UniProt ID: M9NVD3 (373-1296) Flagella binding tail protein
|
|
|
|
| Sample: |
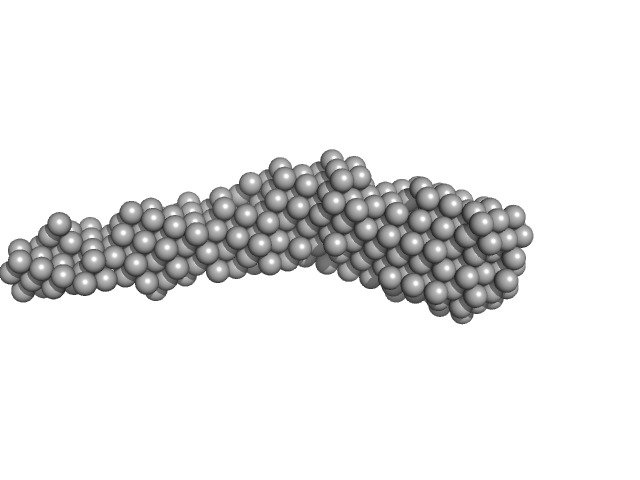
Flagella binding tail protein monomer, 103 kDa Salmonella virus Chi protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.03 % NaN3, 5.0 % glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2017 Apr 4
|
The flagellotropic bacteriophage YSD1 targets Salmonella Typhi with a Chi-like protein tail fibre.
Mol Microbiol (2019)
Dunstan RA, Pickard D, Dougan S, Goulding D, Cormie C, Hardy J, Li F, Grinter R, Harcourt K, Yu L, Song J, Schreiber F, Choudhary J, Clare S, Coulibaly F, Strugnell RA, Dougan G, Lithgow T
|
| RgGuinier |
5.6 |
nm |
| Dmax |
27.4 |
nm |
| VolumePorod |
155 |
nm3 |
|
|
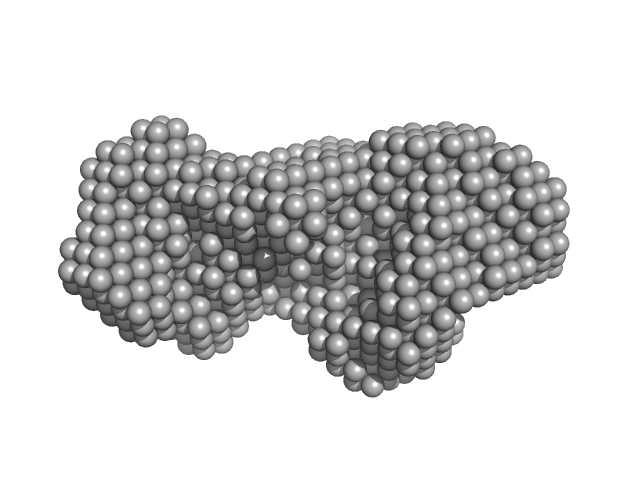
UniProt ID: P31828 (27-931) Zinc protease PqqL
|
|
|

|
| Sample: |
Zinc protease PqqL monomer, 102 kDa Escherichia coli protein
|
| Buffer: |
20 mM Tris HCl, 150 nM NaCl, 0.02 % NaN3, 5% glycerol, pH: 7.8 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2017 Jun 14
|
Protease-associated import systems are widespread in Gram-negative bacteria.
PLoS Genet 15(10):e1008435 (2019)
Grinter R, Leung PM, Wijeyewickrema LC, Littler D, Beckham S, Pike RN, Walker D, Greening C, Lithgow T
|
| RgGuinier |
4.0 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
207 |
nm3 |
|
|
UniProt ID: Q93KQ6 (89-729) Protein kinase YopO
|
|
|
|
| Sample: |
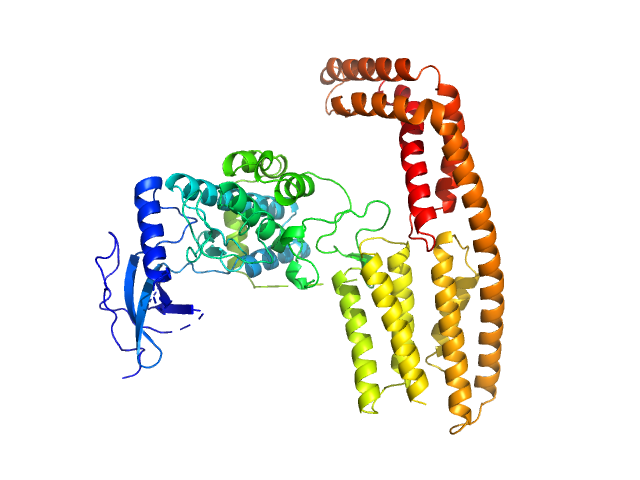
Protein kinase YopO monomer, 63 kDa Yersinia enterocolitica protein
|
| Buffer: |
10 mM Tris-HCl, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 6
|
Studying Conformational Changes of the Yersinia Type-III-Secretion Effector YopO in Solution by Integrative Structural Biology.
