UniProt ID: P11532 (1990-3040) Human dystrophin central domain R16-24 fragment
|
|
|
|
| Sample: |
Human dystrophin central domain R16-24 fragment monomer, 127 kDa protein
|
| Buffer: |
NaP 20 mM, NaCl 300 mM, EDTA 1 mM, Glycérol 2%, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2016 Dec 17
|
Dystrophin SAXS data
Raphael Dos Santos Morais
|
| RgGuinier |
9.3 |
nm |
| Dmax |
34.0 |
nm |
|
|
UniProt ID: P11532 (1990-3040) Human dystrophin central domain R16-24 del45-55 fragment
|
|
|
|
| Sample: |
Human dystrophin central domain R16-24 del45-55 fragment monomer, 58 kDa protein
|
| Buffer: |
NaP 20 mM, NaCl 300 mM, EDTA 1 mM, Glycérol 2%, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Sep 28
|
Dystrophin SAXS data
Raphael Dos Santos Morais
|
| RgGuinier |
4.7 |
nm |
| Dmax |
18.3 |
nm |
|
|
UniProt ID: P11532 (1990-3040) Human dystrophin central domain R16-24 del45-51 fragment
|
|
|
|
| Sample: |
Human dystrophin central domain R16-24 del45-51 fragment monomer, 85 kDa protein
|
| Buffer: |
NaP 20 mM, NaCl 300 mM, EDTA 1 mM, Glycérol 2%, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Jul 13
|
Dystrophin SAXS data
Raphael Dos Santos Morais
|
| RgGuinier |
7.0 |
nm |
| Dmax |
26.2 |
nm |
|
|
UniProt ID: P21224 (30-338) Leukocidin G
UniProt ID: Q5HEH9 (28-351) Leukocidin H
UniProt ID: None (None-None) LukGH neutralizing antibody
UniProt ID: P11215 (143-337) Integrin alpha-M
|
|
|
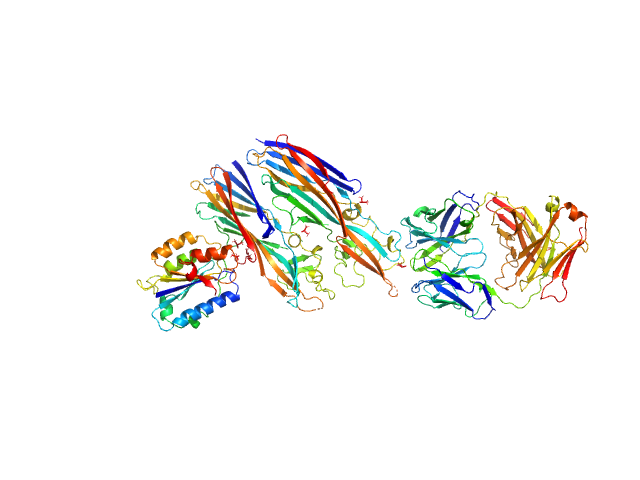
|
| Sample: |
Leukocidin G monomer, 36 kDa Staphylococcus aureus protein
Leukocidin H monomer, 38 kDa Staphylococcus aureus protein
LukGH neutralizing antibody monomer, 47 kDa Homo sapiens protein
Integrin alpha-M monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM Hepes, 300 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Sep 17
|
Molecular mechanism of leukocidin GH-integrin CD11b/CD18 recognition and species specificity.
