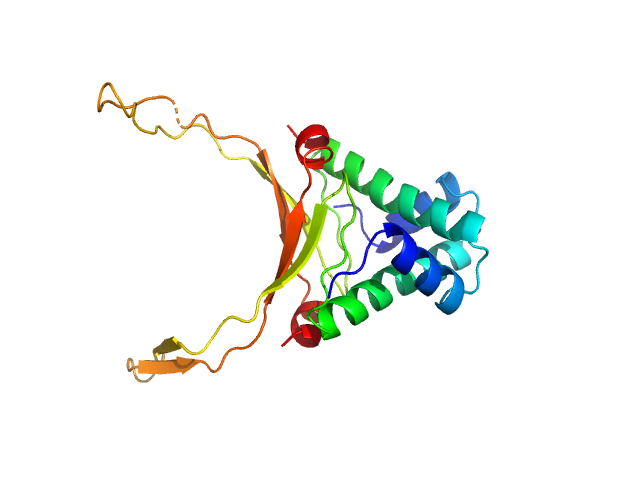
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha, E38K/V42L double mutant
|
|
|
|
| Sample: |
DNA-binding protein HU-alpha, E38K/V42L double mutant decamer, 95 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM DTT, 1 mM PMSF, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Apr 23
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling (supplementary)
Soumya G Remesh
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
53 |
nm3 |
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha, E38K/V42L double mutant
|
|
|
|
| Sample: |
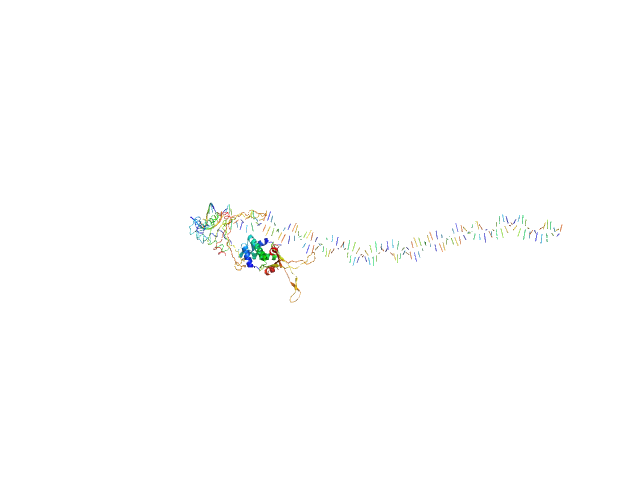
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha, E38K/V42L double mutant tetramer, 38 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM DTT, 1 mM PMSF, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Apr 23
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling (supplementary)
Soumya G Remesh
|
| RgGuinier |
5.7 |
nm |
| Dmax |
31.3 |
nm |
| VolumePorod |
297 |
nm3 |
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha, E38K/V42L double mutant
|
|
|
|
| Sample: |
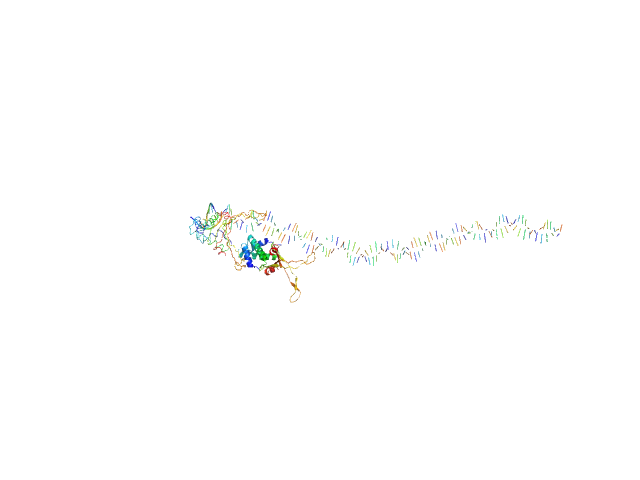
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha, E38K/V42L double mutant tetramer, 38 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM DTT, 1 mM PMSF, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Apr 23
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling (supplementary)
Soumya G Remesh
|
| RgGuinier |
5.7 |
nm |
| Dmax |
25.7 |
nm |
| VolumePorod |
195 |
nm3 |
|
|
UniProt ID: None (None-None) 80bp_DNA Forward
UniProt ID: None (None-None) 80bp_DNA Reverse
UniProt ID: P0ACF0 (1-90) DNA-binding protein HU-alpha, E38K/V42L double mutant
|
|
|

