|
|
|
|

|
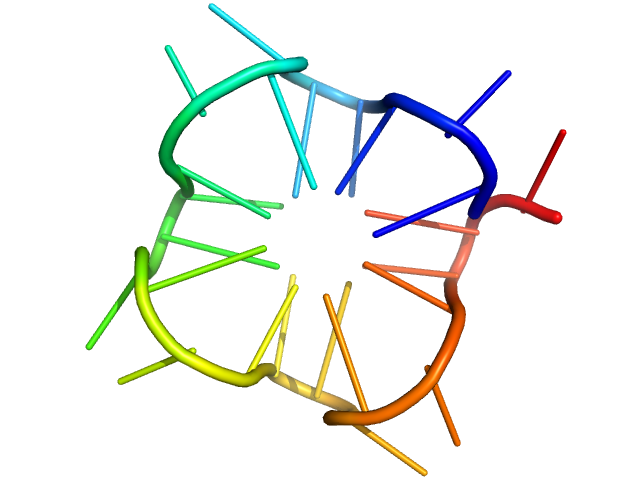
| Sample: |
PolyGU RNA - (GU)12 monomer, 8 kDa synthetic RNA RNA
|
| Buffer: |
20 mM HEPES, 150 mM KCl, pH: 7 |
| Experiment: |
SAXS
data collected at 12-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Jun 10
|
Solution structure of poly(UG) RNA
Journal of Molecular Biology :168340 (2023)
Escobar C, Petersen R, Tonelli M, Fan L, Henzler-Wildman K, Butcher S
|
| RgGuinier |
1.3 |
nm |
| Dmax |
4.7 |
nm |
| VolumePorod |
8 |
nm3 |
|
|
|
|
|
|

|
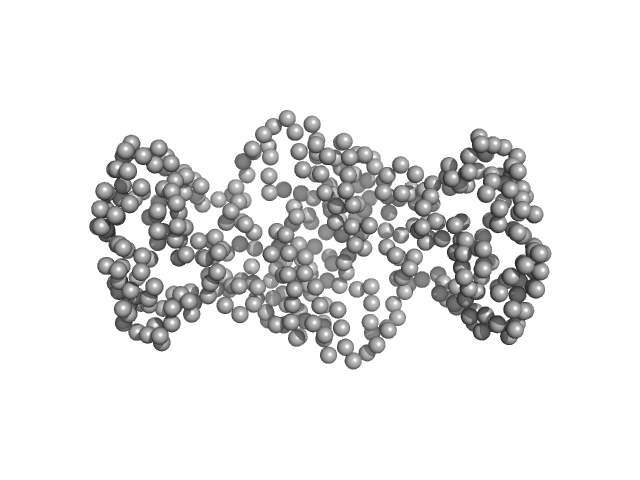
| Sample: |
20 kDa accessory protein dimer, 42 kDa Bacillus thuringiensis serovar … protein
|
| Buffer: |
10 mM Tris, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BL-18, INDUS-2 on 2023 Aug 15
|
20-kDa accessory protein (P20) from Bacillus thuringiensis subsp. israelensis ISPC-12: Purification, characterization, solution scattering and structural analysis
International Journal of Biological Macromolecules :127985 (2023)
Kinkar O, Singh R, Prashar A, Kumar A, Hire R, Makde R
|
| RgGuinier |
3.0 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
67 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris/Tris-HCl, 0.1 mM CaCl2, 1 mM NaN3, 0.2 mM ATP, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Jul 1
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
2.9 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
58 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris/Tris-HCl, 0.1 mM CaCl2, 1 mM NaN3, 1.0 mM ATP, 50 mM KCl, 2 mM MgCl2, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Jul 1
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
15.7 |
nm |
| Dmax |
60.0 |
nm |
| VolumePorod |
4710 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDTZ4_fit1_model1.png)
|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris/Tris-HCl, 2.0 mM EDTA, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Nov 16
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
4.9 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
549 |
nm3 |
|
|
|
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDT25_fit1_model1.png)
|
| Sample: |
Actin, alpha skeletal muscle monomer, 42 kDa Oryctolagus cuniculus protein
|
| Buffer: |
5 mM Tris, 0.1 mM CaCl2, 1 mM NaN3, 0.2 mM ATP, pH: 8.1 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2021 Jul 5
|
Small-angle X-ray scattering structural insights into alternative pathway of actin oligomerization associated with inactivated state
Biochemical and Biophysical Research Communications :149340 (2023)
Ryzhykau Y, Povarova O, Dronova E, Kuklina D, Antifeeva I, Ilyinsky N, Okhrimenko I, Semenov Y, Kuklin A, Ivanovich V, Fonin A, Uversky V, Turoverov K, Kuznetsova I
|
| RgGuinier |
3.1 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Heat-labile enterotoxin B chain monomer, 36 kDa Clostridium perfringens protein
|
| Buffer: |
10 mM HEPES, 100 mM NaCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Nov 18
|
Structural Basis of Clostridium perfringens Enterotoxin Activation and Oligomerization by Trypsin
Toxins 15(11):637 (2023)
Ogbu C, Kapoor S, Vecchio A
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.2 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
|
|
|
|

