|
|
|
|
|
| Sample: |
CDNA FLJ45612 fis, clone BRTHA3025073, highly similar to Actin cross-linking family protein 7 Y259D mutant monomer, 46 kDa Homo sapiens protein
|
| Buffer: |
PBS, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2014 Nov 7
|
In vivo epidermal migration requires focal adhesion targeting of ACF7.
Nat Commun 7:11692 (2016)
Yue J, Zhang Y, Liang WG, Gou X, Lee P, Liu H, Lyu W, Tang WJ, Chen SY, Yang F, Liang H, Wu X
|
| RgGuinier |
3.4 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
55 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Primase D5 protein fragment containing the D5N and helicase domain hexamer, 321 kDa Vaccinia virus protein
|
| Buffer: |
20 mM Tris 150 mM NaCL 5% glycerol 1mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Sep 26
|
Domain Organization of Vaccinia Virus Helicase-Primase D5.
J Virol 90(9):4604-4613 (2016)
Hutin S, Ling WL, Round A, Effantin G, Reich S, Iseni F, Tarbouriech N, Schoehn G, Burmeister WP
|
| RgGuinier |
4.8 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
570 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
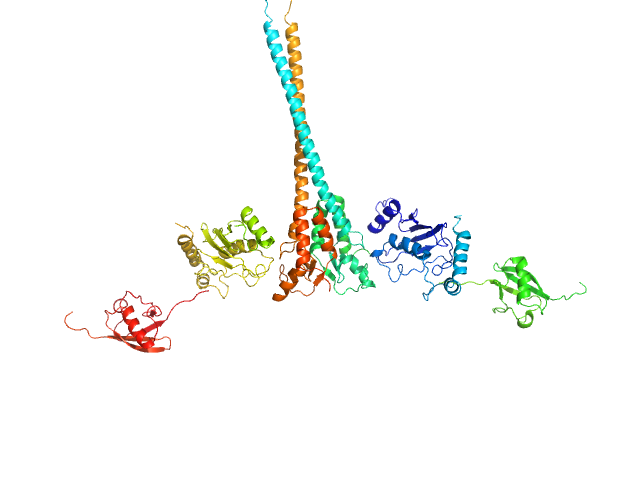
E3 ubiquitin-protein ligase RNF8 dimer, 35 kDa Homo sapiens protein
Ubiquitin-conjugating enzyme E2 N double mutant (C87K, K92A) dimer, 36 kDa Homo sapiens protein
Polyubiquitin-C dimer, 17 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES 200 mM NaCl 0.01 mM ZnSO4 1 mM DTT, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Sep 8
|
RNF8 E3 Ubiquitin Ligase Stimulates Ubc13 E2 Conjugating Activity That Is Essential for DNA Double Strand Break Signaling and BRCA1 Tumor Suppressor Recruitment.
J Biol Chem 291(18):9396-410 (2016)
Hodge CD, Ismail IH, Edwards RA, Hura GL, Xiao AT, Tainer JA, Hendzel MJ, Glover JN
|
| RgGuinier |
4.5 |
nm |
| Dmax |
18.9 |
nm |
| VolumePorod |
119 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
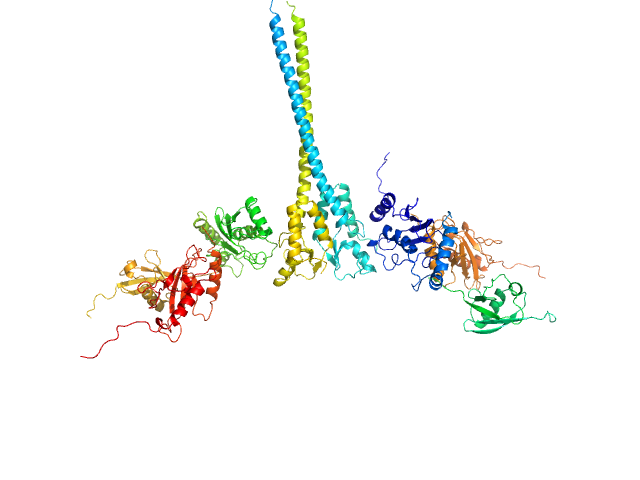
E3 ubiquitin-protein ligase RNF8 dimer, 35 kDa Homo sapiens protein
Ubiquitin-conjugating enzyme E2 N double mutant (C87K, K92A) dimer, 36 kDa Homo sapiens protein
Polyubiquitin-C dimer, 17 kDa Homo sapiens protein
Ubiquitin-conjugating enzyme E2 variant 2 dimer, 34 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES 200 mM NaCl 0.01 mM ZnSO4 1 mM DTT, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Sep 8
|
RNF8 E3 Ubiquitin Ligase Stimulates Ubc13 E2 Conjugating Activity That Is Essential for DNA Double Strand Break Signaling and BRCA1 Tumor Suppressor Recruitment.
