UniProt ID: Q504U8 (651-993) Receptor protein-tyrosine kinase (duplication mutant)
|
|
|
|
| Sample: |
Receptor protein-tyrosine kinase (duplication mutant) monomer, 80 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 250 mM NaCl, 250 mM KCl, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax-007HF, Yale University on 2022 Mar 22
|
The role of kinase domain dimerization in EGFR activation
Zaritza Petrova
|
| RgGuinier |
4.5 |
nm |
| Dmax |
14.9 |
nm |
| VolumePorod |
190 |
nm3 |
|
|
UniProt ID: A0A2V4FWX5 (18-668) Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB
|
|
|

|
| Sample: |
Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB monomer, 73 kDa Serratia marcescens protein
|
| Buffer: |
phosphate-buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2025 Feb 3
|
Structural, biophysical and biochemical studies of CAZYmes involved in exopolysaccharides degradation of microbial biofilms
Universidade de São Paulo Master's thesis (2025)
Amanda Freitas Cruz
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.4 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
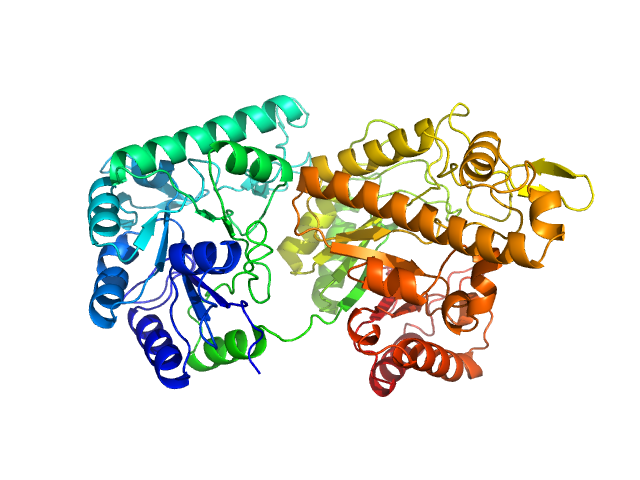
UniProt ID: A0A2V4FWX5 (40-653) Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB
|
|
|
|
| Sample: |
Poly-beta-1,6-N-acetyl-D-glucosamine N-deacetylase PgaB monomer, 69 kDa Serratia marcescens protein
|
| Buffer: |
phosphate-buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2025 Feb 3
|
Structural, biophysical and biochemical studies of CAZYmes involved in exopolysaccharides degradation of microbial biofilms
Universidade de São Paulo Master's thesis (2025)
Amanda Freitas Cruz
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
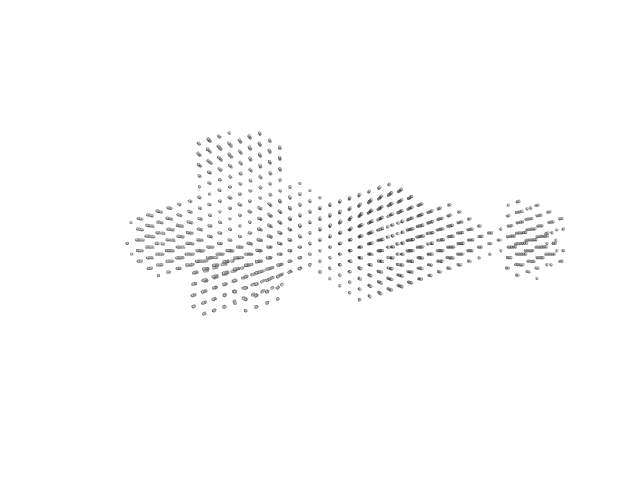
UniProt ID: Q1R2T6 (2-707) Alpha-hemolysin translocation ATP-binding protein HlyB
UniProt ID: A0A236M9V9 (1-1024) HlyA (GFP-Hemolysin with Flag-Tag insertion)
UniProt ID: Q1R2T7 (1-478) Membrane fusion protein (MFP) family protein (R14C; L229V)
UniProt ID: P02930 (1-493) Outer membrane protein TolC
|
|
|

|
| Sample: |
Alpha-hemolysin translocation ATP-binding protein HlyB hexamer, 480 kDa Escherichia coli (strain … protein
HlyA (GFP-Hemolysin with Flag-Tag insertion) monomer, 144 kDa Escherichia coli protein
Membrane fusion protein (MFP) family protein (R14C; L229V) hexamer, 327 kDa Escherichia coli (strain … protein
Outer membrane protein TolC trimer, 161 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM Tris, 150 mM NaCl, 10 mM CaCl2, 0.0063% glyco-diosgenin (GDN), pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Jun 29
|
Molecular Insights into CLD Domain Dynamics and Toxin Recruitment of the HlyA E. coli T1SS
Journal of Molecular Biology :169485 (2025)
Gentile R, Schott-Verdugo S, Khosa S, Günes C, Bonus M, Reiners J, Smits S, Schmitt L, Gohlke H
|
| RgGuinier |
11.4 |
nm |
| Dmax |
46.2 |
nm |
| VolumePorod |
2837 |
nm3 |
|
|
UniProt ID: Q9AES2 (31-1167) IgA protease
|
|
|
|
| Sample: |
IgA protease monomer, 127 kDa Thomasclavelia ramosa protein
|
| Buffer: |
50 mM Tris-HCl pH 7.5, 100 mM NaCl, 2% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Apr 19
|
Molecular basis of Fab-dependent IgA antibody recognition by gut-bacterial metallopeptidases
The EMBO Journal (2025)
Márquez-Moñino M, Martínez Gascueña A, Azzam T, Persson A, Manzanares-Gomez A, Aguillo-Urarte M, Brown T, Montero-Sagarminaga A, Lood R, Naegeli A, Connell S, Sastre D, Sundberg E, Trastoy B
|