Structure 27(9):1416-1426.e3 (2019)
Peter MF, Tuukkanen AT, Heubach CA, Selsam A, Duthie FG, Svergun DI, Schiemann O, Hagelueken G
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
119 |
nm3 |
|
|
UniProt ID: Q93KQ6 (89-729) Protein kinase YopO
UniProt ID: P60709 (1-375) Actin, cytoplasmic 1
|
|
|
|
| Sample: |
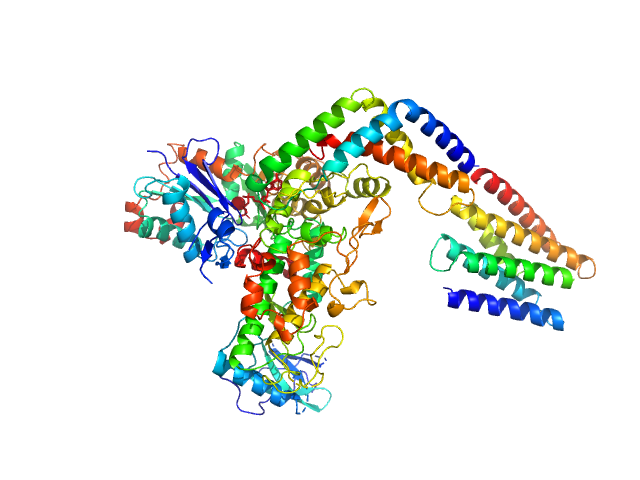
Protein kinase YopO monomer, 63 kDa Yersinia enterocolitica protein
Actin, cytoplasmic 1 monomer, 42 kDa Homo sapiens protein
|
| Buffer: |
10 mM Tris-HCl, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 May 6
|
Studying Conformational Changes of the Yersinia Type-III-Secretion Effector YopO in Solution by Integrative Structural Biology.
Structure 27(9):1416-1426.e3 (2019)
Peter MF, Tuukkanen AT, Heubach CA, Selsam A, Duthie FG, Svergun DI, Schiemann O, Hagelueken G
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.3 |
nm |
| VolumePorod |
147 |
nm3 |
|
|
UniProt ID: P9WNU1 (1-402) Beta sliding clamp
|
|
|
|
| Sample: |
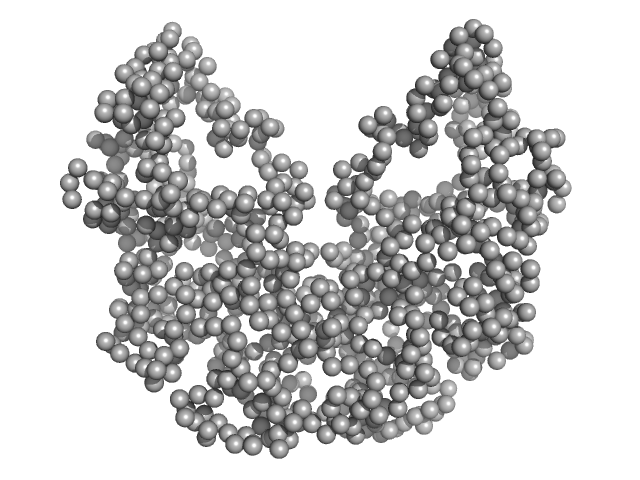
Beta sliding clamp dimer, 86 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris pH 8.0, 200 mM NaCl , 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 15
|
Regulation of futile ligation during early steps of BER in M. tuberculosis is carried out by a β-clamp-XthA-LigA tri-component complex
International Journal of Biological Macromolecules (2022)
Shukla A, Afsar M, Khanam T, Kumar N, Ali F, Kumar S, Jahan F, Ramachandran R
|
| RgGuinier |
3.7 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
186 |
nm3 |
|
|
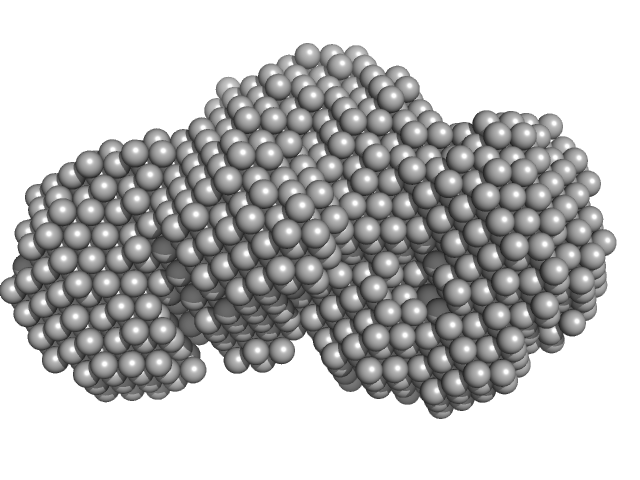
UniProt ID: P9WNU1 (1-402) Beta sliding clamp
UniProt ID: P96273 (1-291) Probable exodeoxyribonuclease III protein XthA (Exonuclease III) (EXO III) (AP endonuclease VI)
|
|
|

|
| Sample: |
Beta sliding clamp dimer, 86 kDa Mycobacterium tuberculosis protein