J Biol Chem 291(18):9396-410 (2016)
Hodge CD, Ismail IH, Edwards RA, Hura GL, Xiao AT, Tainer JA, Hendzel MJ, Glover JN
|
| RgGuinier |
5.4 |
nm |
| Dmax |
23.8 |
nm |
| VolumePorod |
214 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
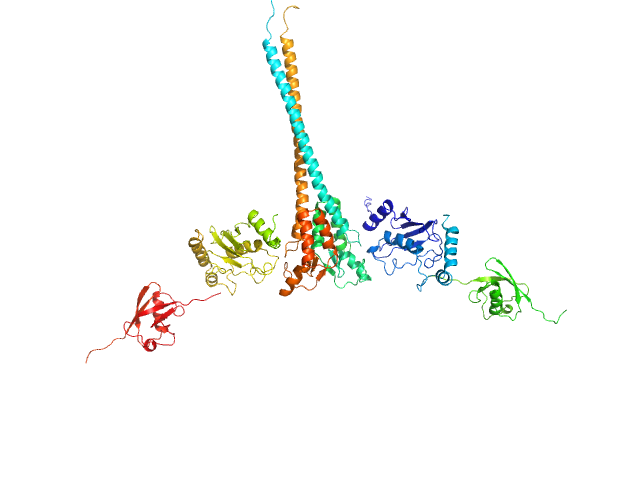
Ubiquitin-conjugating enzyme E2 N double mutant (C87K, K92A) dimer, 36 kDa Homo sapiens protein
Polyubiquitin-C dimer, 17 kDa Homo sapiens protein
E3 ubiquitin-protein ligase RNF8 mutant (L451D) dimer, 35 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES 200 mM NaCl 0.01 mM ZnSO4 1 mM DTT, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Sep 8
|
RNF8 E3 Ubiquitin Ligase Stimulates Ubc13 E2 Conjugating Activity That Is Essential for DNA Double Strand Break Signaling and BRCA1 Tumor Suppressor Recruitment.
J Biol Chem 291(18):9396-410 (2016)
Hodge CD, Ismail IH, Edwards RA, Hura GL, Xiao AT, Tainer JA, Hendzel MJ, Glover JN
|
| RgGuinier |
4.3 |
nm |
| Dmax |
18.9 |
nm |
| VolumePorod |
111 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
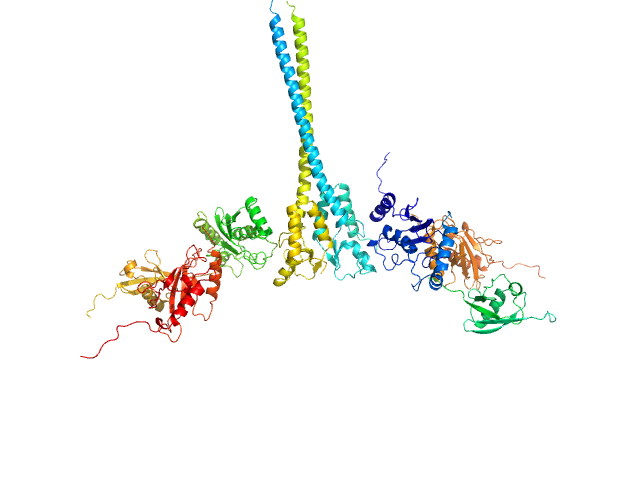
Ubiquitin-conjugating enzyme E2 N double mutant (C87K, K92A) dimer, 36 kDa Homo sapiens protein
Polyubiquitin-C dimer, 17 kDa Homo sapiens protein
Ubiquitin-conjugating enzyme E2 variant 2 dimer, 34 kDa Homo sapiens protein
E3 ubiquitin-protein ligase RNF8 mutant (L451D) dimer, 35 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES 200 mM NaCl 0.01 mM ZnSO4 1 mM DTT, pH: 6.8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2015 Sep 8
|
RNF8 E3 Ubiquitin Ligase Stimulates Ubc13 E2 Conjugating Activity That Is Essential for DNA Double Strand Break Signaling and BRCA1 Tumor Suppressor Recruitment.