|
| Sample: |
80bp_DNA Forward monomer, 25 kDa Escherichia coli DNA
80bp_DNA Reverse monomer, 25 kDa Escherichia coli DNA
DNA-binding protein HU-alpha, E38K/V42L double mutant octamer, 76 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl, 150 mM NaCl, 1 mM DTT, 1 mM PMSF, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Apr 23
|
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling (supplementary)
Soumya G Remesh
|
| RgGuinier |
5.8 |
nm |
| Dmax |
28.1 |
nm |
| VolumePorod |
296 |
nm3 |
|
|
UniProt ID: Q9WTL8-2 (530-625) Aryl hydrocarbon receptor nuclear translocator-like protein 1
|
|
|
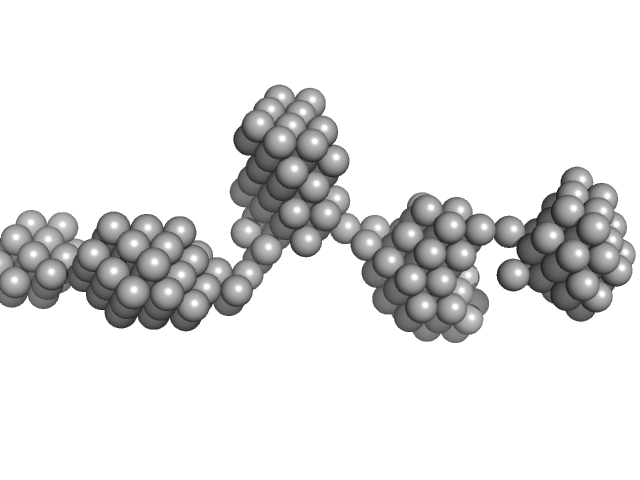
|
| Sample: |
Aryl hydrocarbon receptor nuclear translocator-like protein 1 monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
2.8 |
nm |
| Dmax |
11.0 |
nm |
|
|
UniProt ID: P45481 (586-672) Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C
|
|
|
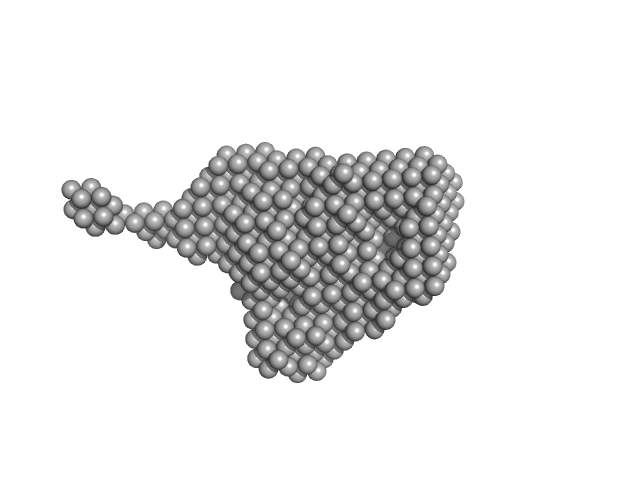

|
| Sample: |
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
1.9 |
nm |
| Dmax |
8.1 |
nm |
|
|
UniProt ID: Q9WTL8-2 (530-625) Aryl hydrocarbon receptor nuclear translocator-like protein 1
UniProt ID: P45481 (586-672) Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C
|
|
|
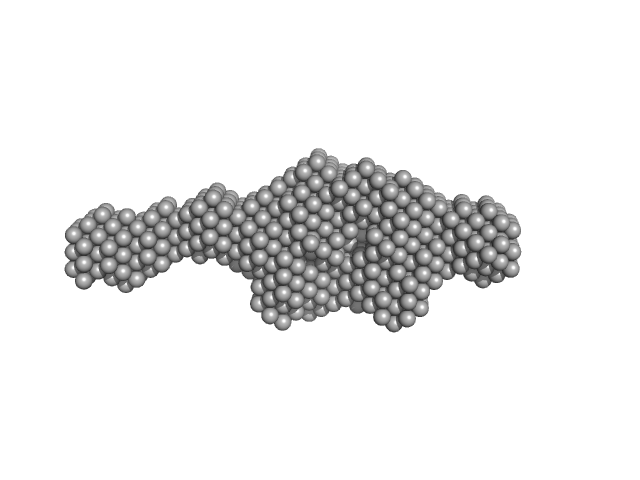

|
| Sample: |
Aryl hydrocarbon receptor nuclear translocator-like protein 1 monomer, 10 kDa Mus musculus protein
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
2.8 |
nm |
| Dmax |
11.5 |
nm |
|
|
UniProt ID: Q9WTL8-2 (517-625) Brain and muscle ARNT-like 1 (G517-L625) including transactivation domain (TAD)
UniProt ID: P45481 (586-672) Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP)
|
|
|
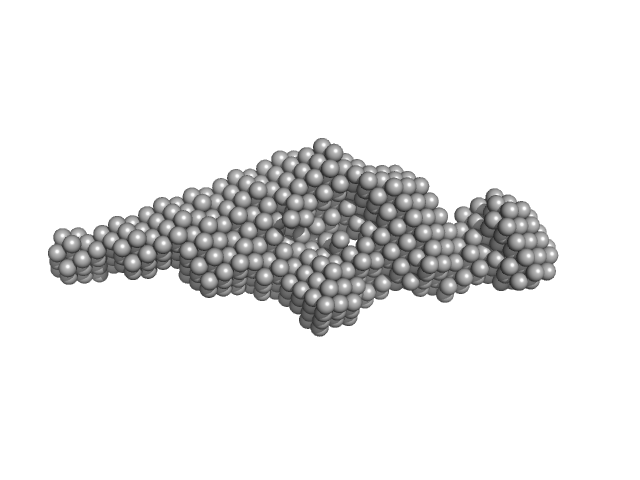

