|
|
|
|
|
| Sample: |
Alpha cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
Alpha cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
Beta Cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
Beta Cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
Gamma Cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
Gamma Cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
Gamma Cyclodextrin monomer, 1 kDa
|
| Buffer: |
water, pH: 7 |
| Experiment: |
SAXS
data collected at 11-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Nov 2
|
Extended q‐range X‐ray Scattering Reveals High‐Resolution Structural Details of Biomacromolecules in Aqueous Solutions
Chemistry – A European Journal (2023)
Chen J, Borkiewicz O, Grishaev A, Zhang F, Bera M, Ruett U, Levin I
|
|
|
|
|
|
|
|
| Sample: |
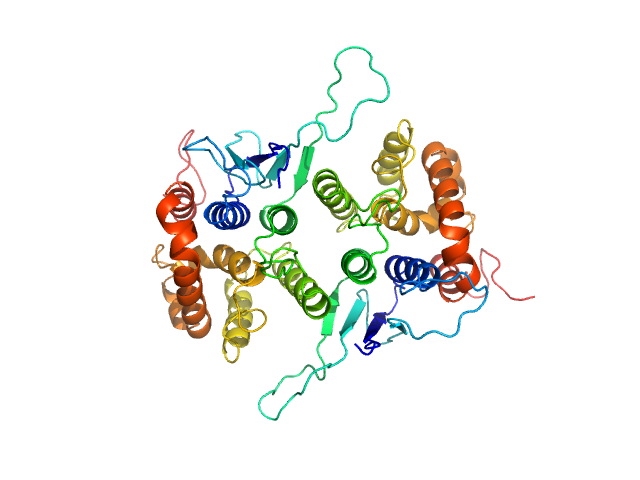
Glutamate--tRNA ligase dimer, 57 kDa Plasmodium berghei (strain … protein
|
| Buffer: |
25 mM HEPES-NaOH, 300 mM NaCl, 5% (v/v) glycerol, 0.005% (m/v) n-dodecyl beta-D-maltoside, 5 mM 2-mercaptoethanol, pH: 7 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2021 Oct 9
|
Solution x-ray scattering highlights discrepancies in Plasmodium multi-aminoacyl-tRNA synthetase complexes.
Protein Sci :e4564 (2023)
Jaramillo Ponce JR, Théobald-Dietrich A, Bénas P, Paulus C, Sauter C, Frugier M
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
96 |
nm3 |
|
|
|
|
|
|
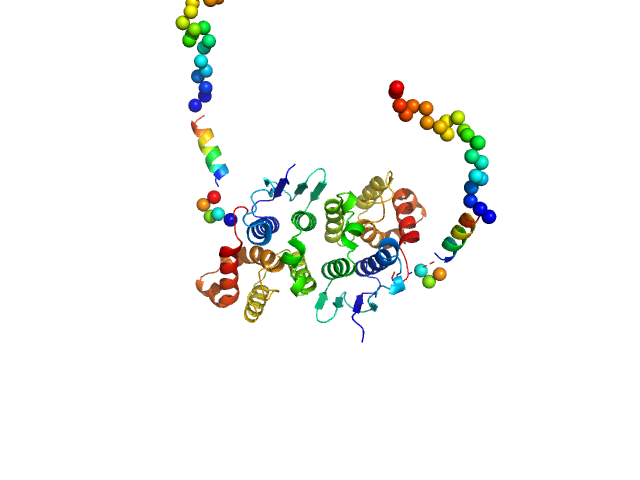
|
| Sample: |
TRNA import protein tRIP dimer, 49 kDa Plasmodium berghei (strain … protein
|
| Buffer: |
25 mM HEPES-NaOH, 300 mM NaCl, 5% (v/v) glycerol, 0.005% (m/v) n-dodecyl beta-D-maltoside, 5 mM 2-mercaptoethanol, pH: 7 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2021 Oct 9
|
Solution x-ray scattering highlights discrepancies in Plasmodium multi-aminoacyl-tRNA synthetase complexes.
Protein Sci :e4564 (2023)
Jaramillo Ponce JR, Théobald-Dietrich A, Bénas P, Paulus C, Sauter C, Frugier M
|
| RgGuinier |
2.9 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
68 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
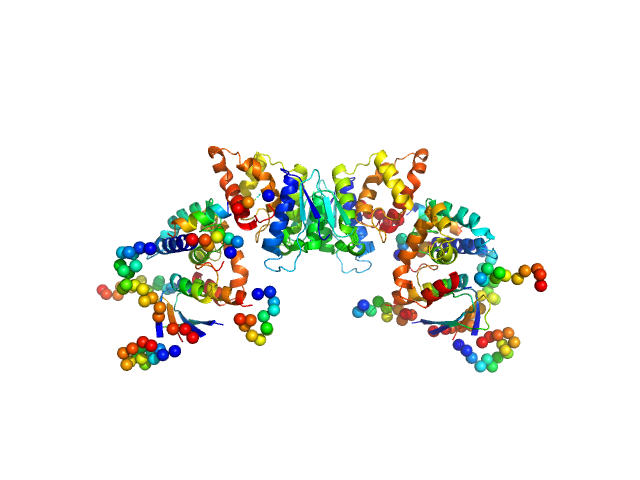
Chimeric construct containing the GST-like domains of P. berghei tRNA import protein and glutamyl-tRNA synthetase dimer, 105 kDa Plasmodium berghei ANKA protein
|
| Buffer: |
25 mM HEPES-NaOH, 300 mM NaCl, 5% (v/v) glycerol, 0.005% (m/v) n-dodecyl beta-D-maltoside, 5 mM 2-mercaptoethanol, pH: 7 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2021 Oct 9
|
Solution x-ray scattering highlights discrepancies in Plasmodium multi-aminoacyl-tRNA synthetase complexes.
Protein Sci :e4564 (2023)
Jaramillo Ponce JR, Théobald-Dietrich A, Bénas P, Paulus C, Sauter C, Frugier M
|
| RgGuinier |
3.9 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
145 |
nm3 |
|
|