UniProt ID: P25440 (71-455) Bromodomain-containing protein 2
|
|
|
|
| Sample: |
Bromodomain-containing protein 2 monomer, 43 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Sep 25
|
Multivalent nucleosome scaffolding by bromodomain and extraterminal domain tandem bromodomains.
J Biol Chem :108289 (2025)
Olp MD, Bursch KL, Wynia-Smith SL, Nuñez R, Goetz CJ, Jackson V, Smith BC
|
| RgGuinier |
5.2 |
nm |
| Dmax |
18.3 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
UniProt ID: Q7DD94 (1-544) YhbX/YhjW/YijP/YjdB family protein (L152F)
|
|
|
|
| Sample: |
YhbX/YhjW/YijP/YjdB family protein (L152F) monomer, 62 kDa Neisseria meningitidis serogroup … protein
|
| Buffer: |
50 mM HEPES, 100 mM NaCl, 0.023% n-Dodecyl β-D-maltoside (DDM), pH: 7 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Nov 26
|
Conformational flexibility of EptA driven by an interdomain helix provides insights for enzyme-substrate recognition.
IUCrJ 8(Pt 5):732-746 (2021)
Anandan A, Dunstan NW, Ryan TM, Mertens HDT, Lim KYL, Evans GL, Kahler CM, Vrielink A
|
| RgGuinier |
4.2 |
nm |
| Dmax |
12.9 |
nm |
| VolumePorod |
304 |
nm3 |
|
|
UniProt ID: B7UM99 (1-233) Translocated intimin receptor Tir
|
|
|
|
| Sample: |
Translocated intimin receptor Tir dimer, 50 kDa Escherichia coli O127:H6 … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Feb 18
|
The pathogen-encoded signalling receptor Tir exploits host-like intrinsic disorder for infection.
Commun Biol 7(1):179 (2024)
Vieira MFM, Hernandez G, Zhong Q, Arbesú M, Veloso T, Gomes T, Martins ML, Monteiro H, Frazão C, Frankel G, Zanzoni A, Cordeiro TN
|
| RgGuinier |
3.5 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
UniProt ID: P23615 (None-None) Transcription elongation factor SPT6
|
|
|
|
| Sample: |
Transcription elongation factor SPT6 monomer, 145 kDa Saccharomyces cerevisiae (strain … protein
|
| Buffer: |
25 mM NaPi; 150mM NaCl; 0.5 mM EDTA; 5% glycerol; 1 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Oct 22
|
Cooperation between intrinsically disordered and ordered regions of Spt6 regulates nucleosome and Pol II CTD binding, and nucleosome assembly.
Nucleic Acids Res (2022)
Kasiliauskaite A, Kubicek K, Klumpler T, Zanova M, Zapletal D, Koutna E, Novacek J, Stefl R
|
| RgGuinier |
4.5 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
249 |
nm3 |
|
|
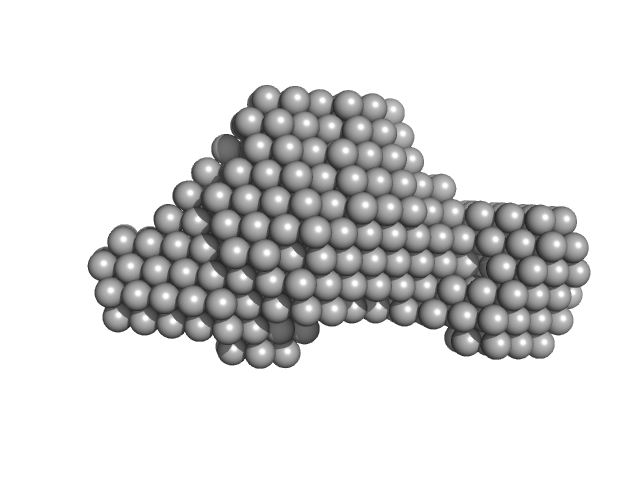
UniProt ID: E0IW31 (90-1520) Accessory colonization factor
|
|
|

|
| Sample: |
Accessory colonization factor monomer, 160 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM citrate-phosphate buffer, 200 mM NaCl, pH: 4.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 27
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari V, Garnett J
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.7 |
nm |
| VolumePorod |
248 |
nm3 |
|
|
