|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDML5_electron_density.png)
|
| Sample: |
Proteoliposomes composed of lipids from A/Puerto Rico/8/34 (H1N1) virus envelope together with the HA LI45 peptides loaded with M1 protein (lipid:M1 molar ration 4:1) monomer, 2000 kDa
|
| Buffer: |
100 mM NaCl, 50 mM MES buffer, pH: 6.8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Nov 26
|
The Cytoplasmic Tail of Influenza A Virus Hemagglutinin and Membrane Lipid Composition Change the Mode of M1 Protein Association with the Lipid Bilayer
Membranes 11(10):772 (2021)
Kordyukova L, Konarev P, Fedorova N, Shtykova E, Ksenofontov A, Loshkarev N, Dadinova L, Timofeeva T, Abramchuk S, Moiseenko A, Baratova L, Svergun D, Batishchev O
|
|
|
|
|
|
|
|
| Sample: |
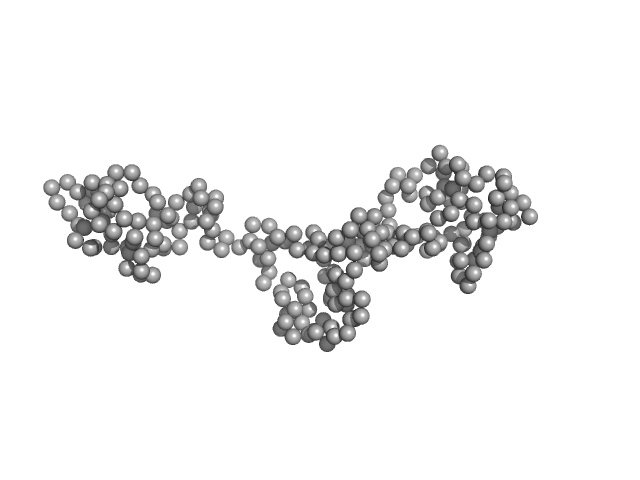
Coronin tetramer, 21 kDa Trypanosoma brucei brucei … protein
|
| Buffer: |
100 mM Tris-HCl, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2019 Oct 17
|
Structural insights into kinetoplastid coronin oligomerization domain and F-actin interaction
Current Research in Structural Biology (2021)
Parihar P, Singh A, Karade S, Sahasrabuddhe A, Pratap J
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
|
|
|
|
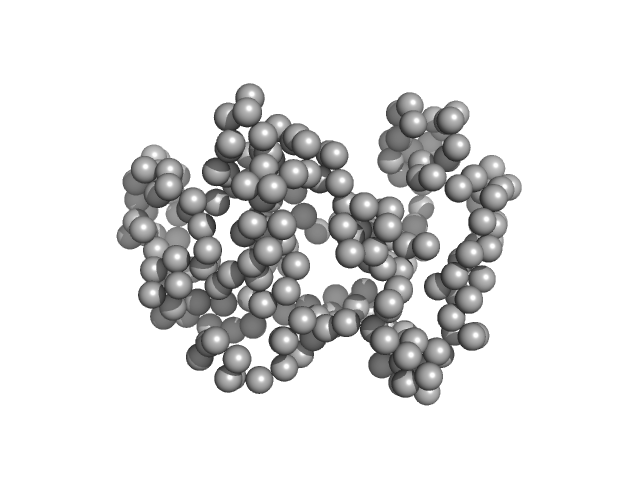
|
| Sample: |
Cell cycle associated protein MOB1, putative monomer, 27 kDa Leishmania donovani (strain … protein
|
| Buffer: |
100 mM Tris-HCl, 100 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2017 Jun 22
|
Structural insights into kinetoplastid coronin oligomerization domain and F-actin interaction
Current Research in Structural Biology (2021)
Parihar P, Singh A, Karade S, Sahasrabuddhe A, Pratap J
|
| RgGuinier |
2.1 |
nm |
| Dmax |
5.6 |
nm |
| VolumePorod |
59 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclic di-AMP synthase CdaA dimer, 44 kDa Bacillus subtilis (strain … protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 21
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
60 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Phosphoglucosamine mutase dimer, 101 kDa Bacillus subtilis (strain … protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 21
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.2 |
