|
|
|
|
|
| Sample: |
C-type lectin mosGCTL-1 dimer, 35 kDa Aedes aegypti protein
|
| Buffer: |
25 mM Citrate, 1 M NaCl, 10 mM CaCl2, pH: 4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Aug 7
|
Conserved dimerization architecture in C-type lectins from virus-vector mosquitoes
Max Renner
|
|
|
|
|
|
|
|
| Sample: |
C-type lectin mosGCTL-1 dimer, 35 kDa Aedes aegypti protein
|
| Buffer: |
25 mM Citrate, 1 M NaCl, 10 mM CaCl2, pH: 4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Aug 7
|
Conserved dimerization architecture in C-type lectins from virus-vector mosquitoes
Max Renner
|
|
|
|
|
|
|
|
| Sample: |
MosGCTL-3 dimer, 36 kDa Aedes aegypti protein
|
| Buffer: |
20 mM Tris, 250 mM NaCl, 2.5 mM CaCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Dec 17
|
Conserved dimerization architecture in C-type lectins from virus-vector mosquitoes
Max Renner
|
|
|
|
|
|
|
|
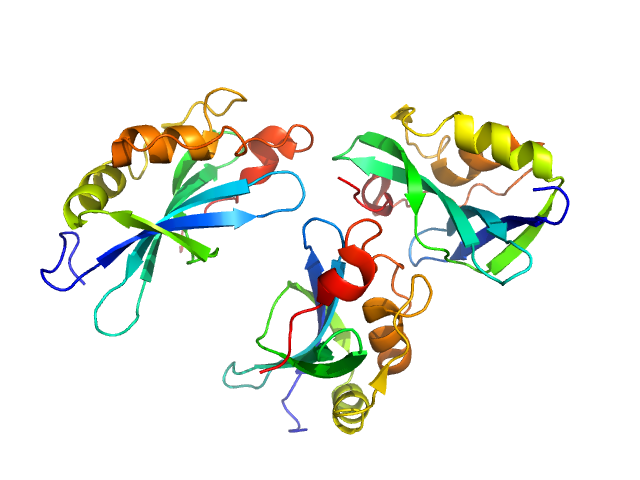
| Sample: |
Genome polyprotein monomer, 15 kDa Aichi virus (strain … protein
|
| Buffer: |
20 mM HEPES,150 mM NaCl, 2 mM TCEP, 1% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 11
|
Structural and functional repurposing of picornaviral H-box 2A proteins
Marion Pichon
|
| RgGuinier |
1.8 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
|
|
|
|
|
