UniProt ID: P43405 (6-269) Tyrosine-protein kinase SYK
UniProt ID: O43914 (88-107) TYRO protein tyrosine kinase-binding protein
|
|
|
|
| Sample: |
Tyrosine-protein kinase SYK monomer, 30 kDa Homo sapiens protein
TYRO protein tyrosine kinase-binding protein monomer, 2 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES, 150 mM NaCl, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2023 Nov 13
|
The mechanism of allosteric activation of SYK kinase derived from multiple phospho-ITAM-bound structures
Structure (2024)
Bradshaw W, Harris G, Gileadi O, Katis V
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
53 |
nm3 |
|
|
UniProt ID: Q9P8F4 (1-860) beta-glucosidase
|
|
|

|
| Sample: |
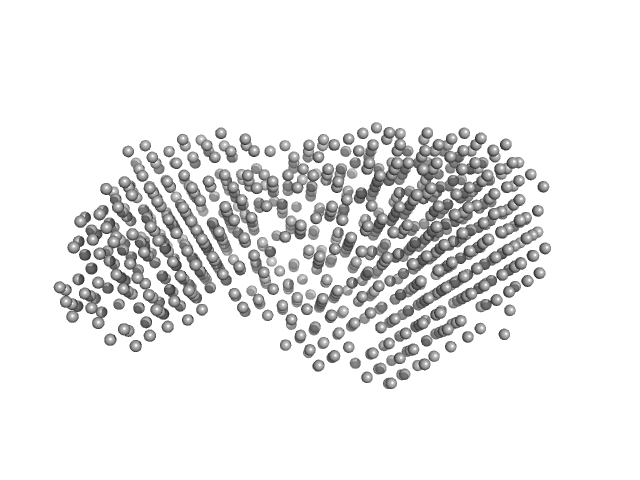
Beta-glucosidase dimer, 187 kDa Aspergillus niger protein
|
| Buffer: |
50 mM sodium acetate, pH: 5 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2024 Feb 15
|
Aspergillus niger beta-glucosidase (bgl1)
Estella Yee
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
253 |
nm3 |
|
|
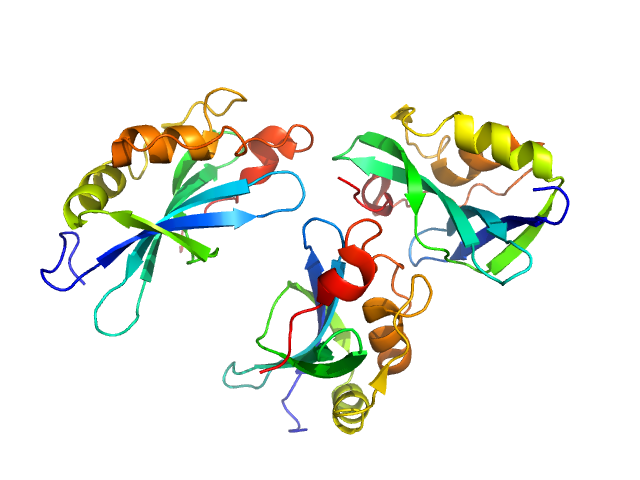
UniProt ID: O91464 (1017-1152) Genome polyprotein
|
|
|
|
| Sample: |
Genome polyprotein monomer, 15 kDa Aichi virus (strain … protein
|
| Buffer: |
20 mM HEPES,150 mM NaCl, 2 mM TCEP, 1% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 11
|
Structural and functional repurposing of picornaviral H-box 2A proteins
Marion Pichon
|
| RgGuinier |