UniProt ID: Q9GZZ9 (57-346) Ubiquitin-like modifier-activating enzyme 5
UniProt ID: P61960 (1-83) Ubiquitin fold modifer 1
|
|
|
|
| Sample: |
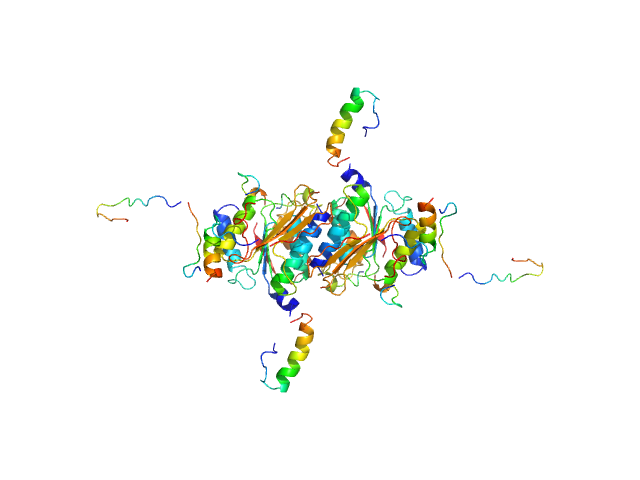
Ubiquitin-like modifier-activating enzyme 5 dimer, 68 kDa Homo sapiens protein
Ubiquitin fold modifer 1 monomer, 9 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, 2 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Oct 29
|
Structure and dynamics of UBA5-UFM1 complex formation showing new insights in the UBA5 activation mechanism
Journal of Structural Biology :107796 (2021)
Fuchs S, Kikhney A, Schubert R, Kaiser C, Liebau E, Svergun D, Betzel C, Perbandt M
|
| RgGuinier |
2.9 |
nm |
| Dmax |
11.0 |
nm |
|
|
UniProt ID: P40189 (22-324) Interleukin-6 receptor subunit beta
UniProt ID: P20809 (32-199) Interleukin-11
UniProt ID: Q14626 (23-319) Interleukin-11 receptor subunit alpha
|
|
|
|
| Sample: |
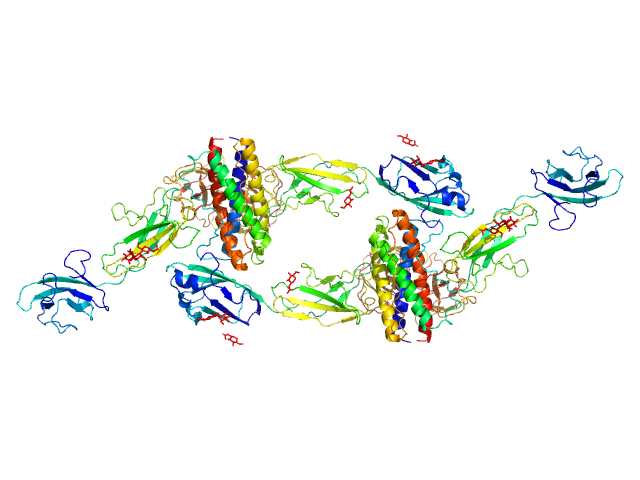
Interleukin-6 receptor subunit beta dimer, 69 kDa Homo sapiens protein
Interleukin-11 dimer, 36 kDa Homo sapiens protein
Interleukin-11 receptor subunit alpha dimer, 64 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.2% sodium azide, pH: 8.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2019 Jun 8
|
Structures of the interleukin 11 signalling complex reveal gp130 dynamics and the inhibitory mechanism of a cytokine variant
Nature Communications 14(1) (2023)
Metcalfe R, Hanssen E, Fung K, Aizel K, Kosasih C, Zlatic C, Doughty L, Morton C, Leis A, Parker M, Gooley P, Putoczki T, Griffin M
|
| RgGuinier |
5.3 |
nm |
| Dmax |
17.6 |
nm |
| VolumePorod |
405 |
nm3 |
|
|
UniProt ID: P02768 (None-None) Albumin
|
|
|
|
| Sample: |
Albumin monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Dec 1
|
Albumin in patients with liver disease shows an altered conformation.
Commun Biol 4(1):731 (2021)
Paar M, Fengler VH, Rosenberg DJ, Krebs A, Stauber RE, Oettl K, Hammel M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.5 |
nm |
|
|
UniProt ID: P0AEK2 (1-244) 3-oxoacyl-[acyl-carrier-protein] reductase FabG
|
|
|
|
| Sample: |
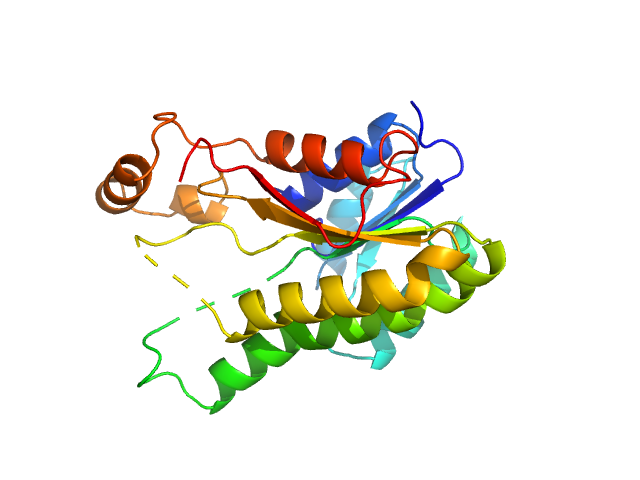
3-oxoacyl-[acyl-carrier-protein] reductase FabG, 26 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 1
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
3.6 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
212 |
nm3 |
|
|
UniProt ID: Q45589 (97-273) Cyclic di-AMP synthase CdaA
UniProt ID: O34824 (1-448) Phosphoglucosamine mutase
|
|
|
|
| Sample: |
Cyclic di-AMP synthase CdaA dimer, 44 kDa Bacillus subtilis (strain … protein
Phosphoglucosamine mutase dimer, 101 kDa Bacillus subtilis (strain … protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 21
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
4.5 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
250 |
nm3 |
|
|