Proc Natl Acad Sci U S A (2019)
Trstenjak N, Milić D, Graewert MA, Rouha H, Svergun D, Djinović-Carugo K, Nagy E, Badarau A
|
| RgGuinier |
5.0 |
nm |
| Dmax |
18.0 |
nm |
| VolumePorod |
178 |
nm3 |
|
|
UniProt ID: P21224 (30-338) Leukocidin G
UniProt ID: Q5HEH9 (28-351) Leukocidin H
UniProt ID: None (None-None) LukGH neutralizing antibody
|
|
|
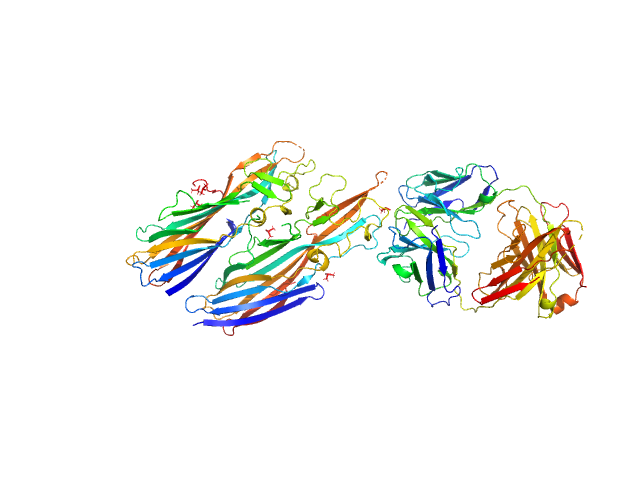
|
| Sample: |
Leukocidin G monomer, 36 kDa Staphylococcus aureus protein
Leukocidin H monomer, 38 kDa Staphylococcus aureus protein
LukGH neutralizing antibody monomer, 47 kDa Homo sapiens protein
|
| Buffer: |
20 mM Hepes, 300 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Sep 17
|
Molecular mechanism of leukocidin GH-integrin CD11b/CD18 recognition and species specificity.
Proc Natl Acad Sci U S A (2019)
Trstenjak N, Milić D, Graewert MA, Rouha H, Svergun D, Djinović-Carugo K, Nagy E, Badarau A
|
| RgGuinier |
4.7 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
152 |
nm3 |
|
|
UniProt ID: Q5E9J8 (1-492) Resistance to inhibitors of cholinesterase 8 homolog A
|
|
|
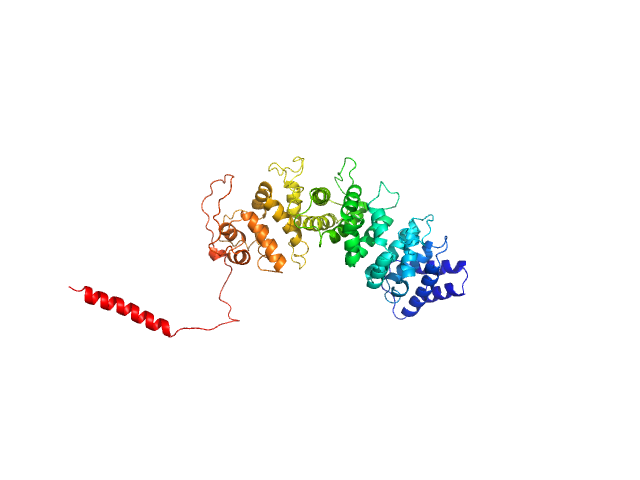
|
| Sample: |
Resistance to inhibitors of cholinesterase 8 homolog A monomer, 56 kDa Bos taurus protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 5 % glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Oct 27
|
Structural underpinnings of Ric8A function as a G-protein α-subunit chaperone and guanine-nucleotide exchange factor.
Nat Commun 10(1):3084 (2019)
Srivastava D, Gakhar L, Artemyev NO
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
88 |
nm3 |
|
|
UniProt ID: Q5E9J8 (1-452) Resistance to inhibitors of cholinesterase 8 homolog A
|
|
|
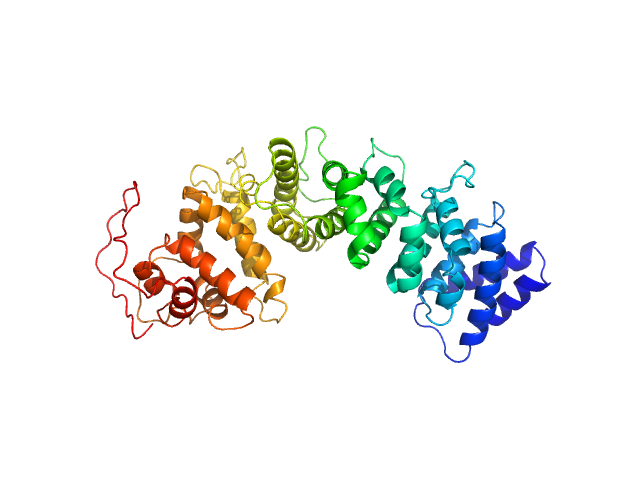
|
| Sample: |
Resistance to inhibitors of cholinesterase 8 homolog A monomer, 51 kDa Bos taurus protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 5 % glycerol, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2018 Oct 27
|
Structural underpinnings of Ric8A function as a G-protein α-subunit chaperone and guanine-nucleotide exchange factor.
