UniProt ID: None (None-None) inverted repeat DNA
UniProt ID: Q96RI1-1 (110-466) Isoform 3 of Bile acid receptor
|
|
|
|
| Sample: |
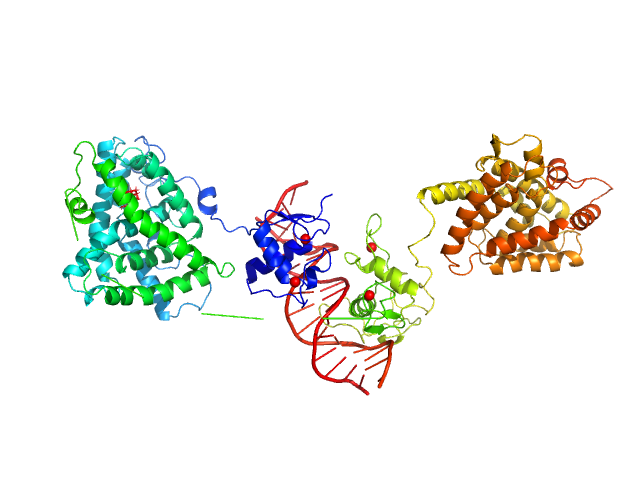
Inverted repeat DNA monomer, 11 kDa DNA
Isoform 3 of Bile acid receptor dimer, 83 kDa Homo sapiens protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 5 % glycerol, 1 mM DTT, 20 µM CDCA, pH: 7.4 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, Pennsylvania State University on 2024 May 9
|
DNA induces non-canonical dimerization of the farnesoid X receptor.
Nucleic Acids Res 54(4) (2026)
Khan SH, Yennawar N, Okafor CD
|
| RgGuinier |
5.1 |
nm |
| Dmax |
23.2 |
nm |
| VolumePorod |
120 |
nm3 |
|
|
UniProt ID: P02766 (21-147) Transthyretin
|
|
|
|
| Sample: |
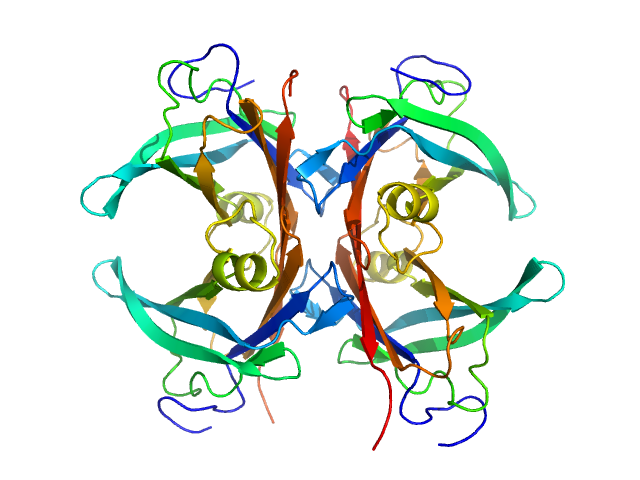
Transthyretin tetramer, 55 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2022 Sep 4
|
Differentiating the solution structures and stability of transthyretin tetramer complexed with tolcapone and tafamidis using SEC-SWAXS and NMR.
J Appl Crystallogr 58(Pt 4):1373-1383 (2025)
Shih O, Feng YC, Agrawal S, Liao KF, Yeh YQ, Chang JW, Yu TY, Jeng US
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.2 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
UniProt ID: P02766 (21-147) Transthyretin
UniProt ID: None (None-None) Tolcapone
|
|
|
|
| Sample: |
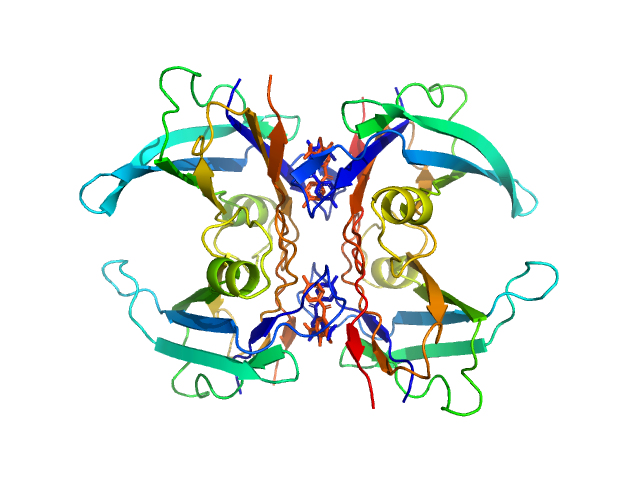
Transthyretin tetramer, 55 kDa Homo sapiens protein
Tolcapone dimer, 1 kDa
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2023 May 19
|
Differentiating the solution structures and stability of transthyretin tetramer complexed with tolcapone and tafamidis using SEC-SWAXS and NMR.