|
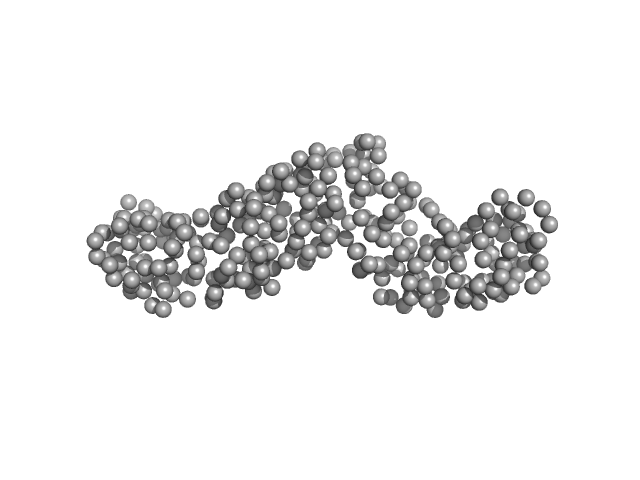
| Sample: |
Heat-labile enterotoxin B chain monomer, 33 kDa Clostridium perfringens protein
|
| Buffer: |
10 mM HEPES, 100 mM NaCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Nov 18
|
Structural Basis of Clostridium perfringens Enterotoxin Activation and Oligomerization by Trypsin
Toxins 15(11):637 (2023)
Ogbu C, Kapoor S, Vecchio A
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Ferritin light chain 24-mer, 479 kDa Equus caballus protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SANS
data collected at SPB/SFX, European XFEL on 2021 May 14
|
Form factor determination of biological molecules with X-ray free electron laser small-angle scattering (XFEL-SAS).
Commun Biol 6(1):1057 (2023)
Blanchet CE, Round A, Mertens HDT, Ayyer K, Graewert M, Awel S, Franke D, Dörner K, Bajt S, Bean R, Custódio TF, de Wijn R, Juncheng E, Henkel A, Gruzinov A, Jeffries CM, Kim Y, Kirkwood H, Kloos M, Knoška J, Koliyadu J, Letrun R, Löw C, Makroczyova J, Mall A, Meijers R, Pena Murillo GE, Oberthür D, Round E, Seuring C, Sikorski M, Vagovic P, Valerio J, Wollweber T, Zhuang Y, Schulz J, Haas H, Chapman HN, Mancuso AP, Svergun D
|
|
|
|
|
|
|
|
| Sample: |
Spike glycoprotein (ACE2 receptor binding domain) monomer, 29 kDa Severe acute respiratory … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SANS
data collected at SPB/SFX, European XFEL on 2021 May 14
|
Form factor determination of biological molecules with X-ray free electron laser small-angle scattering (XFEL-SAS).
Commun Biol 6(1):1057 (2023)
Blanchet CE, Round A, Mertens HDT, Ayyer K, Graewert M, Awel S, Franke D, Dörner K, Bajt S, Bean R, Custódio TF, de Wijn R, Juncheng E, Henkel A, Gruzinov A, Jeffries CM, Kim Y, Kirkwood H, Kloos M, Knoška J, Koliyadu J, Letrun R, Löw C, Makroczyova J, Mall A, Meijers R, Pena Murillo GE, Oberthür D, Round E, Seuring C, Sikorski M, Vagovic P, Valerio J, Wollweber T, Zhuang Y, Schulz J, Haas H, Chapman HN, Mancuso AP, Svergun D
|
|
|