Nat Commun 10(1):3084 (2019)
Srivastava D, Gakhar L, Artemyev NO
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.2 |
nm |
| VolumePorod |
70 |
nm3 |
|
|
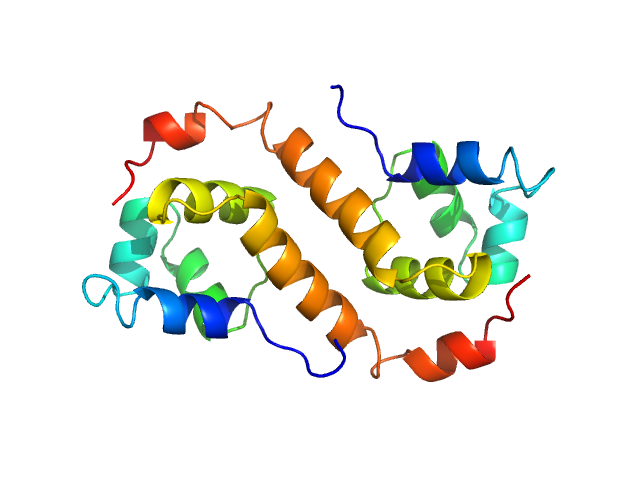
UniProt ID: Q9HVC1 (None-None) Uncharacterized protein
|
|
|
|
| Sample: |
Uncharacterized protein dimer, 22 kDa Pseudomonas aeruginosa protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 5% (v/v) glycerol, and 1 mM PMSF, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Dec 21
|
Structural Insights Into the Transcriptional Regulation of HigBA Toxin–Antitoxin System by Antitoxin HigA in Pseudomonas aeruginosa
Frontiers in Microbiology 10 (2020)
Liu Y, Gao Z, Liu G, Geng Z, Dong Y, Zhang H
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
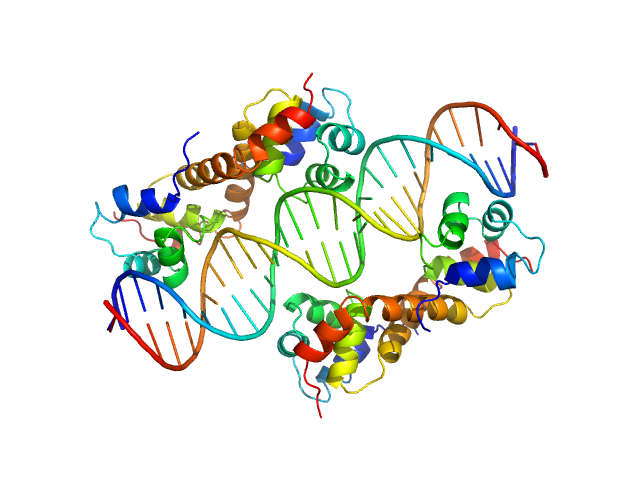
UniProt ID: Q9HVC1 (None-None) Uncharacterized protein
UniProt ID: None (None-None) DNA Duplex
|
|
|
|
| Sample: |
Uncharacterized protein dimer, 22 kDa Pseudomonas aeruginosa protein
DNA Duplex dimer, 20 kDa DNA
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 5% (v/v) glycerol, and 1 mM PMSF, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Sep 19
|
Structural Insights Into the Transcriptional Regulation of HigBA Toxin–Antitoxin System by Antitoxin HigA in Pseudomonas aeruginosa
Frontiers in Microbiology 10 (2020)
Liu Y, Gao Z, Liu G, Geng Z, Dong Y, Zhang H
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
81 |
nm3 |
|
|
UniProt ID: B1H241 (1-452) Resistance to inhibitors of cholinesterase 8 homolog A
|
|
|
|
| Sample: |
Resistance to inhibitors of cholinesterase 8 homolog A monomer, 51 kDa Rattus norvegicus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2018 Apr 24
|
Structure, Function, and Dynamics of the Gα Binding Domain of Ric-8A.
Structure (2019)
Zeng B, Mou TC, Doukov TI, Steiner A, Yu W, Papasergi-Scott M, Tall GG, Hagn F, Sprang SR
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
70 |
nm3 |
|
|