1.8 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
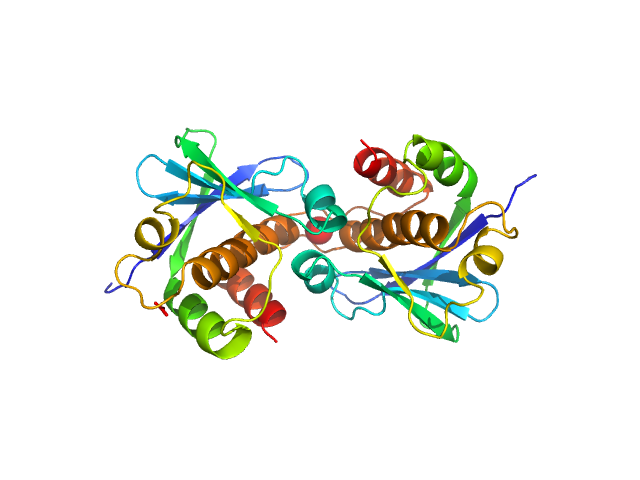
UniProt ID: Q66578 (777-923) Genome polyprotein (I918M)
|
|
|
|
| Sample: |
Genome polyprotein (I918M) dimer, 34 kDa Human parechovirus 1 … protein
|
| Buffer: |
20 mM HEPES,150 mM NaCl, 2 mM TCEP, 1% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 11
|
Structural and functional repurposing of picornaviral H-box 2A proteins
Marion Pichon
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.0 |
nm |
| VolumePorod |
58 |
nm3 |
|
|
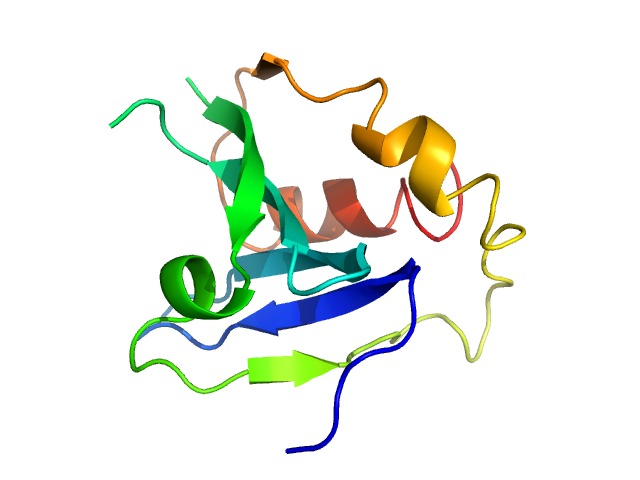
UniProt ID: P53816 (1-132) Phospholipase A and acyltransferase 3
|
|
|
|
| Sample: |
Phospholipase A and acyltransferase 3 monomer, 15 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES,150 mM NaCl, 2 mM TCEP, 1% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Mar 11
|
Structural and functional repurposing of picornaviral H-box 2A proteins
Marion Pichon
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.5 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
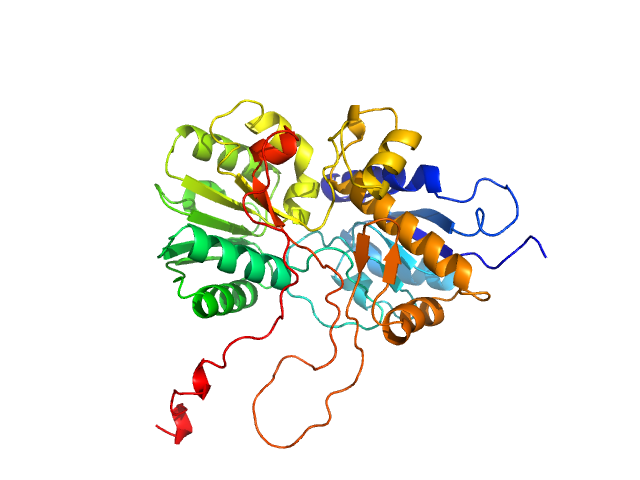
UniProt ID: Q8WPR6 (480-859) adenylate cyclase
|
|
|
|
| Sample: |
Adenylate cyclase monomer, 43 kDa Trypanosoma brucei protein
|
| Buffer: |
50 mM Tris-HCl, 500 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Sep 23
|
Biophysical analysis of the membrane-proximal Venus Flytrap domain of ESAG4 receptor-like adenylate cyclase from Trypanosoma brucei.
