|
|
|
|
|
| Sample: |
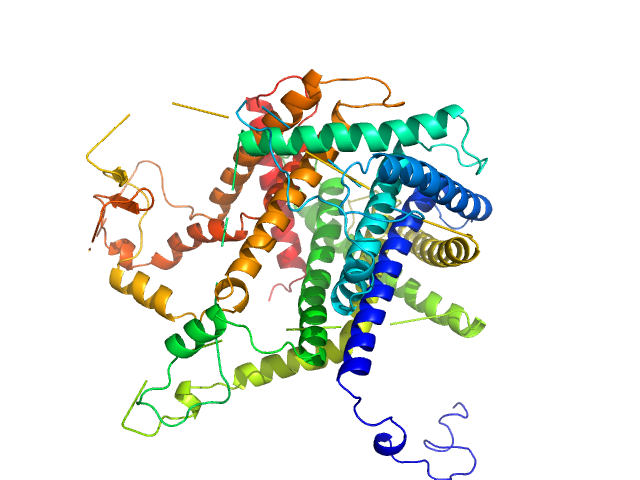
Mitochondrial import inner membrane translocase subunit TIM8 trimer, 29 kDa Saccharomyces cerevisiae protein
Mitochondrial import inner membrane translocase subunit TIM13 trimer, 34 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Apr 21
|
Structural basis of client specificity in mitochondrial membrane-protein chaperones.
Sci Adv 6(51) (2020)
Sučec I, Wang Y, Dakhlaoui O, Weinhäupl K, Jores T, Costa D, Hessel A, Brennich M, Rapaport D, Lindorff-Larsen K, Bersch B, Schanda P
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.9 |
nm |
| VolumePorod |
136 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
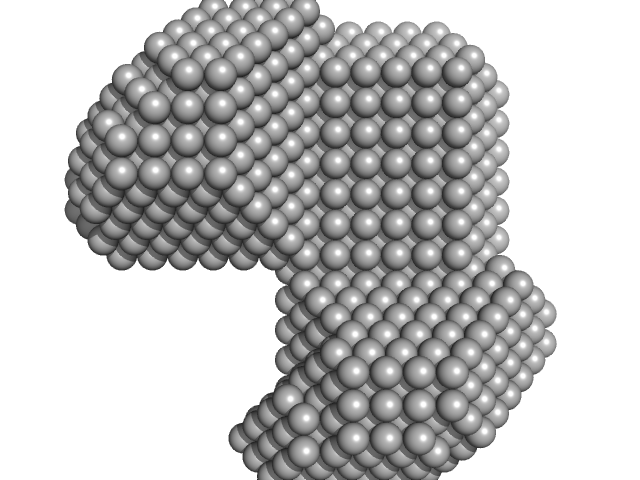
LIM/homeobox protein Lhx3 monomer, 10 kDa Mus musculus protein
|
| Buffer: |
20 mM sodium phosphate monobasic/dibasic, 100 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Nov 2
|
Contrasting DNA-binding behaviour by ISL1 and LHX3 underpins differential gene targeting in neuronal cell specification
Journal of Structural Biology: X :100043 (2020)
Smith N, Wilkinson-White L, Kwan A, Trewhella J, Matthews J
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
|
|
|
|
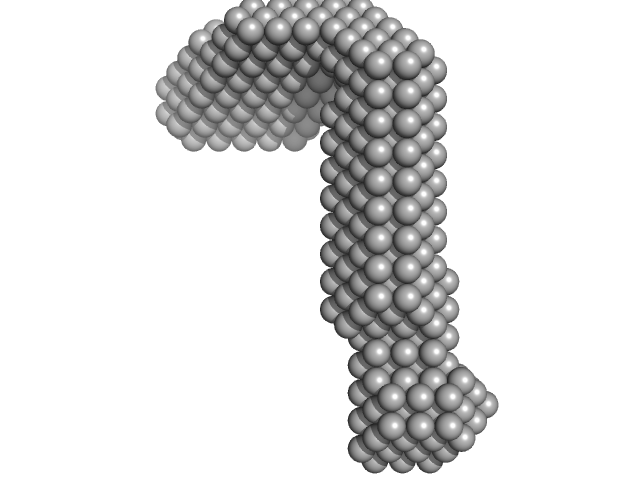
|
| Sample: |
LIM/homeobox protein Lhx3 monomer, 23 kDa Mus musculus protein
Insulin gene enhancer protein ISL-1 monomer, 4 kDa Mus musculus protein
|
| Buffer: |
20 mM sodium phosphate monobasic/dibasic, 100 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Nov 2
|
Contrasting DNA-binding behaviour by ISL1 and LHX3 underpins differential gene targeting in neuronal cell specification