| RgGuinier |
6.4 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
208 |
nm3 |
|
|
UniProt ID: Q9AES2 (31-878) IgA protease
|
|
|
|
| Sample: |
IgA protease monomer, 96 kDa Thomasclavelia ramosa protein
|
| Buffer: |
50 mM Tris-HCl pH 7.5, 100 mM NaCl, 2% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Apr 19
|
Molecular basis of Fab-dependent IgA antibody recognition by gut-bacterial metallopeptidases
The EMBO Journal (2025)
Márquez-Moñino M, Martínez Gascueña A, Azzam T, Persson A, Manzanares-Gomez A, Aguillo-Urarte M, Brown T, Montero-Sagarminaga A, Lood R, Naegeli A, Connell S, Sastre D, Sundberg E, Trastoy B
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
101 |
nm3 |
|
|
UniProt ID: Q9AES2 (323-878) IgA protease
|
|
|
|
| Sample: |
IgA protease monomer, 64 kDa Thomasclavelia ramosa protein
|
| Buffer: |
50 mM Tris-HCl pH 7.5, 100 mM NaCl, 2% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Apr 19
|
Molecular basis of Fab-dependent IgA antibody recognition by gut-bacterial metallopeptidases
The EMBO Journal (2025)
Márquez-Moñino M, Martínez Gascueña A, Azzam T, Persson A, Manzanares-Gomez A, Aguillo-Urarte M, Brown T, Montero-Sagarminaga A, Lood R, Naegeli A, Connell S, Sastre D, Sundberg E, Trastoy B
|
| RgGuinier |
3.0 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
UniProt ID: A0AAE9RTQ7 (43-265) Virulence factor family protein
|
|
|
|
| Sample: |
Virulence factor family protein monomer, 24 kDa Brucella abortus protein
|
| Buffer: |
50 mM Tris pH 8.0, 200 mM NaCl, 5% v/v glycerol, pH: |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2023 Jul 12
|
Crystal structure of the virulence protein J (VirJ) domain 1 from Brucella abortus.
Acta Crystallogr F Struct Biol Commun (2025)
Dugelay C, Ferrarin S, Terradot L
|
| RgGuinier |
1.8 |
nm |
| Dmax |
6.1 |
nm |
| VolumePorod |
39 |
nm3 |
|
|
UniProt ID: P51587 (1212-1551) Breast cancer type 2 susceptibility protein (BRC repeats 2-4)
UniProt ID: Q06609 (1-339) DNA repair protein RAD51 homolog 1 (F86E A89E)
|
|
|
|
| Sample: |
Breast cancer type 2 susceptibility protein (BRC repeats 2-4) monomer, 39 kDa Homo sapiens protein
DNA repair protein RAD51 homolog 1 (F86E A89E) trimer, 111 kDa Homo sapiens protein
|
| Buffer: |
20 mM K₂HPO₄/KH₂PO₄, 100 mM NaCl, 200 mM Li₂SO₄, 5% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Dec 2
|
Breaking new ground into RAD51–BRC repeats interplay in Homologous Recombination
(2025)
Rinaldi F, Franco P, Veronesi M, Romeo E, Bresciani V, Varignani G, Catalano F, Bernetti M, Masetti M, Langer J, Girotto S, Cavalli A
|
| RgGuinier |
6.5 |
nm |
| Dmax |
28.5 |
nm |
| VolumePorod |
334 |
nm3 |
|
|
UniProt ID: P51587 (1212-1551) Breast cancer type 2 susceptibility protein (BRC repeats 2-4)
UniProt ID: Q06609 (1-339) DNA repair protein RAD51 homolog 1 (F86E A89E)
|
|
|
|
| Sample: |
Breast cancer type 2 susceptibility protein (BRC repeats 2-4) monomer, 39 kDa Homo sapiens protein
DNA repair protein RAD51 homolog 1 (F86E A89E) trimer, 111 kDa Homo sapiens protein
|
| Buffer: |
20 mM K₂HPO₄/KH₂PO₄, 100 mM NaCl, 200 mM Li₂SO₄, 5% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Dec 2
|
Breaking new ground into RAD51–BRC repeats interplay in Homologous Recombination
(2025)
Rinaldi F, Franco P, Veronesi M, Romeo E, Bresciani V, Varignani G, Catalano F, Bernetti M, Masetti M, Langer J, Girotto S, Cavalli A
|
| RgGuinier |
6.5 |
nm |
| Dmax |
28.5 |
nm |
| VolumePorod |
473 |
nm3 |
|
|