UniProt ID: Q01962 (33-713) Peroxisomal biogenesis factor 8
UniProt ID: P33292 (1-276) Peroxisomal targeting signal receptor
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDXB4_fit1_model1.png)
|
| Sample: |
Peroxisomal biogenesis factor 8 monomer, 78 kDa Komagataella pastoris protein
Peroxisomal targeting signal receptor monomer, 32 kDa Komagataella pastoris protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Aug 17
|
Structure of Pex8 in complex with peroxisomal receptor Pex5 reveals its essential role in peroxisomal cargo translocation
(2025)
Ekal L, Wendscheck D, David Y, Chojnowski G, Jeffries C, Mullapudi E, Schuldiner M, Warscheid B, Zalckvar E, Wilmanns M
|
| RgGuinier |
4.6 |
nm |
| Dmax |
19.0 |
nm |
| VolumePorod |
177 |
nm3 |
|
|
UniProt ID: Q01962 (33-713) Peroxisomal biogenesis factor 8
UniProt ID: P33292 (259-576) Peroxisomal targeting signal receptor
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDXC4_fit1_model1.png)
|
| Sample: |
Peroxisomal biogenesis factor 8 monomer, 78 kDa Komagataella pastoris protein
Peroxisomal targeting signal receptor monomer, 36 kDa Komagataella pastoris protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Aug 17
|
Structure of Pex8 in complex with peroxisomal receptor Pex5 reveals its essential role in peroxisomal cargo translocation
(2025)
Ekal L, Wendscheck D, David Y, Chojnowski G, Jeffries C, Mullapudi E, Schuldiner M, Warscheid B, Zalckvar E, Wilmanns M
|
| RgGuinier |
4.2 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
141 |
nm3 |
|
|
UniProt ID: P23246 (214-707) Splicing factor, proline- and glutamine-rich
|
|
|
|
| Sample: |
Splicing factor, proline- and glutamine-rich dimer, 153 kDa Homo sapiens protein
|
| Buffer: |
150 mM KCl, 1.5% glycerol, 20 mM HEPES, 1 mM DTT (in 100% H2O), pH: 7.4 |
| Experiment: |
SANS
data collected at Quokka - Small Angle Neutron Scattering, Australian Centre for Neutron Scattering (ANSTO) on 2023 Jan 23
|
Structural dynamics of IDR interactions in human SFPQ and implications for liquid–liquid phase separation
Acta Crystallographica Section D Structural Biology 81(7):357-379 (2025)
Koning H, Lai V, Sethi A, Chakraborty S, Ang C, Fox A, Duff A, Whitten A, Marshall A, Bond C
|
| RgGuinier |
8.7 |
nm |
| Dmax |
30.7 |
nm |
| VolumePorod |
241000 |
nm3 |
|
|
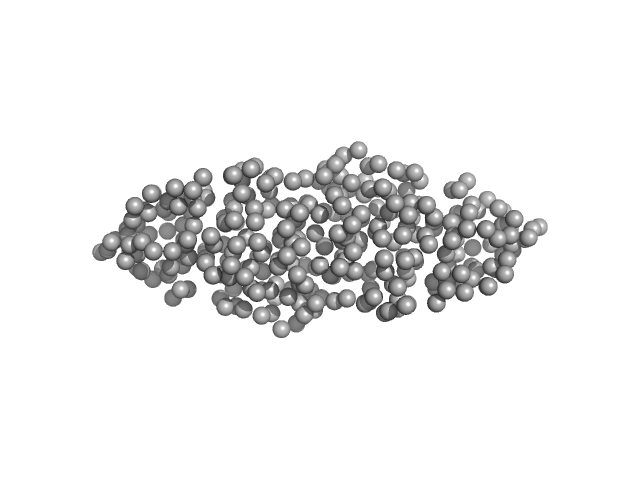
UniProt ID: Q1D0B6 (1-120) Mutual gliding motility protein C
|
|
|

|
| Sample: |
Mutual gliding motility protein C dimer, 32 kDa Myxococcus xanthus (strain … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 10% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Mar 31
|
Structural and biophysical insights into RomR, MglB and MglC interactions involved in regulating cell polarity in Myxococcus xanthus
Journal of Biological Chemistry :110907 (2025)
Kodesia A, Kapoor S, Thakur K
|
| RgGuinier |
2.3 |
nm |
| Dmax |
9.8 |
nm |
| VolumePorod |
49 |
nm3 |
|
|
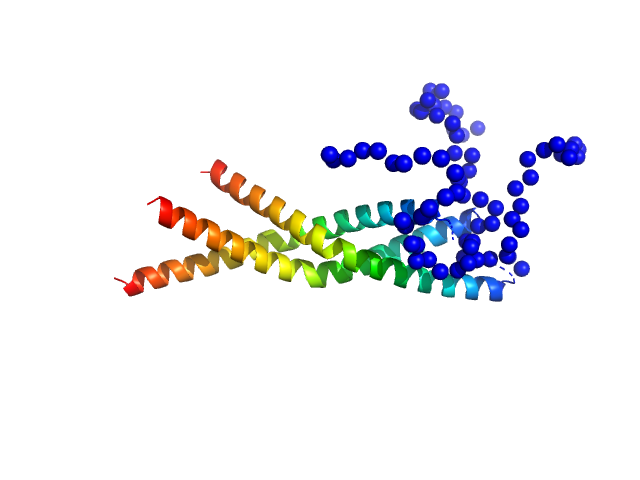
UniProt ID: A0AAE6G1Y1 (372-421) Two-component system response regulator
|
|
|
|
| Sample: |
Two-component system response regulator trimer, 26 kDa Myxococcus xanthus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Mar 31
|
Structural and biophysical insights into RomR, MglB and MglC interactions involved in regulating cell polarity in Myxococcus xanthus
Journal of Biological Chemistry :110907 (2025)
Kodesia A, Kapoor S, Thakur K
|
| RgGuinier |
