UniProt ID: Q9Y6M1 (1-599) Insulin-like growth factor 2 mRNA-binding protein 2
UniProt ID: None (None-None) Cox7B RNA
|
|
|
|
| Sample: |
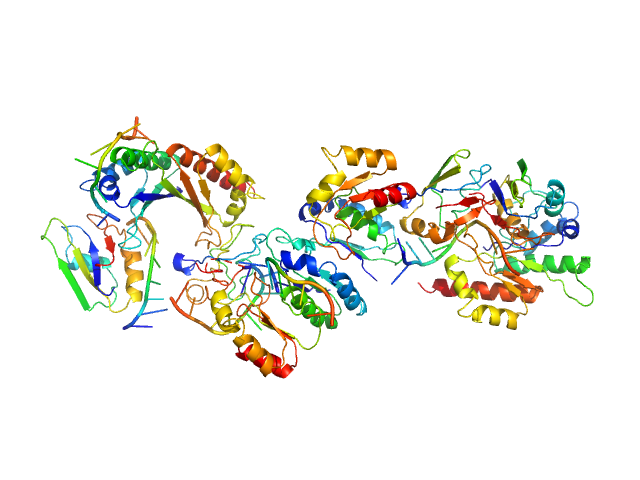
Insulin-like growth factor 2 mRNA-binding protein 2 dimer, 132 kDa Homo sapiens protein
Cox7B RNA monomer, 22 kDa RNA
|
| Buffer: |
20 mM HEPES, 2 mM MgCl2, 1 M NaCl, 10% glycerol v/v, 2 mM β-mercaptoethanol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2023 Nov 16
|
Structural insights into IMP2 dimerization and RNA binding
Raktim Roy
|
| RgGuinier |
3.8 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
UniProt ID: A0A150M835 (17-670) Acylamino-acid-releasing enzyme (I277L, V491A)
|
|
|
|
| Sample: |
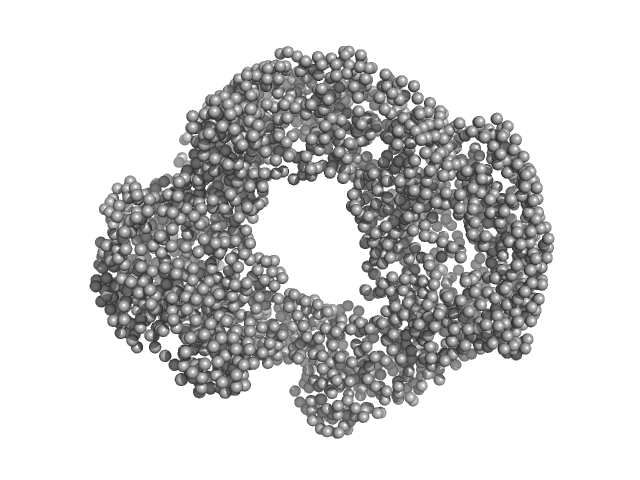
Acylamino-acid-releasing enzyme (I277L, V491A) tetramer, 308 kDa Geobacillus stearothermophilus protein
|
| Buffer: |
10 mM Tris, 100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BL-18, INDUS-2 on 2023 Aug 11
|
Structural and functional analysis of S9C peptidases of Geobacillus stearothermophilus
Dr. Rahul Singh
|
| RgGuinier |
5.4 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
424 |
nm3 |
|
|
UniProt ID: P42224 (1-750) Signal transducer and activator of transcription 1-alpha/beta
|
|
|
|
| Sample: |
Signal transducer and activator of transcription 1-alpha/beta tetramer, 349 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES-NaOH, 150 mM NaCl, 3% glycerol, 2 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2022 Dec 11
|
Structural analysis reveals how tetrameric tyrosine-phosphorylated STAT1 is targeted by the rabies virus P-protein
Science Signaling 18(878) (2025)
Sugiyama A, Minami M, Ugajin K, Inaba-Inoue S, Yabuno N, Takekawa Y, Xiaomei S, Takei S, Sasaki M, Nomai T, Jiang X, Kita S, Maenaka K, Hirose M, Yao M, Gooley P, Moseley G, Sugita Y, Ose T
|
| RgGuinier |
6.2 |
nm |
| Dmax |
21.2 |
nm |
| VolumePorod |
800 |
nm3 |
|
|
UniProt ID: P42224 (1-750) Signal transducer and activator of transcription 1-alpha/beta
UniProt ID: Q9IPJ8 (1-297) Phosphoprotein
|
|
|
|
| Sample: |