J Appl Crystallogr 58(Pt 4):1373-1383 (2025)
Shih O, Feng YC, Agrawal S, Liao KF, Yeh YQ, Chang JW, Yu TY, Jeng US
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
107 |
nm3 |
|
|
UniProt ID: P02766 (21-147) Transthyretin
UniProt ID: None (None-None) Tafamidis
|
|
|
|
| Sample: |
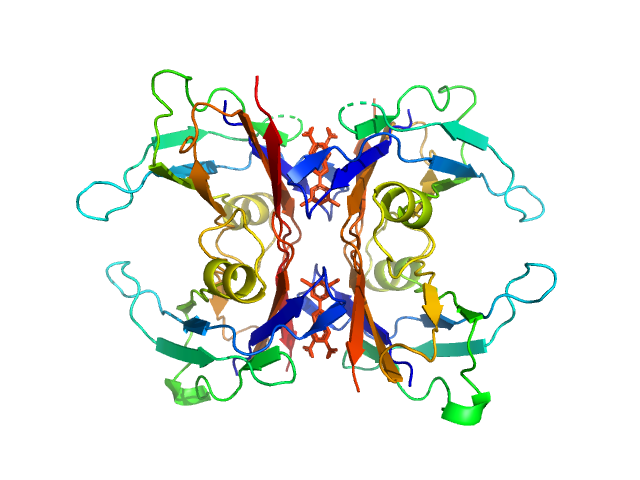
Transthyretin tetramer, 55 kDa Homo sapiens protein
Tafamidis dimer, 1 kDa
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at 13A, Taiwan Photon Source, NSRRC on 2023 May 19
|
Differentiating the solution structures and stability of transthyretin tetramer complexed with tolcapone and tafamidis using SEC-SWAXS and NMR.
J Appl Crystallogr 58(Pt 4):1373-1383 (2025)
Shih O, Feng YC, Agrawal S, Liao KF, Yeh YQ, Chang JW, Yu TY, Jeng US
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
69 |
nm3 |
|
|
UniProt ID: J7I2T6 (20-200) Perivitellin ovorubin-1
UniProt ID: J7HZ90 (26-210) Perivitellin ovorubin-2
UniProt ID: A0A2T7NVP5 (21-194) Uncharacterized protein (ovorubin-3, short)
UniProt ID: A0A2T7NVP6 (24-203) Perivitellin protein (ovorubin-4, form 1)
UniProt ID: A0A2T7NVP5 (419-602) Uncharacterized protein (ovorubin-5)
|
|
|
|
| Sample: |
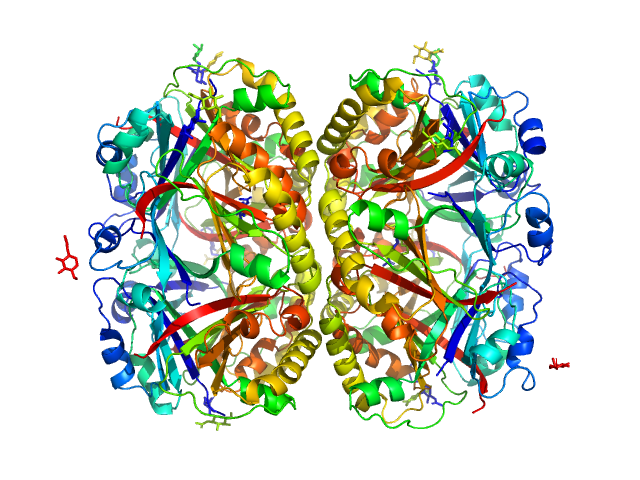
Perivitellin ovorubin-1 dimer, 41 kDa Pomacea canaliculata protein
Perivitellin ovorubin-2 dimer, 42 kDa Pomacea canaliculata protein
Uncharacterized protein (ovorubin-3, short) dimer, 40 kDa Pomacea canaliculata protein
Perivitellin protein (ovorubin-4, form 1) dimer, 40 kDa Pomacea canaliculata protein
Uncharacterized protein (ovorubin-5) dimer, 41 kDa Pomacea canaliculata protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at TPS13A, NSRRC on 2025 Mar 26
|
Structure of the chromoprotein ovorubin from the golden apple snail ( Pomacea canaliculata
