion of a plesiotypic disulfide bond?
J Biol Chem (2019)
Leth JM, Mertens HDT, Leth-Espensen KZ, Jørgensen TJD, Ploug M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
55 |
nm3 |
|
|
UniProt ID: Q03405 (None-None) Urokinase plasminogen activator surface receptor
UniProt ID: P00749 (1-143) Urokinase-type plasminogen activator (Amino Terminal Fragment)
|
|
|

|
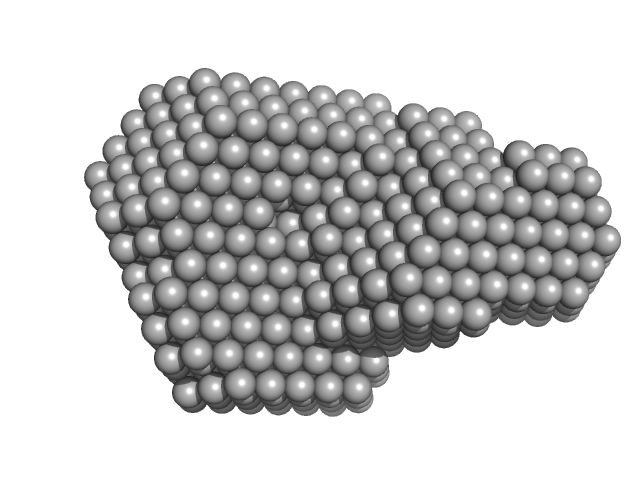
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
Urokinase-type plasminogen activator (Amino Terminal Fragment) monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
20 mM PBS, 5 %(v/v) glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Dec 1
|
Did evolution create a flexible ligand-binding cavity in the urokinase receptor through deletion of a plesiotypic disulfide bond?
J Biol Chem (2019)
Leth JM, Mertens HDT, Leth-Espensen KZ, Jørgensen TJD, Ploug M
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: Q03405 (None-None) Urokinase plasminogen activator surface receptor
|
|
|

|
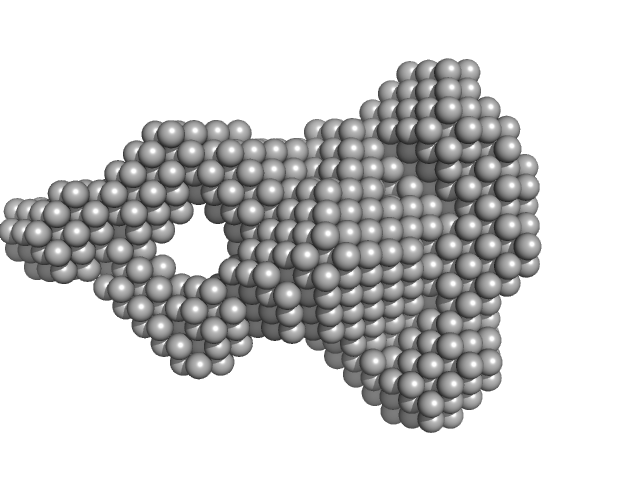
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
|
| Buffer: |
20 mM PBS, 5 %(v/v) glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 5
|
Did evolution create a flexible ligand-binding cavity in the urokinase receptor through deletion of a plesiotypic disulfide bond?
J Biol Chem (2019)
Leth JM, Mertens HDT, Leth-Espensen KZ, Jørgensen TJD, Ploug M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
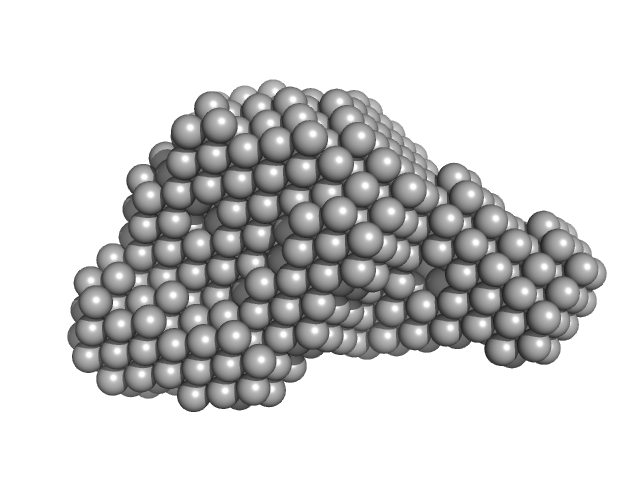
UniProt ID: Q03405 (None-None) Urokinase plasminogen activator surface receptor
UniProt ID: P00749 (1-143) Urokinase-type plasminogen activator (Amino Terminal Fragment)
|
|
|

|
| Sample: |
Urokinase plasminogen activator surface receptor monomer, 37 kDa Homo sapiens protein
Urokinase-type plasminogen activator (Amino Terminal Fragment) monomer, 16 kDa Homo sapiens protein
|
| Buffer: |
20 mM PBS, 5 %(v/v) glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 5
|
Did evolution create a flexible ligand-binding cavity in the urokinase receptor through deletion of a plesiotypic disulfide bond?
J Biol Chem (2019)
Leth JM, Mertens HDT, Leth-Espensen KZ, Jørgensen TJD, Ploug M
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
UniProt ID: P01137 (30-390) Human Latent Transforming Growth Factor beta 1
|
|
|
|
| Sample: |
Human Latent Transforming Growth Factor beta 1 dimer, 86 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Oct 4
|
Structural consequences of transforming growth factor beta-1 activation from near-therapeutic X-ray doses.
J Synchrotron Radiat 26(Pt 4):967-979 (2019)
Stachowski T, Grant TD, Snell EH
|
| RgGuinier |
3.8 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
200 |
nm3 |
|
|