J Biol Chem 291(18):9396-410 (2016)
Hodge CD, Ismail IH, Edwards RA, Hura GL, Xiao AT, Tainer JA, Hendzel MJ, Glover JN
|
| RgGuinier |
5.2 |
nm |
| Dmax |
23.8 |
nm |
| VolumePorod |
192 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
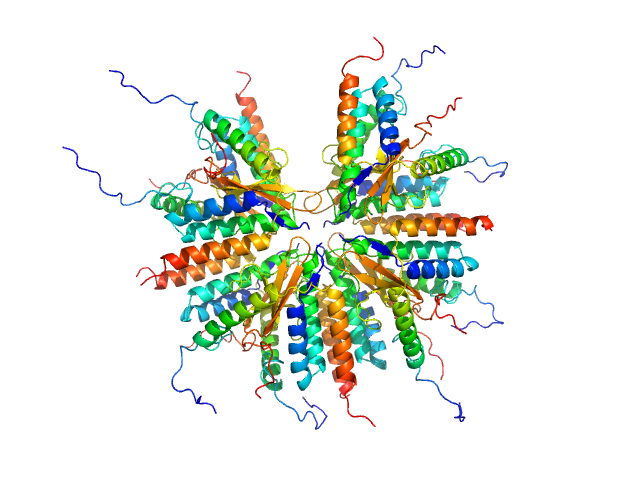
Plasmid stabilization protein ParE 16-mer, 188 kDa Escherichia coli protein
Uncharacterized protein (Antitoxin) 16-mer, 135 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Dec 9
|
A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
J Mol Biol 428(8):1589-603 (2016)
Sterckx YG, Jové T, Shkumatov AV, Garcia-Pino A, Geerts L, De Kerpel M, Lah J, De Greve H, Van Melderen L, Loris R
|
| RgGuinier |
3.8 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
312 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
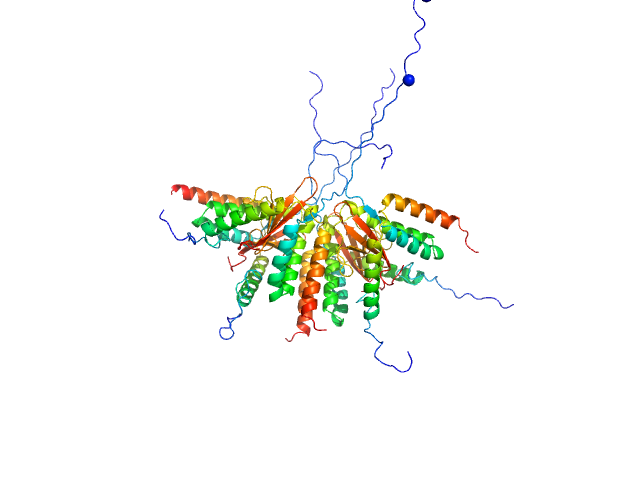
Plasmid stabilization protein ParE octamer, 102 kDa Escherichia coli protein
Uncharacterized protein (Antitoxin) octamer, 68 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris-HCl 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 Feb 5
|
A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
J Mol Biol 428(8):1589-603 (2016)
Sterckx YG, Jové T, Shkumatov AV, Garcia-Pino A, Geerts L, De Kerpel M, Lah J, De Greve H, Van Melderen L, Loris R
|
| RgGuinier |
3.3 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
166 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Plasmid stabilization protein ParE monomer, 13 kDa Escherichia coli protein
Uncharacterized protein monomer, 6 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
50 mM Tris-HCl, 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 Feb 5
|
A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
J Mol Biol 428(8):1589-603 (2016)
Sterckx YG, Jové T, Shkumatov AV, Garcia-Pino A, Geerts L, De Kerpel M, Lah J, De Greve H, Van Melderen L, Loris R
|
| RgGuinier |
2.2 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
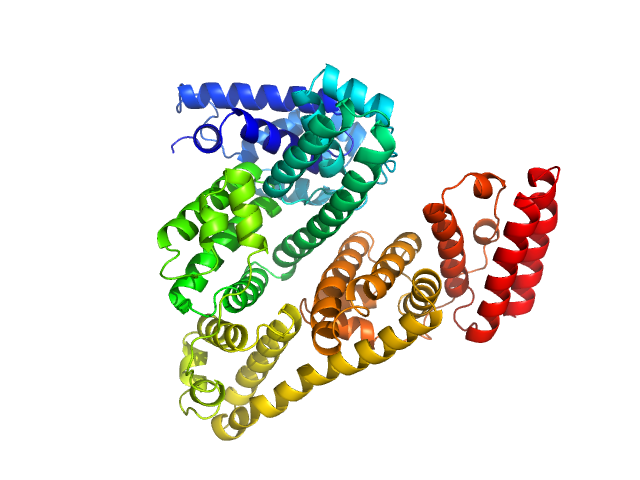
Bovine serum albumin, monomer monomer, 66 kDa Bos taurus protein
|
| Buffer: |
20 mM Tris 142 mM NaCl 5 % Glycerol 1mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 22
|
Bovine Serum Albumin measured by SEC-SAXS
Martha Brennich
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
118 |
nm3 |
|
|