2.7 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
43 |
nm3 |
|
|
UniProt ID: A0AAE6G1Y1 (328-421) Two-component system response regulator
|
|
|
|
| Sample: |
Two-component system response regulator trimer, 39 kDa Myxococcus xanthus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 10% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Mar 31
|
Structural and biophysical insights into RomR, MglB and MglC interactions involved in regulating cell polarity in Myxococcus xanthus
Journal of Biological Chemistry :110907 (2025)
Kodesia A, Kapoor S, Thakur K
|
| RgGuinier |
4.1 |
nm |
| Dmax |
16.7 |
nm |
| VolumePorod |
81 |
nm3 |
|
|
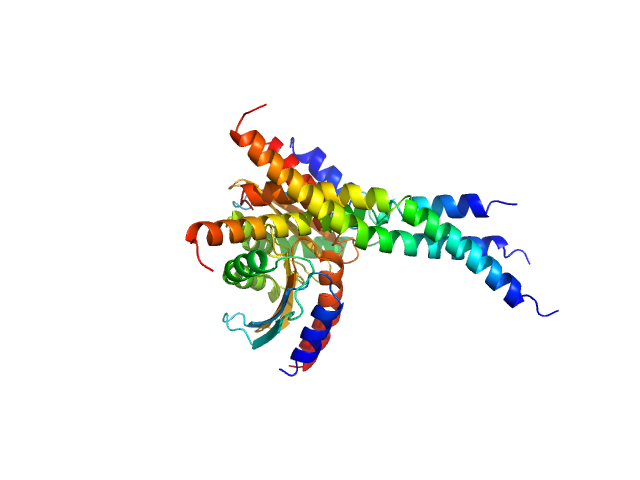
UniProt ID: Q1D0B6 (1-120) GTPase
UniProt ID: A0AAE6G1Y1 (None-None) Two-component system response regulator
|
|
|
|
| Sample: |
GTPase dimer, 32 kDa Myxococcus xanthus (strain … protein
Two-component system response regulator trimer, 26 kDa Myxococcus xanthus DK … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 10% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Mar 31
|
Structural and biophysical insights into RomR, MglB and MglC interactions involved in regulating cell polarity in Myxococcus xanthus
Journal of Biological Chemistry :110907 (2025)
Kodesia A, Kapoor S, Thakur K
|
| RgGuinier |
2.8 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
91 |
nm3 |
|
|
UniProt ID: Q1D0B6 (1-120) GTPase
UniProt ID: A0AAE6G1Y1 (328-421) Two-component system response regulator
|
|
|
|
| Sample: |
GTPase dimer, 32 kDa Myxococcus xanthus (strain … protein
Two-component system response regulator trimer, 39 kDa Myxococcus xanthus protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 10% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Mar 31
|
Structural and biophysical insights into RomR, MglB and MglC interactions involved in regulating cell polarity in Myxococcus xanthus
Journal of Biological Chemistry :110907 (2025)
Kodesia A, Kapoor S, Thakur K
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
79 |
nm3 |
|
|
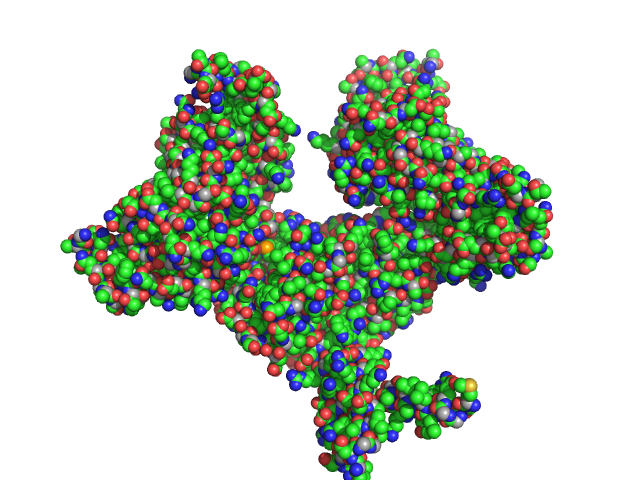
UniProt ID: A0AAE6G1Y1 (None-None) Two-component system response regulator
UniProt ID: Q1D0B6 (None-None) Mutual gliding motility protein C
UniProt ID: Q1DB03 (11-133) Gliding motility protein MglB
|
|
|

|
| Sample: |
Two-component system response regulator trimer, 26 kDa Myxococcus xanthus DK … protein
Mutual gliding motility protein C dimer, 32 kDa Myxococcus xanthus (strain … protein
Gliding motility protein MglB dimer, 32 kDa Myxococcus xanthus (strain … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 10% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Mar 31
|
Structural and biophysical insights into RomR, MglB and MglC interactions involved in regulating cell polarity in Myxococcus xanthus
Journal of Biological Chemistry :110907 (2025)
Kodesia A, Kapoor S, Thakur K
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.6 |
nm |
| VolumePorod |
204 |
nm3 |
|
|
UniProt ID: Q504U8 (651-993) Receptor protein-tyrosine kinase (duplication mutant)
|
|
|
|
| Sample: |
Receptor protein-tyrosine kinase (duplication mutant) monomer, 80 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 250 mM NaCl, 250 mM KCl, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax-007HF, Yale University on 2022 Mar 22
|
The role of kinase domain dimerization in EGFR activation
Zaritza Petrova
|
| RgGuinier |
4.9 |
nm |
| Dmax |
16.4 |
nm |
| VolumePorod |
163 |
nm3 |
|
|