Probable exodeoxyribonuclease III protein XthA (Exonuclease III) (EXO III) (AP endonuclease VI) monomer, 33 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 15
|
Regulation of futile ligation during early steps of BER in M. tuberculosis is carried out by a β-clamp-XthA-LigA tri-component complex
International Journal of Biological Macromolecules (2022)
Shukla A, Afsar M, Khanam T, Kumar N, Ali F, Kumar S, Jahan F, Ramachandran R
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.3 |
nm |
| VolumePorod |
98 |
nm3 |
|
|
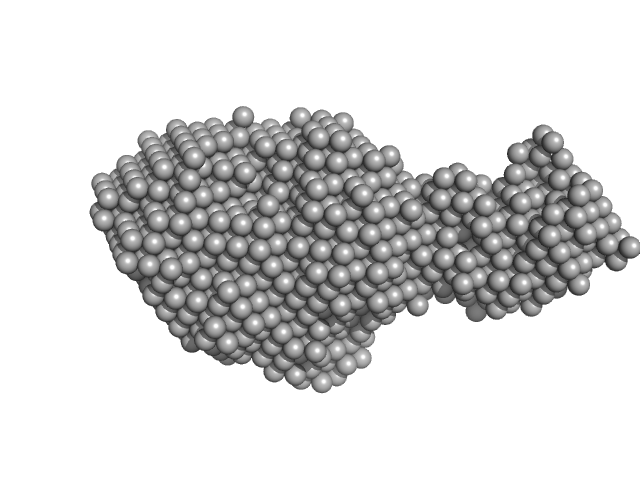
UniProt ID: P9WNU1 (1-402) Beta sliding clamp
UniProt ID: A0A0T9L251 (1-289) Probable exodeoxyribonuclease III protein XthA
UniProt ID: P9WNV1 (1-691) DNA ligase A
|
|
|

|
| Sample: |
Beta sliding clamp dimer, 86 kDa Mycobacterium tuberculosis protein
Probable exodeoxyribonuclease III protein XthA monomer, 31 kDa Mycobacterium tuberculosis protein
DNA ligase A monomer, 76 kDa Mycobacterium tuberculosis protein
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 17
|
Regulation of futile ligation during early steps of BER in M. tuberculosis is carried out by a β-clamp-XthA-LigA tri-component complex
International Journal of Biological Macromolecules (2022)
Shukla A, Afsar M, Khanam T, Kumar N, Ali F, Kumar S, Jahan F, Ramachandran R
|
| RgGuinier |
5.8 |
nm |
| Dmax |
25.3 |
nm |
| VolumePorod |
501 |
nm3 |
|
|
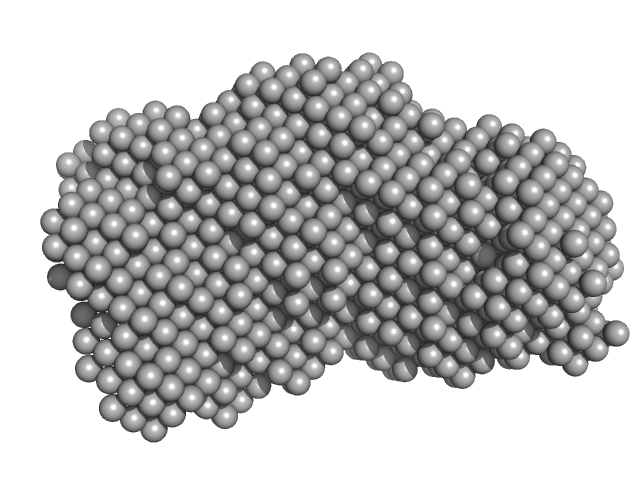
UniProt ID: A0A0T9L251 (1-289) Probable exodeoxyribonuclease III protein XthA
UniProt ID: P9WNV1 (None-None) DNA ligase A
UniProt ID: P9WNU1 (1-402) Beta sliding clamp
UniProt ID: None (None-None) DNA ligase A nicked DNA substrate
|
|
|

|
| Sample: |
Probable exodeoxyribonuclease III protein XthA monomer, 31 kDa Mycobacterium tuberculosis protein