Mol Biochem Parasitol 260:111653 (2024)
Alves DO, Geens R, da Silva Arruda HR, Jennen L, Corthaut S, Wuyts E, de Andrade GC, Prosdocimi F, Cordeiro Y, Pires JR, Vieira LR, de Oliveira GAP, Sterckx YG, Salmon D
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.6 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
UniProt ID: P0AEX9 (27-384) Maltose/maltodextrin-binding periplasmic protein (D108A, K109A, E198A, N199A, K265A)
UniProt ID: P9WPH5 (19-409) Uncharacterized protein Rv2242
|
|
|
|
| Sample: |
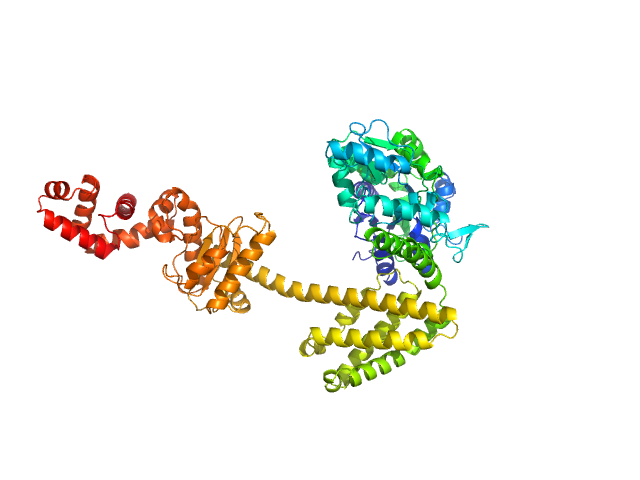
Maltose/maltodextrin-binding periplasmic protein (D108A, K109A, E198A, N199A, K265A) monomer, 40 kDa Escherichia coli (strain … protein
Uncharacterized protein Rv2242 monomer, 43 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris pH 7.5, 150 mM NaCl, pH: |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2024 Apr 3
|
Domain architecture of the Mycobacterium tuberculosis MabR (Rv2242), a member of the PucR transcription factor family
Heliyon :e40494 (2024)
Megalizzi V, Tanina A, Grosse C, Mirgaux M, Legrand P, Mirandela G, Wohlkönig A, Bifani P, Wintjens R
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
113 |
nm3 |
|
|
UniProt ID: P0AEX9 (27-384) Maltose/maltodextrin-binding periplasmic protein (D108A, K109A, E198A, N199A, K265A)
UniProt ID: P9WPH5 (19-409) Uncharacterized protein Rv2242
|
|
|
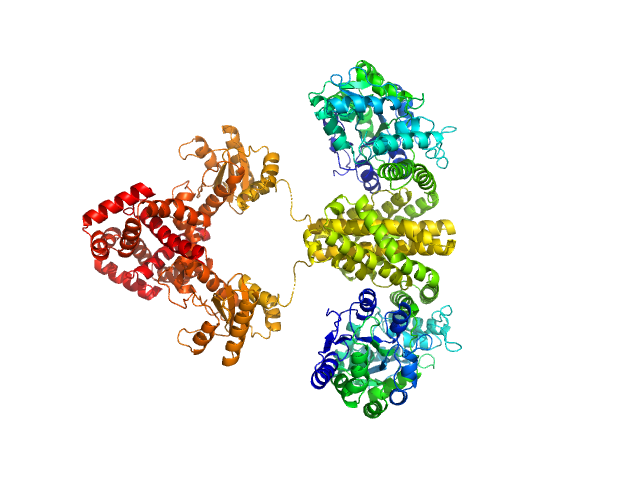
|
| Sample: |
Maltose/maltodextrin-binding periplasmic protein (D108A, K109A, E198A, N199A, K265A) dimer, 80 kDa Escherichia coli (strain … protein
Uncharacterized protein Rv2242 dimer, 86 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris pH 7.5, 150 mM NaCl, pH: |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2024 Apr 3
|
Domain architecture of the Mycobacterium tuberculosis MabR (Rv2242), a member of the PucR transcription factor family