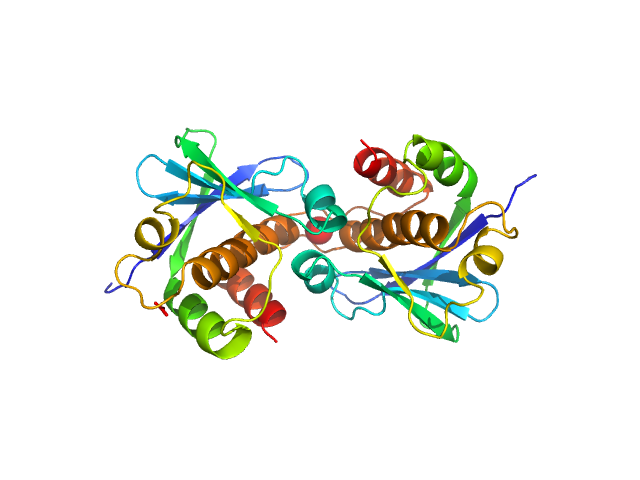
| Sample: |
Genome polyprotein (I918M) dimer, 34 kDa Human parechovirus 1 … protein
|
| Buffer: |
20 mM HEPES,150 mM NaCl, 2 mM TCEP, 1% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 11
|
Structural and functional repurposing of picornaviral H-box 2A proteins
Marion Pichon
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
58 |
nm3 |
|
|
|
|
|
|
|
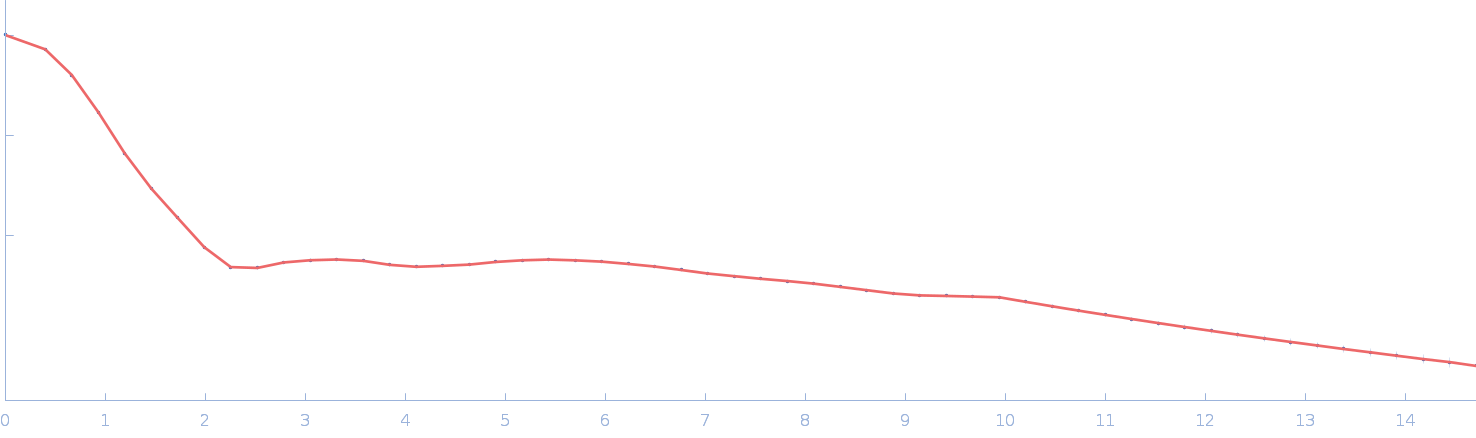
| Sample: |
Chromochloris zofingiensis Hexokinase-1 monomer, 55 kDa Chromochloris zofingiensis protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 10 mM MgCl2, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2023 Mar 25
|
Chromochloris zofingiensis Hexokinase-1 (HXK-1)
Estella Yee
|
| RgGuinier |
2.6 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
|
|
|
|
|
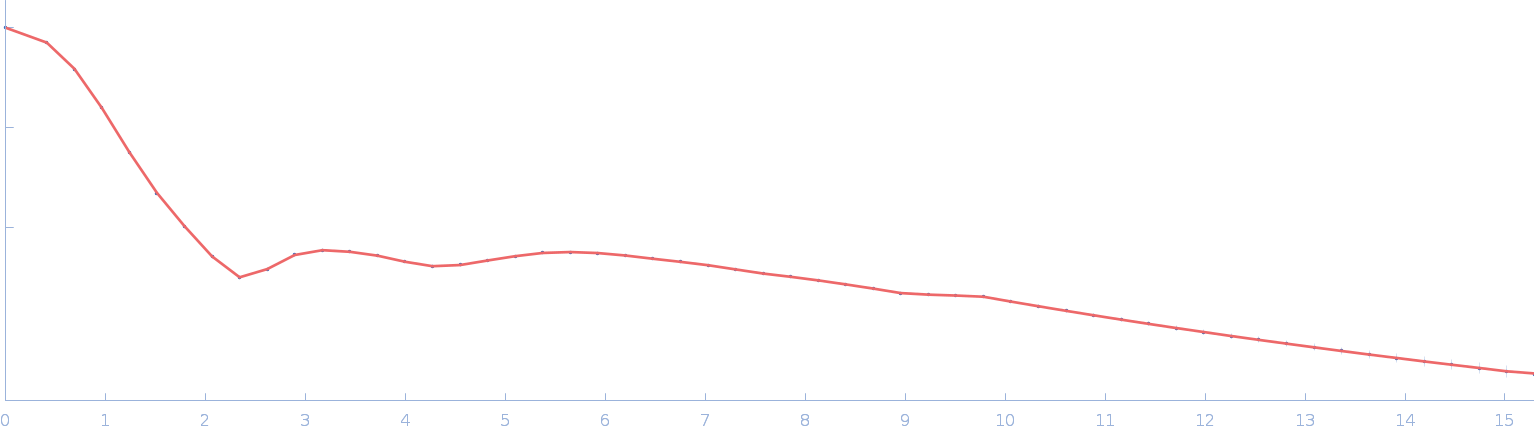
| Sample: |
Chromochloris zofingiensis Hexokinase-1 monomer, 55 kDa Chromochloris zofingiensis protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 10 mM MgCl2, 1 mM TCEP, 200 mM glucose, pH: 8 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2023 Mar 22
|
Chromochloris zofingiensis Hexokinase-1 (HXK-1)
Estella Yee
|
| RgGuinier |
2.5 |
nm |
| Dmax |
7.6 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
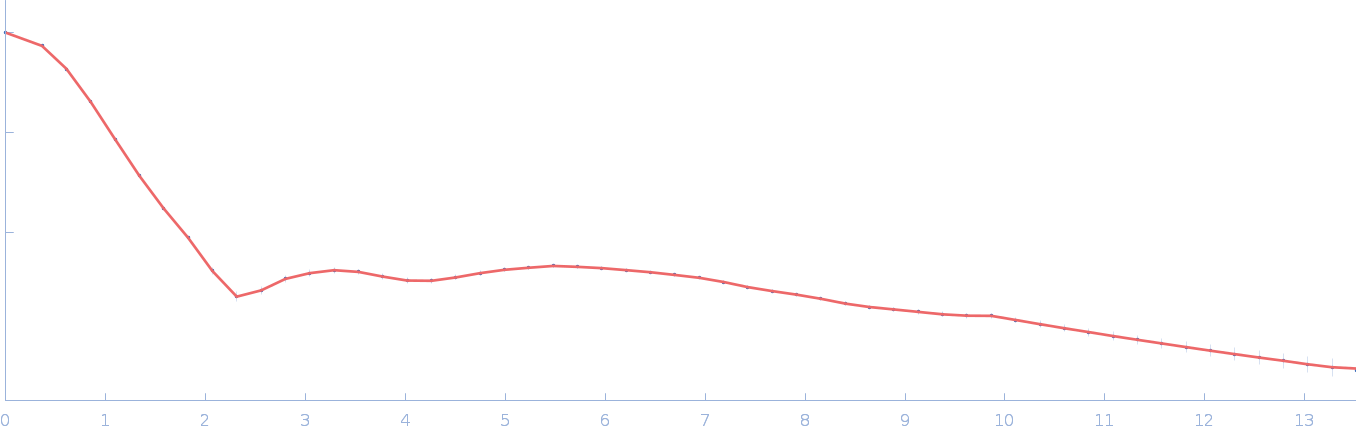
