|
|
|
|
|
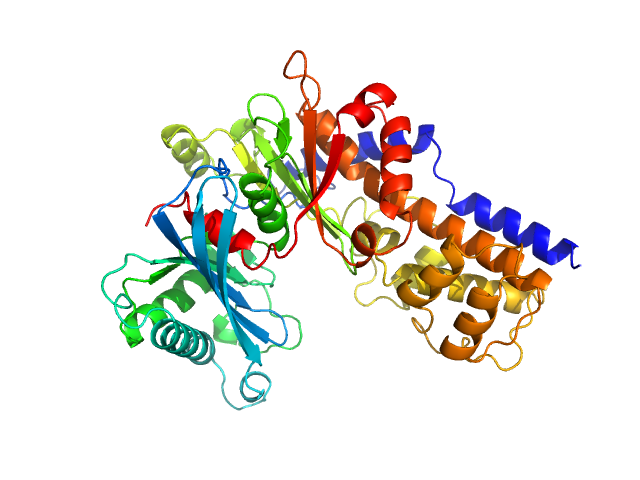
| Sample: |
Glucokinase-1 (I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, 10 mM D-mannose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|
|
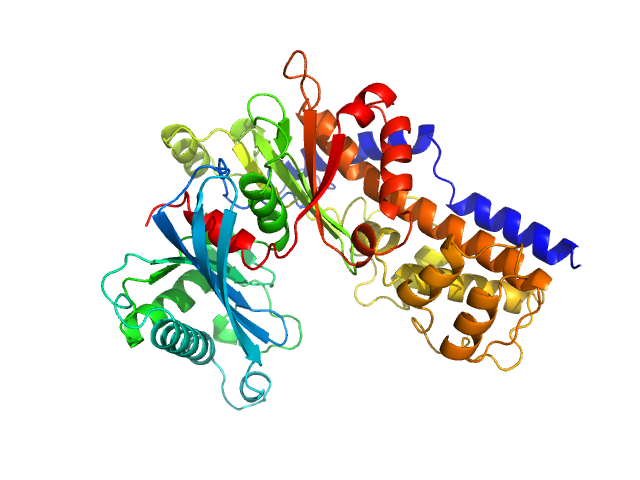
| Sample: |
Glucokinase-1 (I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris/HCl, 1 mM DTT, 10 mM D-glucose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.4 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|
|
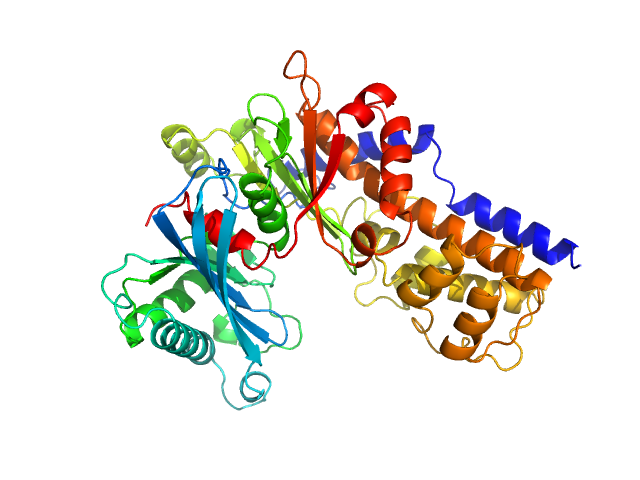
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
|
|
|
|
|
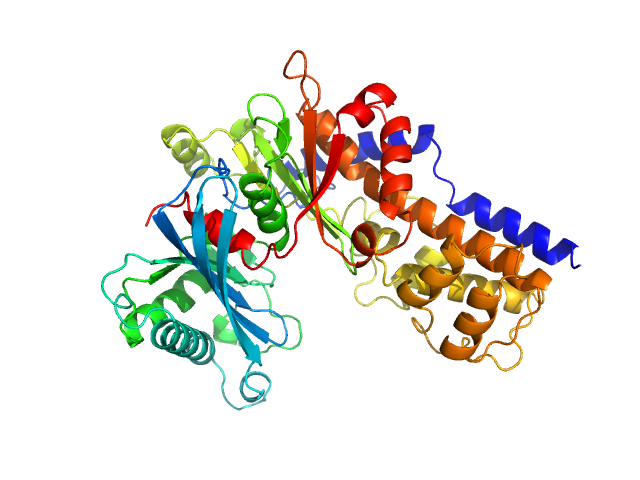
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
|
|
|
|
|
|
|
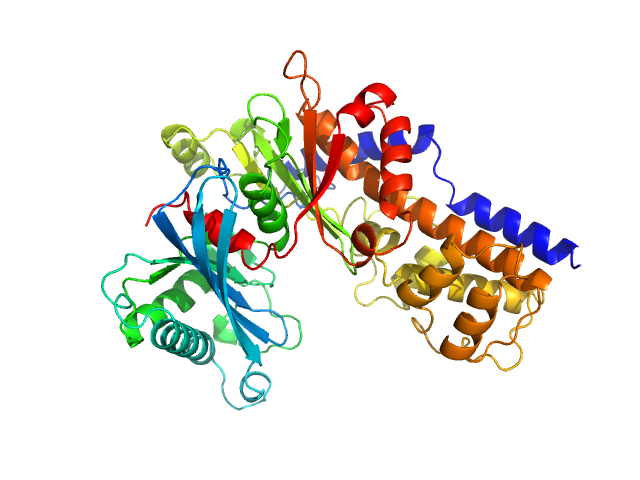
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
|
|
|
|
|
|
|
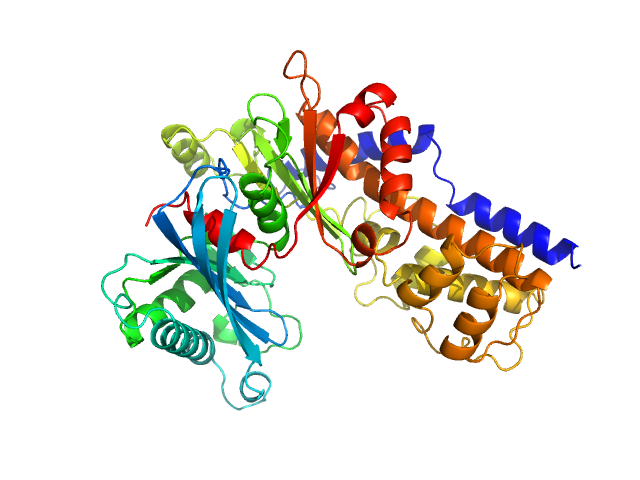