DNA ligase A monomer, 76 kDa Mycobacterium tuberculosis protein
Beta sliding clamp dimer, 86 kDa Mycobacterium tuberculosis protein
DNA ligase A nicked DNA substrate dimer, 16 kDa DNA
|
| Buffer: |
50 mM Tris pH 8.0, 200 mM NaCl , 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 17
|
Regulation of futile ligation during early steps of BER in M. tuberculosis is carried out by a β-clamp-XthA-LigA tri-component complex
International Journal of Biological Macromolecules (2022)
Shukla A, Afsar M, Khanam T, Kumar N, Ali F, Kumar S, Jahan F, Ramachandran R
|
| RgGuinier |
5.8 |
nm |
| Dmax |
19.1 |
nm |
| VolumePorod |
496 |
nm3 |
|
|
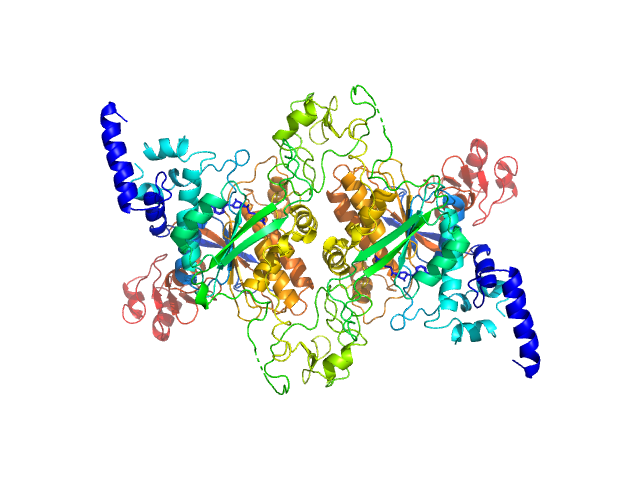
UniProt ID: O25315 (1-598) Adenine specific DNA methyltransferase (Mod)
|
|
|
|
| Sample: |
Adenine specific DNA methyltransferase (Mod) dimer, 137 kDa Helicobacter pylori protein
|
| Buffer: |
25 mM Tris, 250 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at ID14-3, ESRF on 2017 Jul 9
|
Tetramerization at low pH licenses DNA methylation activity of M.HpyAXI in the presence of acid stress.
J Mol Biol (2019)
Narayanan N, Banerjee A, Jain D, Kulkarni DS, Sharma R, Nirwal S, Rao DN, Nair DT
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
143 |
nm3 |
|
|
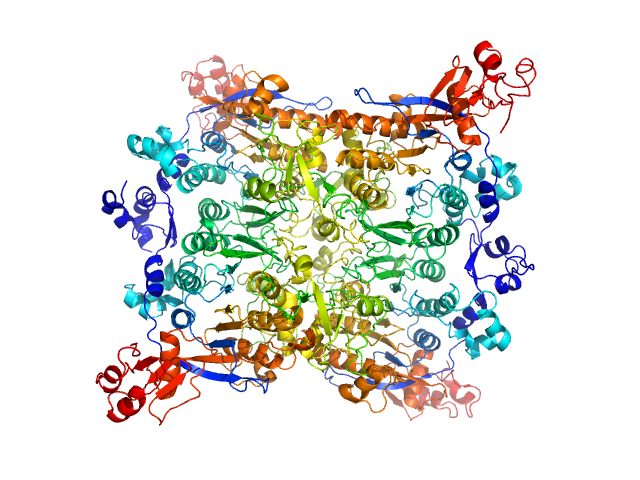
UniProt ID: O25315 (1-598) Adenine specific DNA methyltransferase (Mod)
|
|
|
|
| Sample: |
Adenine specific DNA methyltransferase (Mod) tetramer, 273 kDa Helicobacter pylori protein
|
| Buffer: |
25 mM citrate, 250 mM NaCl, pH: 5.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Dec 1
|
Tetramerization at low pH licenses DNA methylation activity of M.HpyAXI in the presence of acid stress.
J Mol Biol (2019)
Narayanan N, Banerjee A, Jain D, Kulkarni DS, Sharma R, Nirwal S, Rao DN, Nair DT
|
| RgGuinier |
5.0 |
nm |
| Dmax |
19.1 |
nm |
| VolumePorod |
316 |
nm3 |
|
|