|
| Sample: |
Brain and muscle ARNT-like 1 (G517-L625) including transactivation domain (TAD) monomer, 11 kDa Mus musculus protein
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP) monomer, 10 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
3.1 |
nm |
| Dmax |
13.7 |
nm |
|
|
UniProt ID: P45481 (586-672) Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP)
UniProt ID: Q9WTL8-2 (490-625) Brain and muscle ARNT-like 1 (G490-L625) including transactivation domain (TAD)
|
|
|
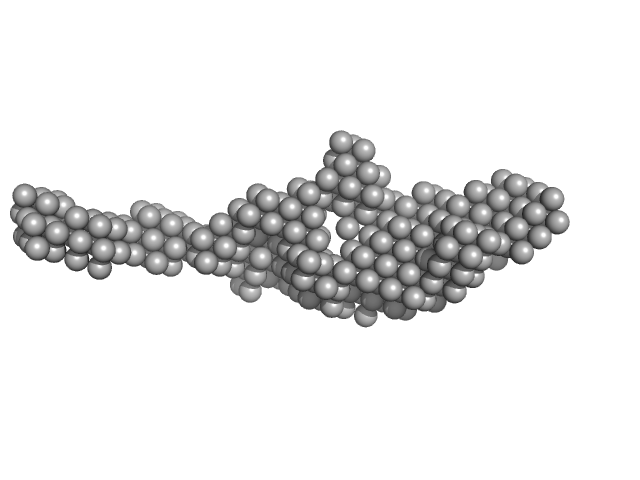

|
| Sample: |
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP) monomer, 10 kDa Mus musculus protein
Brain and muscle ARNT-like 1 (G490-L625) including transactivation domain (TAD) monomer, 14 kDa Mus musculus protein
|
| Buffer: |
25 mM Hepes, 150 NaCl, 1 mM DTT, 5% Glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
3.8 |
nm |
| Dmax |
17.5 |
nm |
|
|
UniProt ID: P45481 (586-672) Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C
UniProt ID: None (None-None) 1-{4-[4-chloro-3-(trifluoromethyl)phenyl]-4-hydroxypiperidin-1-yl}-3-sulfanylpropan-1-one
|
|
|
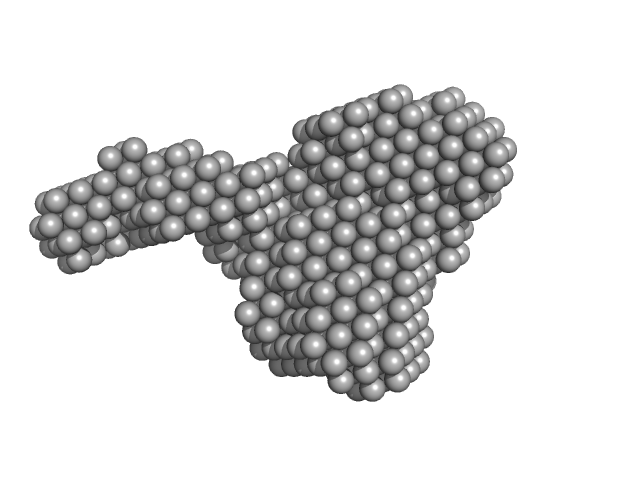

|
| Sample: |
Kinase-inducible domain interacting (KIX) domain of CREB-binding protein (CBP), mutation L664C monomer, 10 kDa Mus musculus protein
1-{4-[4-chloro-3-(trifluoromethyl)phenyl]-4-hydroxypiperidin-1-yl}-3-sulfanylpropan-1-one monomer, 0 kDa
|
| Buffer: |
25 mM Hepes, 150 NaCl, 5% Glycerol, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 27
|
Structural and mechanistic insights into the interaction of the circadian transcription factor BMAL1 with the KIX domain of the CREB-binding protein.
J Biol Chem (2019)
Garg A, Orru R, Ye W, Distler U, Chojnacki JE, Köhn M, Tenzer S, Sönnichsen C, Wolf E
|
| RgGuinier |
1.7 |
nm |
| Dmax |
6.4 |
nm |
|
|