UniProt ID: F4J9Q6 (1-136) Peptidyl-prolyl cis-trans isomerase FKBP43
|
|
|
|
| Sample: |
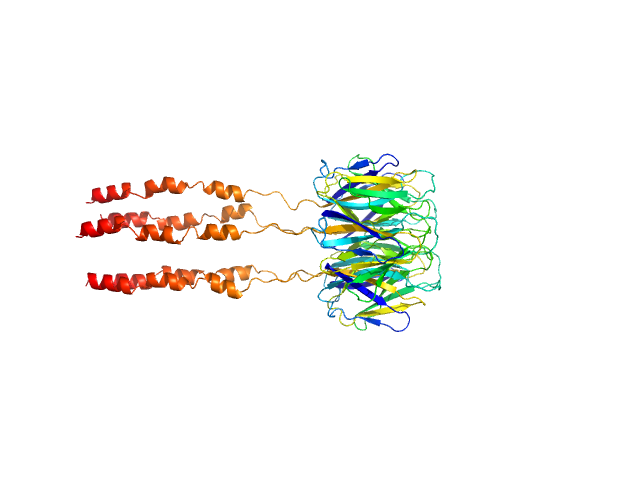
Peptidyl-prolyl cis-trans isomerase FKBP43 pentamer, 81 kDa Arabidopsis thaliana protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 28
|
The plant nucleoplasmin AtFKBP43 needs its extended arms for histone interaction.
Biochim Biophys Acta Gene Regul Mech 1865(7):194872 (2022)
Singh AK, Saharan K, Baral S, Vasudevan D
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
164 |
nm3 |
|
|
UniProt ID: Q5VST9 (1071-1254) Obscurin
|
|
|
|
| Sample: |
Obscurin monomer, 21 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, 0.35 mM NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 May 18
|
The N-terminus of obscurin is flexible in solution.
Proteins (2022)
Mauriello GE, Moncure GE, Nowzari RA, Miller CJ, Wright NT
|
| RgGuinier |
2.8 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
UniProt ID: Q63HQ2 (831-1017) Pikachurin
|
|
|

|
| Sample: |
Pikachurin monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Mar 3
|
Structure of the photoreceptor synaptic assembly of the extracellular matrix protein pikachurin with the orphan receptor GPR179
Science Signaling 16(795) (2023)
Patil D, Pantalone S, Cao Y, Laboute T, Novick S, Singh S, Savino S, Faravelli S, Magnani F, Griffin P, Singh A, Forneris F, Martemyanov K
|
| RgGuinier |
2.2 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
48 |
nm3 |
|
|
UniProt ID: Q9BZ95 (1070-1423) Histone-lysine N-methyltransferase NSD3
|
|
|
|
| Sample: |
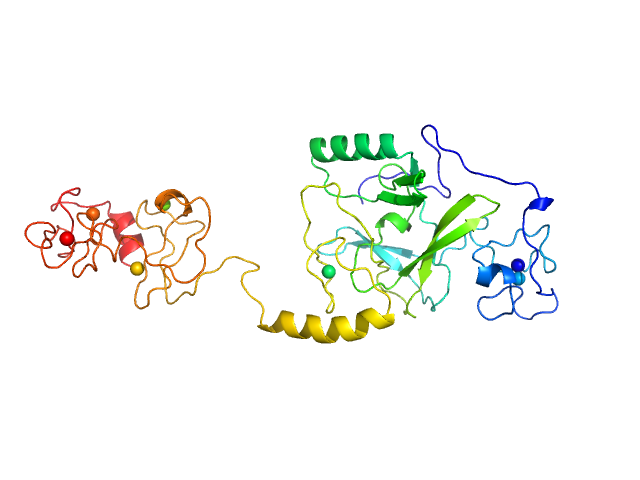
Histone-lysine N-methyltransferase NSD3 monomer, 40 kDa Homo sapiens protein
|
| Buffer: |
0.5 M NaCl, 20 mM Tris-HCl, 5 mM DTT, pH: 8.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Nov 25
|
Structural insights into the C-terminus of the histone-lysine N-methyltransferase NSD3 by small-angle X-ray scattering.