Chromochloris zofingiensis Hexokinase-1 (E291W) monomer, 55 kDa Chromochloris zofingiensis protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 10 mM MgCl2, 1 mM TCEP, 200 mM glucose, pH: 8 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2023 Mar 22
|
Chromochloris zofingiensis Hexokinase-1 (HXK-1)
Estella Yee
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
87 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
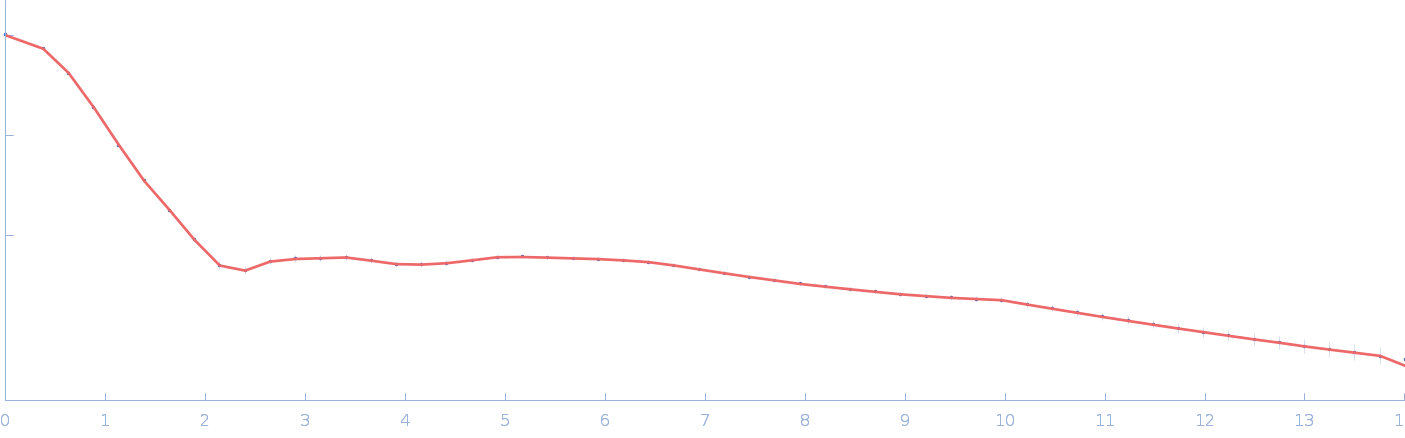
Chromochloris zofingiensis Hexokinase-1 (E291W) monomer, 55 kDa Chromochloris zofingiensis protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 10 mM MgCl2, 1 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2023 Feb 10
|
Chromochloris zofingiensis Hexokinase-1 (HXK-1)
Estella Yee
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
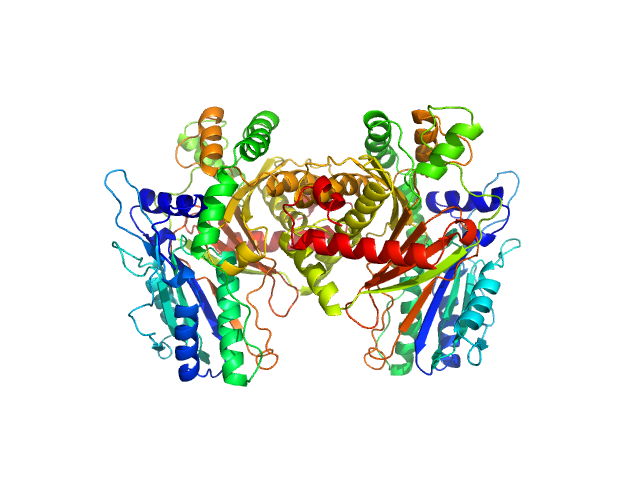
Condensation Domain from Coprococcus Eutactus dimer, 108 kDa Coprococcus eutactus ATCC … protein
|
| Buffer: |
10mM HEPES pH 7.5, 50mM NaCl, 0.2mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2022 Dec 16
|
Structure of a Stand-Alone Homodimeric NRPS Condensation Domain Reveals Occlusion of the Canonical Carrier-Protein Interface
(2026)
Singh J, Grant T, Gulick A
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
184 |
nm3 |
|
|