|
|
|
|
|
| Sample: |
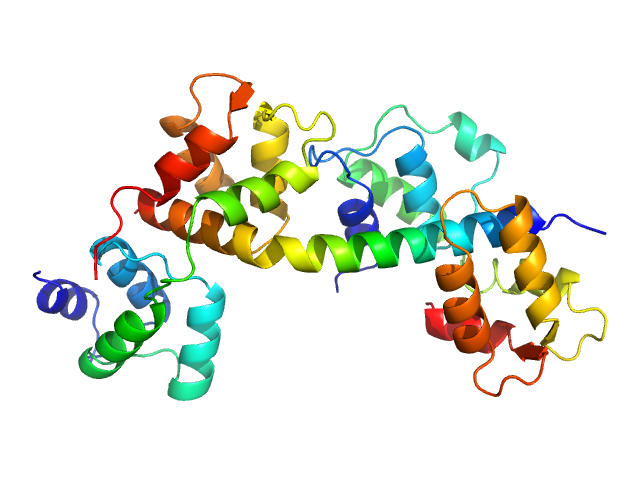
...asma gondii essential light chain 1 monomer, 15 kDa ...asma gondii protein
Myosin A monomer, 5 kDa ...asma gondii protein
...asma gondii myosin light chain, full length monomer, 24 kDa ...asma gondii protein
|
| Buffer: |
...aCl, 0.5 mM TCEP, pH: 7.5
|
| Experiment: |
...AXS
data collected at EMBL P12, ...A III on 2019 Sep 30
|
...al role of essential light chains in the apicomplexan glideosome.
Commun Biol 3(1):568 (2020)
...azicky S, Dhamotharan K, Kaszuba K, Mertens HDT, Gilberger T, Svergun D, Kosinski J, Weininger U, Löw C
|
| RgGuinier |
3.2 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
69 |
nm3 |
|