)
Protein Science 35(1) (2025)
Wilasluck P, Saw W, Tran B, Sabat G, Shih O, Hengphasatporn K, Shigeta Y, Vinayavekhin N, Wangkanont K
|
| RgGuinier |
4.1 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
400 |
nm3 |
|
|
UniProt ID: Q2FDS6 (1-276) Uncharacterized hydrolase SAUSA300_2518
|
|
|
|
| Sample: |
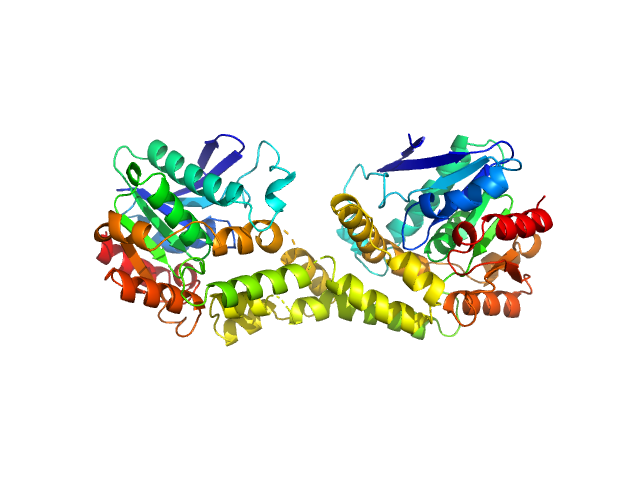
Uncharacterized hydrolase SAUSA300_2518 dimer, 62 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
10 mM HEPES, 50 mM NaCl, 0.1% w/v NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2025 Mar 26
|
Unique structural and ligand-binding properties of the Staphylococcus aureus
serine hydrolase FphE
Proceedings of the National Academy of Sciences 123(13) (2026)
Jo J, Upadhyay T, You X, Bennett J, Lee H, Bogyo M, Fellner M
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
UniProt ID: A0A0H2XFN6 (1-253) S-formylglutathione hydrolase
|
|
|
|
| Sample: |
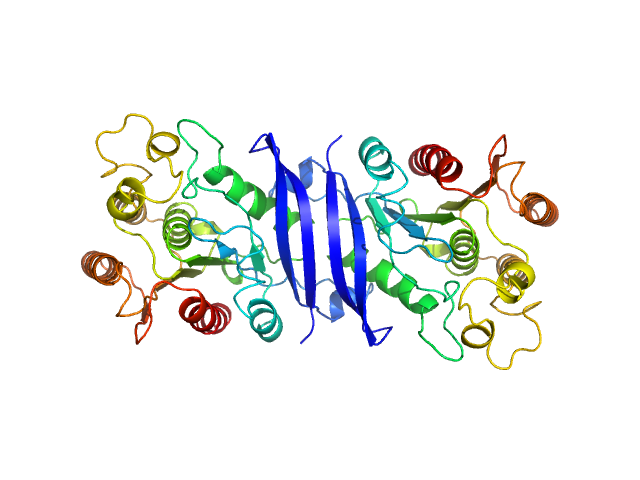
S-formylglutathione hydrolase dimer, 59 kDa Staphylococcus aureus (strain … protein
|
| Buffer: |
10 mM HEPES, 50 mM NaCl, 0.1% w/v NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2025 Mar 26
|
Unique structural and ligand-binding properties of the Staphylococcus aureus
serine hydrolase FphE
Proceedings of the National Academy of Sciences 123(13) (2026)
Jo J, Upadhyay T, You X, Bennett J, Lee H, Bogyo M, Fellner M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
99 |
nm3 |
|
|
UniProt ID: P33292 (1-576) Peroxisomal targeting signal receptor
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDX84_fit1_model1.png)
|
| Sample: |
Peroxisomal targeting signal receptor monomer, 65 kDa Komagataella pastoris protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 May 15
