UniProt ID: P40818 (7-1110) Ubiquitin carboxyl-terminal hydrolase 8
|
|
|

|
| Sample: |
Ubiquitin carboxyl-terminal hydrolase 8 monomer, 127 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Jun 18
|
Autoinhibition of ubiquitin-specific protease 8: insights into domain interactions and mechanisms of regulation
Journal of Biological Chemistry :107727 (2024)
Caba C, Black M, Liu Y, DaDalt A, Mallare J, Fan L, Harding R, Wang Y, Vacratsis P, Huang R, Zhuang Z, Tong Y
|
| RgGuinier |
8.4 |
nm |
| Dmax |
31.8 |
nm |
| VolumePorod |
327 |
nm3 |
|
|
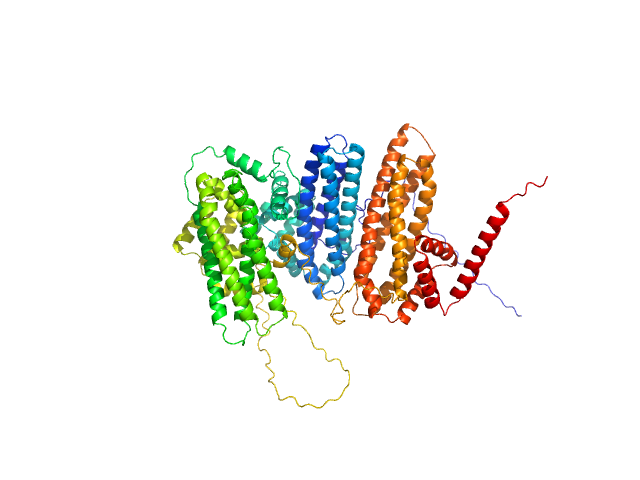
UniProt ID: F4IDY5 (1-831) Zinc finger protein BRUTUS-like At1g18910
|
|
|
|
| Sample: |
Zinc finger protein BRUTUS-like At1g18910 monomer, 98 kDa Arabidopsis thaliana protein
|
| Buffer: |
10 mM MES, 15 mM NaCl, pH: 6.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Feb 8
|
Iron-sensing and redox properties of the hemerythrin-like domains of Arabidopsis BRUTUS and BRUTUS-LIKE2 proteins.
Nat Commun 16(1):3865 (2025)
Pullin J, Rodríguez-Celma J, Franceschetti M, Mundy JEA, Svistunenko DA, Bradley JM, Le Brun NE, Balk J
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
162 |
nm3 |
|
|
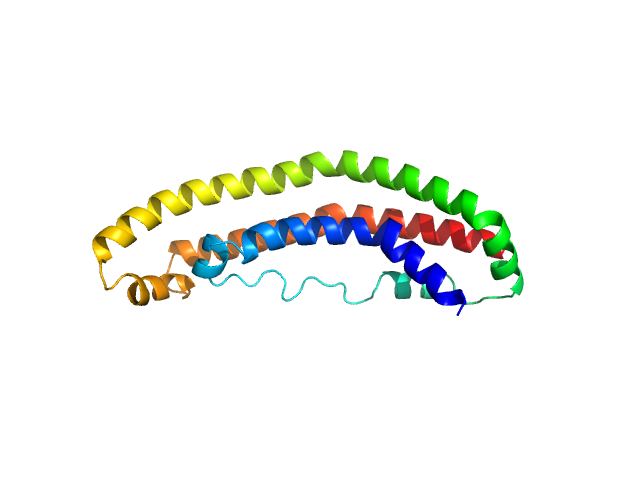
UniProt ID: P02647 (25-208) Apolipoprotein A-I
|
|
|
|
| Sample: |
Apolipoprotein A-I monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.1% sodium azide, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2021 Mar 20
|
The structure of the apolipoprotein A-I monomer provides insights into its oligomerisation and lipid-binding mechanisms
Journal of Molecular Biology :169394 (2025)
Tou H, Rosenes Z, Khandokar Y, Zlatic C, Metcalfe R, Mok Y, Morton C, Gooley P, Griffin M
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
27 |
nm3 |
|
|
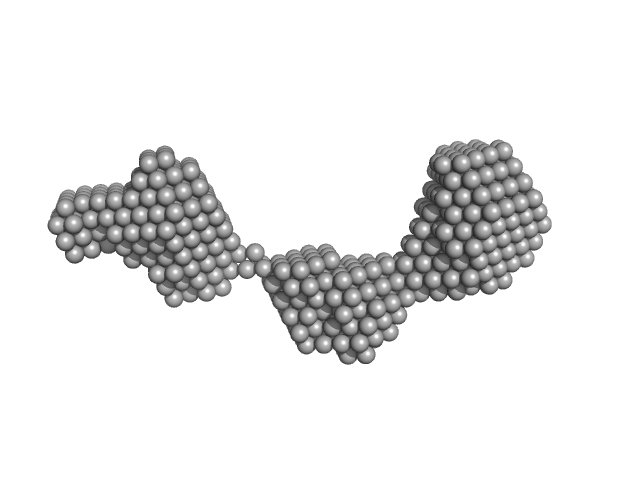
UniProt ID: P02647 (25-208) Apolipoprotein A-I
|
|
|

|
| Sample: |
Apolipoprotein A-I dimer, 43 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.1% sodium azide, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2021 Mar 20
|
The structure of the apolipoprotein A-I monomer provides insights into its oligomerisation and lipid-binding mechanisms
Journal of Molecular Biology :169394 (2025)
Tou H, Rosenes Z, Khandokar Y, Zlatic C, Metcalfe R, Mok Y, Morton C, Gooley P, Griffin M
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
58 |
nm3 |
|
|
UniProt ID: P02647 (25-208) Apolipoprotein A-I
|
|
|
|
| Sample: |
Apolipoprotein A-I monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.1% sodium azide, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Mar 1
|
The structure of the apolipoprotein A-I monomer provides insights into its oligomerisation and lipid-binding mechanisms
Journal of Molecular Biology :169394 (2025)
Tou H, Rosenes Z, Khandokar Y, Zlatic C, Metcalfe R, Mok Y, Morton C, Gooley P, Griffin M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