Signal transducer and activator of transcription 1-alpha/beta tetramer, 349 kDa Homo sapiens protein
Phosphoprotein monomer, 33 kDa Rabies virus (strain … protein
|
| Buffer: |
10 mM HEPES-NaOH, 150 mM NaCl, 3% glycerol, 2 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2022 Dec 11
|
Structural analysis reveals how tetrameric tyrosine-phosphorylated STAT1 is targeted by the rabies virus P-protein
Science Signaling 18(878) (2025)
Sugiyama A, Minami M, Ugajin K, Inaba-Inoue S, Yabuno N, Takekawa Y, Xiaomei S, Takei S, Sasaki M, Nomai T, Jiang X, Kita S, Maenaka K, Hirose M, Yao M, Gooley P, Moseley G, Sugita Y, Ose T
|
| RgGuinier |
6.5 |
nm |
| Dmax |
26.0 |
nm |
| VolumePorod |
857 |
nm3 |
|
|
UniProt ID: P42224 (1-750) Signal transducer and activator of transcription 1-alpha/beta
UniProt ID: None (None-None) Signal transducer and activator of transcription 1 binding DNA
|
|
|
|
| Sample: |
Signal transducer and activator of transcription 1-alpha/beta tetramer, 349 kDa Homo sapiens protein
Signal transducer and activator of transcription 1 binding DNA dimer, 11 kDa Homo sapiens DNA
|
| Buffer: |
10 mM HEPES-NaOH, 150 mM NaCl, 3% glycerol, 2 mM DTT, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2022 Dec 11
|
Structural analysis reveals how tetrameric tyrosine-phosphorylated STAT1 is targeted by the rabies virus P-protein
Science Signaling 18(878) (2025)
Sugiyama A, Minami M, Ugajin K, Inaba-Inoue S, Yabuno N, Takekawa Y, Xiaomei S, Takei S, Sasaki M, Nomai T, Jiang X, Kita S, Maenaka K, Hirose M, Yao M, Gooley P, Moseley G, Sugita Y, Ose T
|
| RgGuinier |
6.1 |
nm |
| Dmax |
19.4 |
nm |
| VolumePorod |
761 |
nm3 |
|
|
UniProt ID: Q88GF9 (1-50) H-NS family protein MvaT
|
|
|
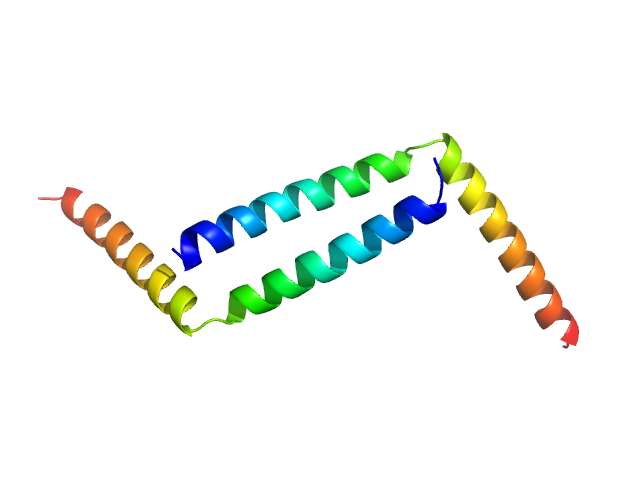
|
| Sample: |
H-NS family protein MvaT dimer, 12 kDa Pseudomonas putida (strain … protein
|
| Buffer: |
20 mM Tris-HCl, 500 mM NaCl, 10% glycerol, 500 mM Imidazole, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2019 May 24
|
Structural characterization of the native oligomerization mode of MvaT proteins in Pseudomonas.