|
Structure of Pex8 in complex with peroxisomal receptor Pex5 reveals its essential role in peroxisomal cargo translocation
(2025)
Ekal L, Wendscheck D, David Y, Chojnowski G, Jeffries C, Mullapudi E, Schuldiner M, Warscheid B, Zalckvar E, Wilmanns M
|
| RgGuinier |
5.0 |
nm |
| Dmax |
21.4 |
nm |
| VolumePorod |
131 |
nm3 |
|
|
UniProt ID: Q01962 (33-713) Peroxisomal biogenesis factor 8
|
|
|
|
| Sample: |
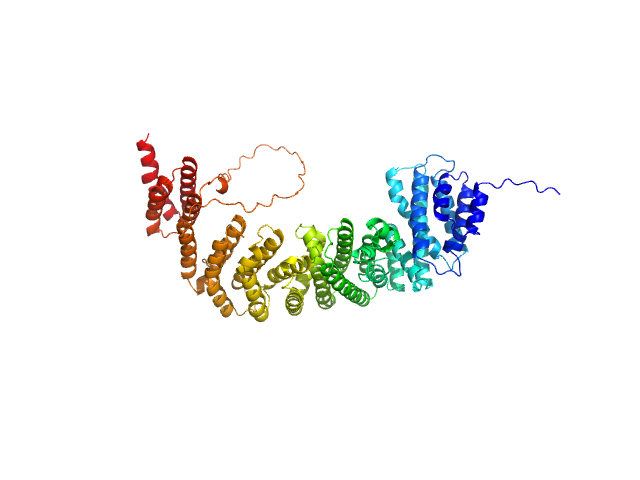
Peroxisomal biogenesis factor 8 monomer, 78 kDa Komagataella pastoris protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Aug 17
|
Structure of Pex8 in complex with peroxisomal receptor Pex5 reveals its essential role in peroxisomal cargo translocation
(2025)
Ekal L, Wendscheck D, David Y, Chojnowski G, Jeffries C, Mullapudi E, Schuldiner M, Warscheid B, Zalckvar E, Wilmanns M
|
| RgGuinier |
3.6 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
117 |
nm3 |
|
|
UniProt ID: P33292 (1-576) Peroxisomal targeting signal receptor
UniProt ID: Q01962 (33-713) Peroxisomal biogenesis factor 8
|
|
|
![OTHER [STATIC IMAGE] model](/media/pdb_file/SASDXA4_fit1_model1.png)
|
| Sample: |
Peroxisomal targeting signal receptor monomer, 65 kDa Komagataella pastoris protein
Peroxisomal biogenesis factor 8 monomer, 78 kDa Komagataella pastoris protein
|
| Buffer: |
50 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Aug 17
|
Structure of Pex8 in complex with peroxisomal receptor Pex5 reveals its essential role in peroxisomal cargo translocation
(2025)
Ekal L, Wendscheck D, David Y, Chojnowski G, Jeffries C, Mullapudi E, Schuldiner M, Warscheid B, Zalckvar E, Wilmanns M
|
| RgGuinier |
5.1 |
nm |
| Dmax |
23.5 |
nm |
| VolumePorod |
247 |
nm3 |
|
|