32 |
nm3 |
|
|
UniProt ID: P02647 (25-208) Apolipoprotein A-I (G50R)
|
|
|
|
| Sample: |
Apolipoprotein A-I (G50R) monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.1% sodium azide, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2023 Jun 27
|
The structure of the apolipoprotein A-I monomer provides insights into its oligomerisation and lipid-binding mechanisms
Journal of Molecular Biology :169394 (2025)
Tou H, Rosenes Z, Khandokar Y, Zlatic C, Metcalfe R, Mok Y, Morton C, Gooley P, Griffin M
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
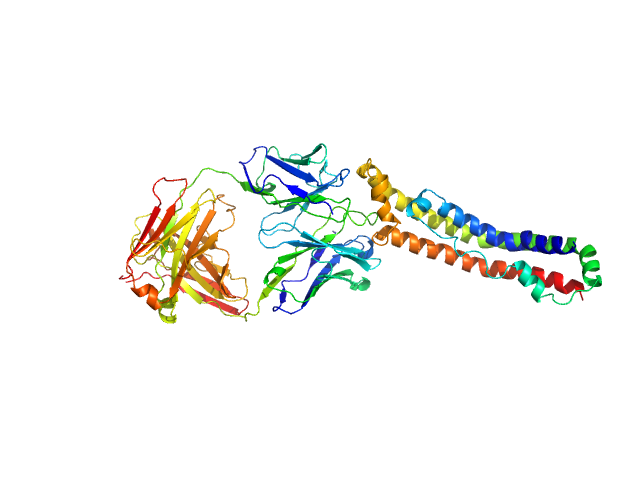
UniProt ID: P02647 (25-267) Apolipoprotein A-I
UniProt ID: None (None-None) Antigen-binding fragment 55201
|
|
|
|
| Sample: |
Apolipoprotein A-I monomer, 28 kDa Homo sapiens protein
Antigen-binding fragment 55201 monomer, 45 kDa Mus musculus protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.1% sodium azide, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2021 Jun 19
|
The structure of the apolipoprotein A-I monomer provides insights into its oligomerisation and lipid-binding mechanisms
Journal of Molecular Biology :169394 (2025)
Tou H, Rosenes Z, Khandokar Y, Zlatic C, Metcalfe R, Mok Y, Morton C, Gooley P, Griffin M
|
| RgGuinier |
4.6 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
99 |
nm3 |
|
|
UniProt ID: O65258 (56-542) Beta-amylase 2, chloroplastic
|
|
|
|
| Sample: |
Beta-amylase 2, chloroplastic tetramer, 229 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HEPES, 100 mM KCl, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Sep 17
|
Potassium cations expand the conformation ensemble of Arabidopsis thaliana β-amylase2 (BAM2).
MicroPubl Biol 2024 (2024)
Sholes A, Asongakap R, Jaconski S, Monroe J, Berndsen CE
|
| RgGuinier |
4.9 |
nm |
| Dmax |
16.4 |
nm |
| VolumePorod |
226 |
nm3 |
|
|
UniProt ID: O65258 (56-542) Beta-amylase 2, chloroplastic
|
|
|
|
| Sample: |
Beta-amylase 2, chloroplastic tetramer, 229 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HEPES, 100 mM KCl, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Sep 17
|
Potassium cations expand the conformation ensemble of Arabidopsis thaliana β-amylase2 (BAM2).
MicroPubl Biol 2024 (2024)
Sholes A, Asongakap R, Jaconski S, Monroe J, Berndsen CE
|
| RgGuinier |
4.9 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
234 |
nm3 |
|
|
UniProt ID: O65258 (56-542) Beta-amylase 2, chloroplastic
|
|
|
|
| Sample: |
Beta-amylase 2, chloroplastic tetramer, 229 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM HEPES, 100 mM KCl, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Sep 17
|
Potassium cations expand the conformation ensemble of Arabidopsis thaliana β-amylase2 (BAM2).
MicroPubl Biol 2024 (2024)
Sholes A, Asongakap R, Jaconski S, Monroe J, Berndsen CE
|
| RgGuinier |
4.7 |
nm |
| Dmax |
15.1 |
nm |
| VolumePorod |
218 |
nm3 |
|
|