Microbiol Spectr :e0023526 (2026)
Vasileva D, Suzuki-Minakuchi C, Arakawa T, Moriwaki Y, Yonezawa K, Shimizu N, Fujimoto Z, Terada T, Okada K, Nojiri H
|
| RgGuinier |
2.9 |
nm |
| Dmax |
14.0 |
nm |
|
|
UniProt ID: Q9UQM7 (345-475) Calcium/calmodulin-dependent protein kinase type II subunit alpha
|
|
|
|
| Sample: |
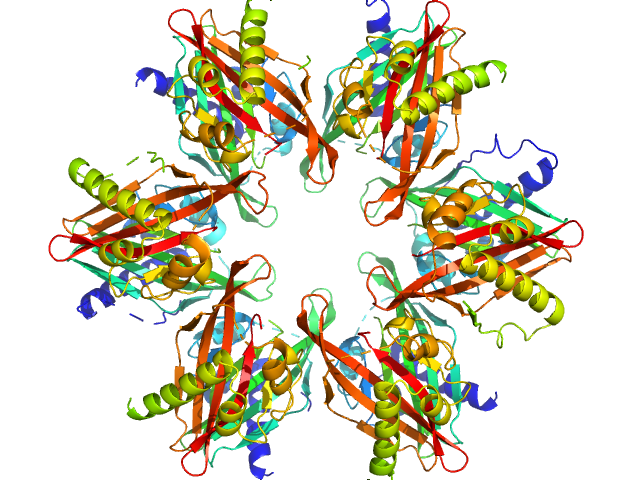
Calcium/calmodulin-dependent protein kinase type II subunit alpha dodecamer, 186 kDa Homo sapiens protein
|
| Buffer: |
20 mM MES, 150 mM NaCl, 1 mM DTT, pH: 6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Oct 3
|
Ligand‐induced CaMKIIα
hub Trp403 flip, hub domain stacking, and modulation of kinase activity
Protein Science 33(10) (2024)
Narayanan D, Larsen A, Gauger S, Adafia R, Hammershøi R, Hamborg L, Bruus‐Jensen J, Griem‐Krey N, Gee C, Frølund B, Stratton M, Kuriyan J, Kastrup J, Langkilde A, Wellendorph P, Solbak S
|
| RgGuinier |
4.5 |
nm |
| Dmax |
15.2 |
nm |
| VolumePorod |
367 |
nm3 |
|
|
UniProt ID: Q9UQM7 (345-475) Calcium/calmodulin-dependent protein kinase type II subunit alpha
|
|
|
|
| Sample: |
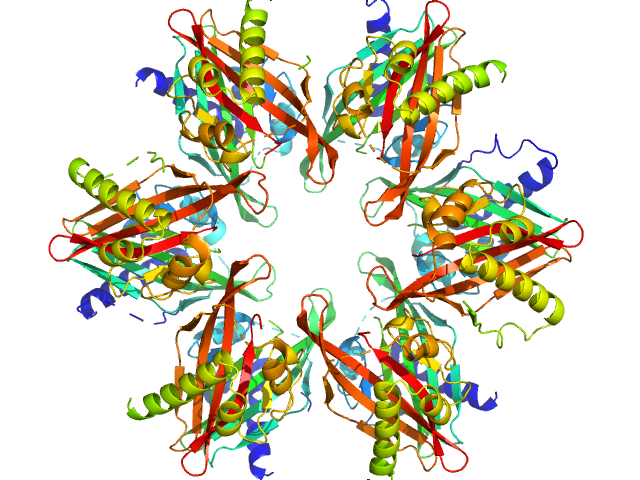
Calcium/calmodulin-dependent protein kinase type II subunit alpha dodecamer, 186 kDa Homo sapiens protein
|
| Buffer: |
20 mM MES, 150 mM NaCl, 1 mM DTT, pH: 6 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, University of Copenhagen, Department of Drug Design and Pharmacology on 2020 Nov 19
|
Ligand‐induced CaMKIIα
hub Trp403 flip, hub domain stacking, and modulation of kinase activity
Protein Science 33(10) (2024)
Narayanan D, Larsen A, Gauger S, Adafia R, Hammershøi R, Hamborg L, Bruus‐Jensen J, Griem‐Krey N, Gee C, Frølund B, Stratton M, Kuriyan J, Kastrup J, Langkilde A, Wellendorph P, Solbak S
|
| RgGuinier |
4.4 |
nm |
| Dmax |
14.9 |
nm |
| VolumePorod |
351 |
nm3 |
|
|
UniProt ID: Q9UQM7 (345-475) Calcium/calmodulin-dependent protein kinase type II subunit alpha