Journal of Structural Biology: X :100043 (2020)
Smith N, Wilkinson-White L, Kwan A, Trewhella J, Matthews J
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
39 |
nm3 |
|
|
|
|
|
|
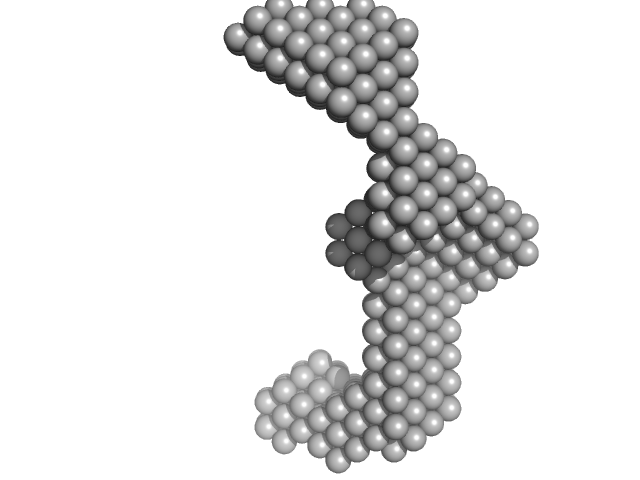
|
| Sample: |
Insulin gene enhancer protein ISL-1 monomer, 12 kDa Mus musculus protein
LIM/homeobox protein Lhx3 monomer, 9 kDa Mus musculus protein
|
| Buffer: |
20 mM sodium phosphate monobasic/dibasic, 100 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Nov 2
|
Contrasting DNA-binding behaviour by ISL1 and LHX3 underpins differential gene targeting in neuronal cell specification
Journal of Structural Biology: X :100043 (2020)
Smith N, Wilkinson-White L, Kwan A, Trewhella J, Matthews J
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
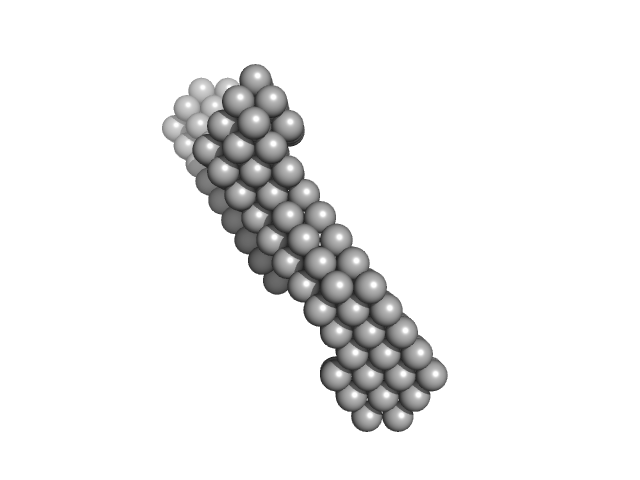
Insulin gene enhancer protein ISL-1 monomer, 14 kDa Mus musculus protein
LIM/homeobox protein Lhx3 monomer, 23 kDa Mus musculus protein
|
| Buffer: |
20 mM sodium phosphate monobasic/dibasic, 100 mM NaCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Nov 2
|
Contrasting DNA-binding behaviour by ISL1 and LHX3 underpins differential gene targeting in neuronal cell specification
Journal of Structural Biology: X :100043 (2020)
Smith N, Wilkinson-White L, Kwan A, Trewhella J, Matthews J
|
| RgGuinier |
3.4 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
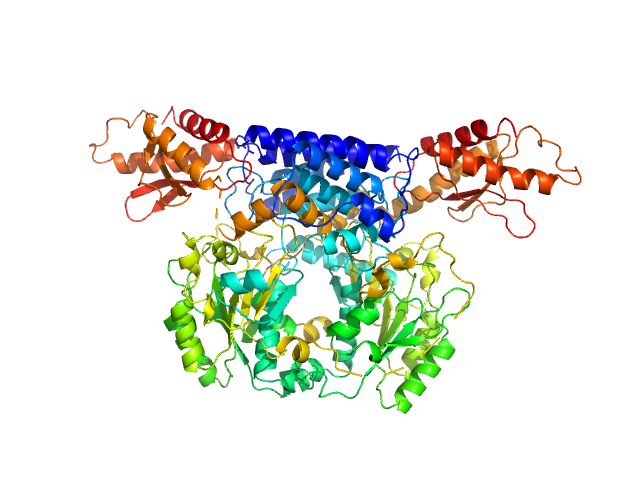
Cysteine sulfinic acid decarboxylase dimer, 117 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Feb 4
|
Structure and substrate specificity determinants of the taurine biosynthetic enzyme cysteine sulphinic acid decarboxylase.