Heliyon :e40494 (2024)
Megalizzi V, Tanina A, Grosse C, Mirgaux M, Legrand P, Mirandela G, Wohlkönig A, Bifani P, Wintjens R
|
| RgGuinier |
3.7 |
nm |
| Dmax |
10.8 |
nm |
| VolumePorod |
345 |
nm3 |
|
|
UniProt ID: P0AEX9 (27-384) Maltose/maltodextrin-binding periplasmic protein (D108A, K109A, E198A, N199A, K265A)
UniProt ID: P9WPH5 (19-409) Uncharacterized protein Rv2242
|
|
|
|
| Sample: |
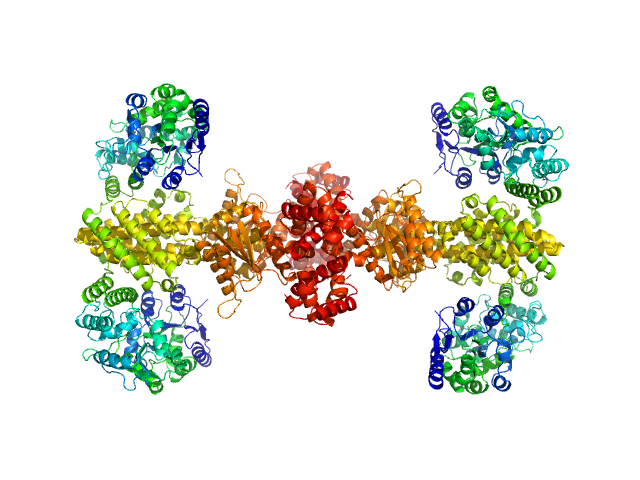
Maltose/maltodextrin-binding periplasmic protein (D108A, K109A, E198A, N199A, K265A) tetramer, 161 kDa Escherichia coli (strain … protein
Uncharacterized protein Rv2242 tetramer, 171 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris pH 7.5, 150 mM NaCl, pH: |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2024 Apr 3
|
Domain architecture of the Mycobacterium tuberculosis MabR (Rv2242), a member of the PucR transcription factor family
Heliyon :e40494 (2024)
Megalizzi V, Tanina A, Grosse C, Mirgaux M, Legrand P, Mirandela G, Wohlkönig A, Bifani P, Wintjens R
|
| RgGuinier |
5.2 |
nm |
| Dmax |
15.8 |
nm |
| VolumePorod |
826 |
nm3 |
|
|
UniProt ID: O43186 (31-107) Cone-rod homeobox protein
UniProt ID: None (None-None) Ret4 sense strand
UniProt ID: None (None-None) Ret4 antisense strand
|
|
|
|
| Sample: |
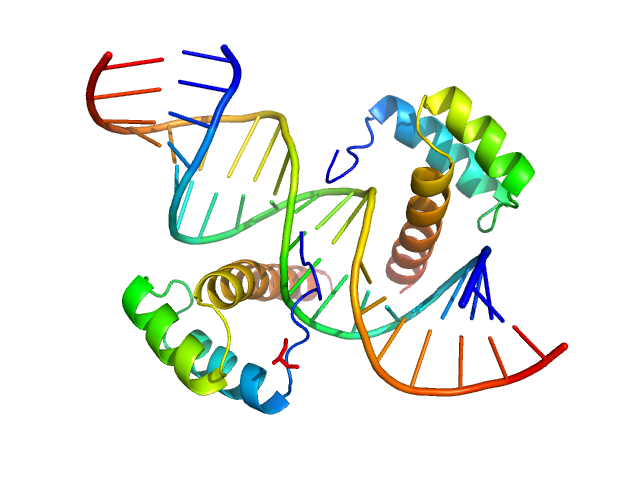
Cone-rod homeobox protein, 20 kDa Homo sapiens protein
Ret4 sense strand, 6 kDa synthetic construct DNA
Ret4 antisense strand, 6 kDa synthetic construct DNA
|
| Buffer: |
20 mM Tris, 150 mM KCl, 5 % glycerol, 1 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Mar 25
|
Molecular basis of CRX/DNA recognition and stoichiometry at the Ret4 response element
Structure (2024)
Srivastava D, Gowribidanur-Chinnaswamy P, Gaur P, Spies M, Swaroop A, Artemyev N
|
| RgGuinier |
2.2 |
nm |
| Dmax |
6.8 |
nm |
| VolumePorod |
37 |
nm3 |
|
|