UniProt ID: None (None-None) 2-(6-(4-chlorophenyl)imidazo[1,2-b]pyridazine-2-yl)acetic acid
|
|
|
|
| Sample: |
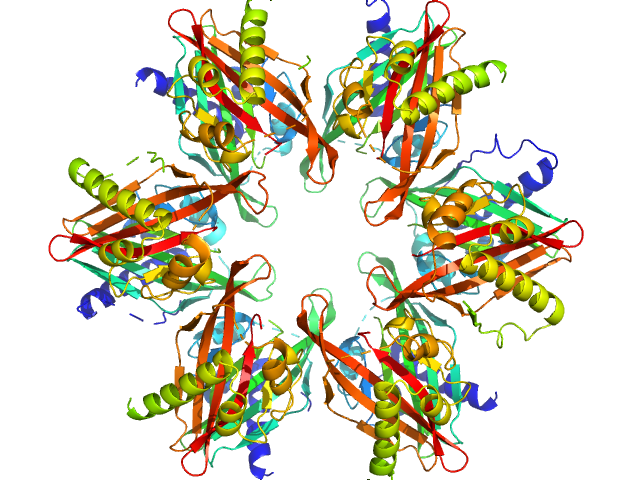
Calcium/calmodulin-dependent protein kinase type II subunit alpha dodecamer, 186 kDa Homo sapiens protein
2-(6-(4-chlorophenyl)imidazo[1,2-b]pyridazine-2-yl)acetic acid monomer, 0 kDa
|
| Buffer: |
20 mM MES, 150 mM NaCl, 1 mM DTT, 200 µM PIPA, pH: 6 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, University of Copenhagen, Department of Drug Design and Pharmacology on 2022 Nov 30
|
Ligand‐induced CaMKIIα
hub Trp403 flip, hub domain stacking, and modulation of kinase activity
Protein Science 33(10) (2024)
Narayanan D, Larsen A, Gauger S, Adafia R, Hammershøi R, Hamborg L, Bruus‐Jensen J, Griem‐Krey N, Gee C, Frølund B, Stratton M, Kuriyan J, Kastrup J, Langkilde A, Wellendorph P, Solbak S
|
| RgGuinier |
4.8 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
396 |
nm3 |
|
|
UniProt ID: Q9UQM7 (345-475) Calcium/calmodulin-dependent protein kinase type II subunit alpha
UniProt ID: None (None-None) 2-(6-(4-chlorophenyl)imidazo[1,2-b]pyridazine-2-yl)acetic acid
|
|
|
|
| Sample: |
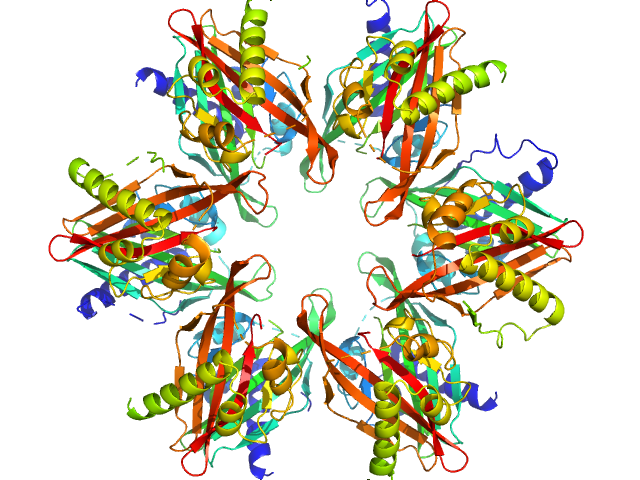
Calcium/calmodulin-dependent protein kinase type II subunit alpha dodecamer, 186 kDa Homo sapiens protein
2-(6-(4-chlorophenyl)imidazo[1,2-b]pyridazine-2-yl)acetic acid monomer, 0 kDa
|
| Buffer: |
20 mM MES, 150 mM NaCl, 1 mM DTT, 200 µM PIPA, pH: 6 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, University of Copenhagen, Department of Drug Design and Pharmacology on 2023 May 30
|
Ligand‐induced CaMKIIα
hub Trp403 flip, hub domain stacking, and modulation of kinase activity
Protein Science 33(10) (2024)
Narayanan D, Larsen A, Gauger S, Adafia R, Hammershøi R, Hamborg L, Bruus‐Jensen J, Griem‐Krey N, Gee C, Frølund B, Stratton M, Kuriyan J, Kastrup J, Langkilde A, Wellendorph P, Solbak S
|
| RgGuinier |
5.5 |
nm |
| Dmax |
27.0 |
nm |
|
|