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, 10 mM D-fructose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
|
|
|
|
|
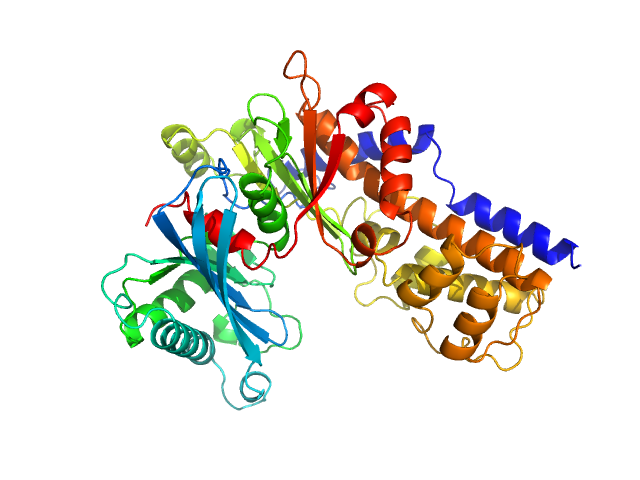
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris-HCl, 1 mM DTT, 10 mM D-mannose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|
|
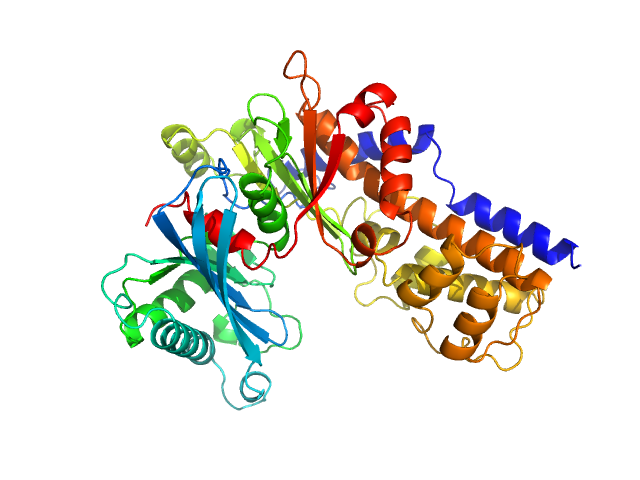
| Sample: |
Glucokinase-1 (H304Q, I356D, Y419D, H420D) monomer, 54 kDa Kluyveromyces lactis (strain … protein
|
| Buffer: |
10 mM Tris/HCl, 1 mM DTT, 10 mM D-glucose, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 14
|
Sugar binding to Kluyveromyces lactis glucokinase 1 (KlGlk1)
Renato H. Weiße
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.6 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|
|
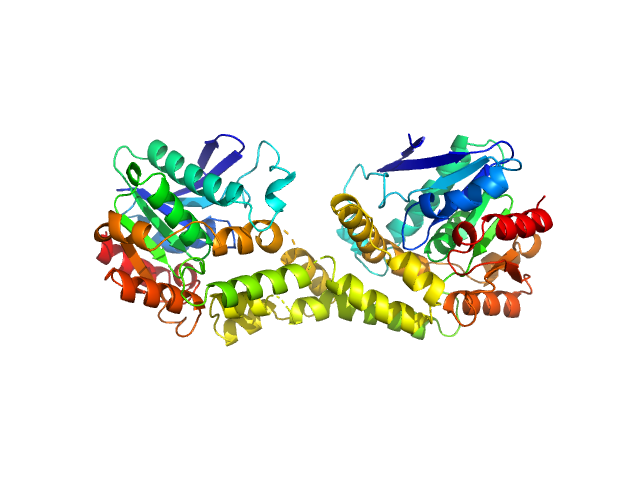
| Sample: |
Uncharacterized hydrolase SAUSA300_2518 dimer, 62 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