nm |
| VolumePorod |
140 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Cyclic di-AMP synthase CdaA dimer, 44 kDa Bacillus subtilis (strain … protein
Phosphoglucosamine mutase dimer, 101 kDa Bacillus subtilis (strain … protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 21
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
4.5 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
250 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
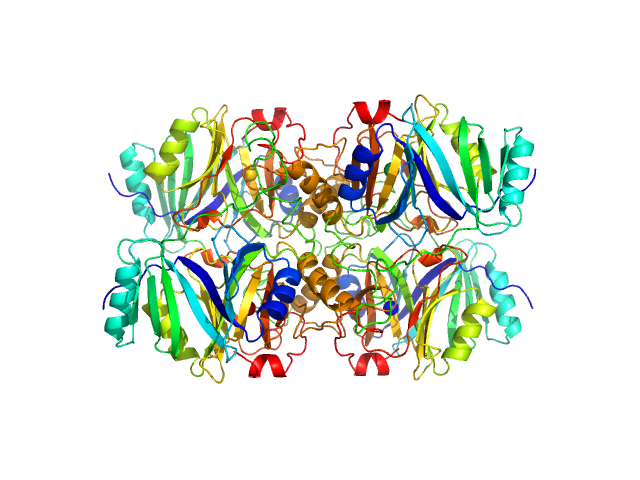
Metapyrocatechase tetramer, 139 kDa Novosphingobium sp. AAP93 protein
|
| Buffer: |
20 mM Tris-HCl 250 mM NaCl, pH 8.0, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2019 Dec 20
|
Shape-function of a novel metapyrocatechase, RW4-MPC: Metagenomics to SAXS data based insight into deciphering regulators of function.
Int J Biol Macromol 188:1012-1024 (2021)
Vasudeva G, Sidhu C, Kalidas N, Ashish, Pinnaka AK
|
| RgGuinier |
3.7 |
nm |
| Dmax |
10.2 |
nm |
|
|
|
|
|
|
|
| Sample: |
Cyclic di-AMP synthase CdaA dimer, 44 kDa Bacillus subtilis (strain … protein
Phosphoglucosamine mutase dimer, 83 kDa Bacillus subtilis protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Aug 2
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.1 |
nm |
| VolumePorod |
156 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
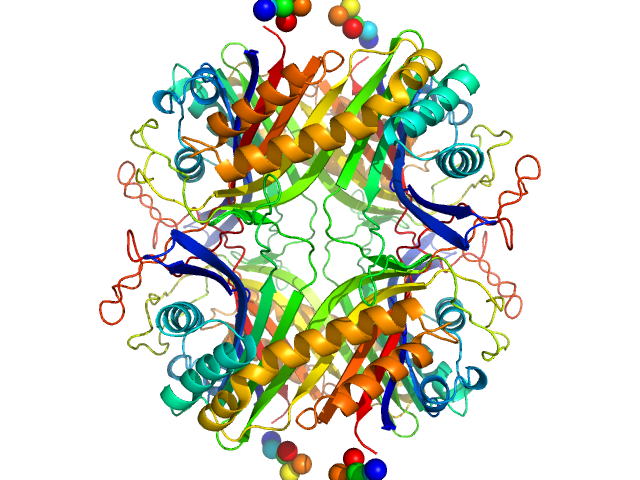
Urate Oxidase (Uricase) from Aspergillus flavus tetramer, 137 kDa Aspergillus flavus protein
|
| Buffer: |
20 mM Tris. 150 mM NaCl, 1 mM EDTA, 5 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2019 Jul 1
|
Urate Oxidase (Uricase) from Aspergillus flavus
Tsutomu Matsui
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
225 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Iron-sulfur cluster assembly protein trimer, 42 kDa Plasmodium falciparum (isolate … protein
|
| Buffer: |
50 mM Tris-Cl, 300 mM NaCl, 5% Glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2019 May 5
|
[Fe-S] cluster biogenesis and unusual assembly of the ISC scaffold complex in Plasmodium falciparum mitochondrion
Ravishankar Ramachandran
|
| RgGuinier |
5.4 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
177 |
nm3 |
|
|