J Struct Biol :107674 (2020)
Mahootchi E, Raasakka A, Luan W, Muruganandam G, Loris R, Haavik J, Kursula P
|
| RgGuinier |
3.4 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
198 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Modular nanotransporter with a melanocyte stimulating hormone ligand module monomer, 70 kDa synthetic construct protein
|
| Buffer: |
150 mM NaCl, 10 mM phosphate buffer saline, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Nov 24
|
Low-resolution structures of modular nanotransporters shed light on their functional activity
Acta Crystallographica Section D Structural Biology 76(12):1270-1279 (2020)
Khramtsov Y, Vlasova A, Vlasov A, Rosenkranz A, Ulasov A, Ryzhykau Y, Kuklin A, Orekhov A, Eydlin I, Georgiev G, Gordeliy V, Sobolev A
|
| RgGuinier |
3.7 |
nm |
| Dmax |
13.3 |
nm |
| VolumePorod |
127 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
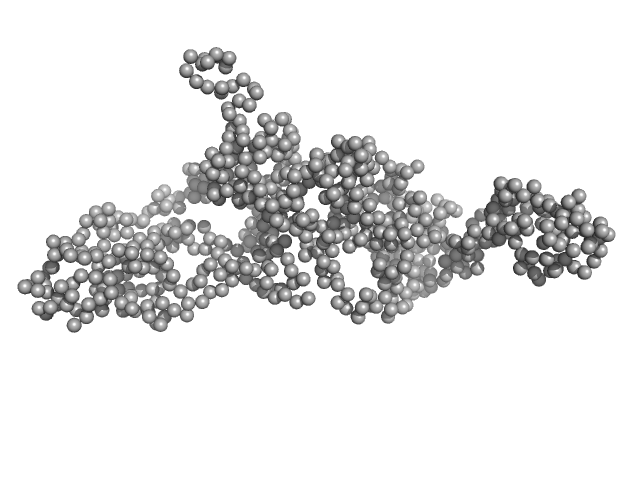
Modular nanotransporter with an epidermal growth factor ligand module monomer, 76 kDa synthetic construct protein
|
| Buffer: |
150 mM NaCl, 10 mM phosphate buffer saline, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Nov 24
|
Low-resolution structures of modular nanotransporters shed light on their functional activity
Acta Crystallographica Section D Structural Biology 76(12):1270-1279 (2020)
Khramtsov Y, Vlasova A, Vlasov A, Rosenkranz A, Ulasov A, Ryzhykau Y, Kuklin A, Orekhov A, Eydlin I, Georgiev G, Gordeliy V, Sobolev A
|
| RgGuinier |
4.2 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
161 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
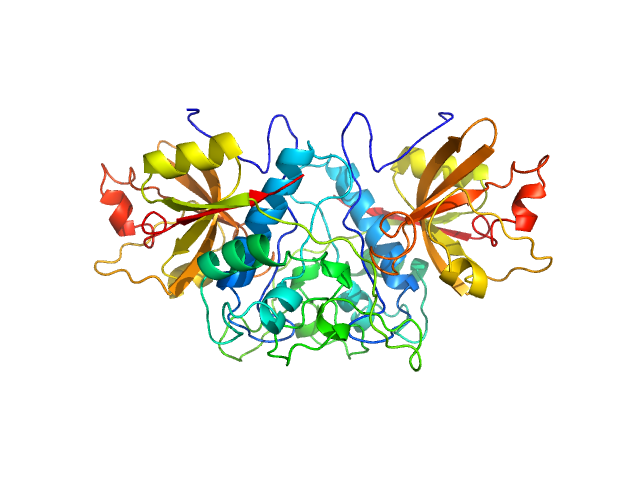
Cathepsin Z dimer, 54 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium acetate, 1 mM EDTA, pH: 5.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Dec 6
|
Human cathepsin X/Z is a biologically active homodimer.
Biochim Biophys Acta Proteins Proteom :140567 (2020)
Dolenc I, Štefe I, Turk D, Taler-Verčič A, Turk B, Turk V, Stoka V
|
| RgGuinier |
2.6 |
nm |
| Dmax |
6.5 |
nm |
| VolumePorod |
89 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
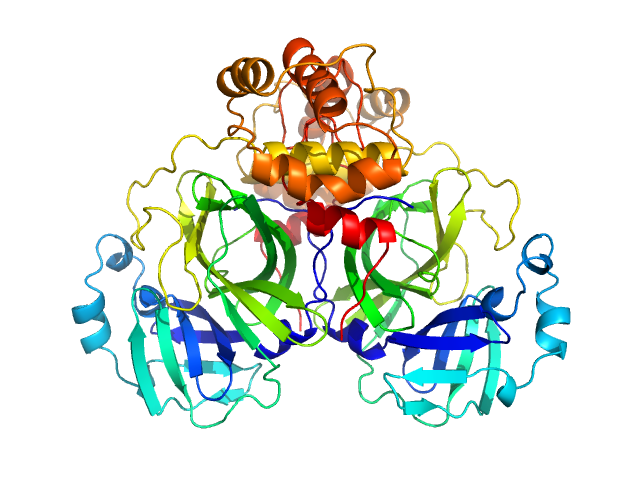
3C-like proteinase from SARS-CoV-2 replicase polyprotein 1a dimer, 68 kDa Severe acute respiratory … protein
|
| Buffer: |
50 mM Tris, 1 mM DTT, 1 mM EDTA, pH: 7.4 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, University of British Columbia on 2020 Jun 1
|
Crystallographic structure of wild-type SARS-CoV-2 main protease acyl-enzyme intermediate with physiological C-terminal autoprocessing site.
Nat Commun 11(1):5877 (2020)
Lee J, Worrall LJ, Vuckovic M, Rosell FI, Gentile F, Ton AT, Caveney NA, Ban F, Cherkasov A, Paetzel M, Strynadka NCJ
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.8 |
nm |
| VolumePorod |
93 |
nm3 |
|
|