10 mM HEPES, 50 mM NaCl, 0.1% w/v NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2025 Mar 26
|
Unique structural and ligand-binding properties of the Staphylococcus aureus
serine hydrolase FphE
Proceedings of the National Academy of Sciences 123(13) (2026)
Jo J, Upadhyay T, You X, Bennett J, Lee H, Bogyo M, Fellner M
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
|
|
|
|
|
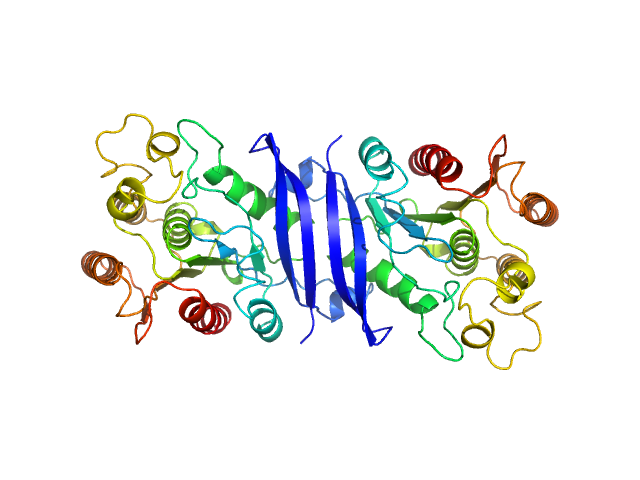
| Sample: |
S-formylglutathione hydrolase dimer, 59 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
10 mM HEPES, 50 mM NaCl, 0.1% w/v NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2025 Mar 26
|
Unique structural and ligand-binding properties of the Staphylococcus aureus
serine hydrolase FphE
Proceedings of the National Academy of Sciences 123(13) (2026)
Jo J, Upadhyay T, You X, Bennett J, Lee H, Bogyo M, Fellner M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
99 |
nm3 |
|
|