Front Mol Biosci 11:1191246 (2024)
Belviso BD, Shen Y, Carrozzini B, Morishita M, di Luccio E, Caliandro R
|
| RgGuinier |
3.4 |
nm |
| Dmax |
13.2 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
UniProt ID: P74102 (2-317) Orange carotenoid-binding protein
|
|
|

|
| Sample: |
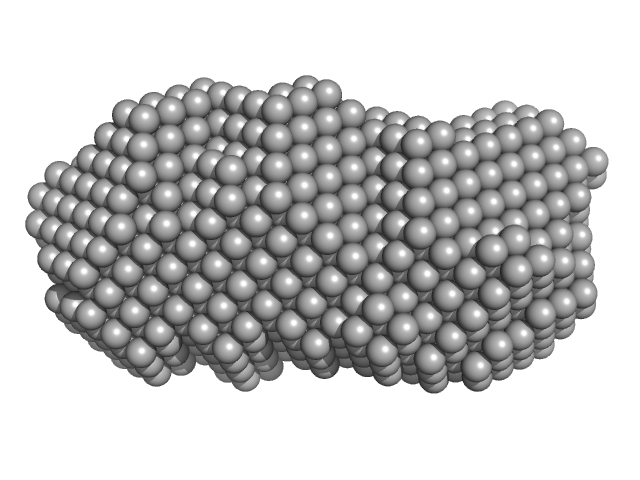
Orange carotenoid-binding protein dimer, 70 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Tris, 150 mM NaCL, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 23
|
Oligomerization processes limit photoactivation and recovery of the orange carotenoid protein.
Biophys J 121(15):2849-2872 (2022)
Andreeva EA, Niziński S, Wilson A, Levantino M, De Zitter E, Munro R, Muzzopappa F, Thureau A, Zala N, Burdzinski G, Sliwa M, Kirilovsky D, Schirò G, Colletier JP
|
| RgGuinier |
2.8 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: Q8U3D2 (5-770) Piwi domain-containing protein
|
|
|
|
| Sample: |
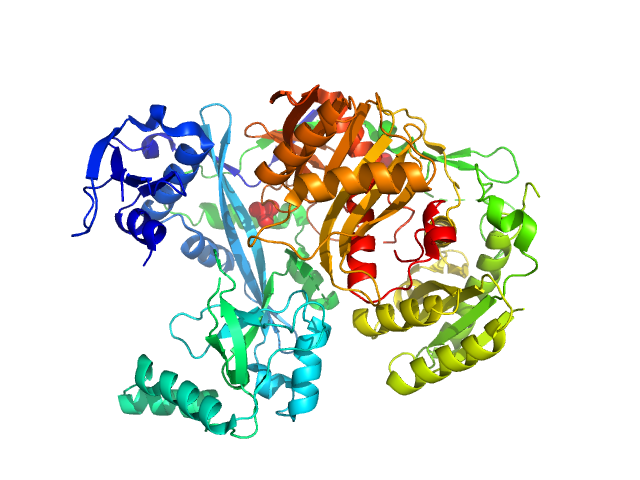
Piwi domain-containing protein monomer, 91 kDa Pyrococcus furiosus (strain … protein
|
| Buffer: |
20 mM Tris–HCl, 250 mM NaCl, 2mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2021 May 7
|
Argonaute protein SAXS investigation
lirong zheng
|
| RgGuinier |
2.9 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
142 |
nm3 |
|
|
UniProt ID: None (719-1706) Mono-acylated variant of a genetic fusion comprising the C-terminal folding scaffold of RTX block V (residues 1562–1681 of CyaA) and the CyaA residues 719–1294
|
|
|
|
| Sample: |
Mono-acylated variant of a genetic fusion comprising the C-terminal folding scaffold of RTX block V (residues 1562–1681 of CyaA) and the CyaA residues 719–1294 monomer, 78 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM NaCl, 3 mM CaCl2, 20 mM Tris, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2022 Feb 16
|
Acyl chains stabilize the acylated domain and determine the receptor-mediated interaction of the Bordetella adenylate cyclase toxin with cell membrane.
J Biol Chem :110392 (2025)
Espinosa-Vinals C, Stransky J, Osicka R, Osickova A, Jurnecka D, Sebo P, Bumba L
|
| RgGuinier |
5.0 |
nm |
| Dmax |
25.8 |
nm |
| VolumePorod |
178 |
nm3 |
|
|
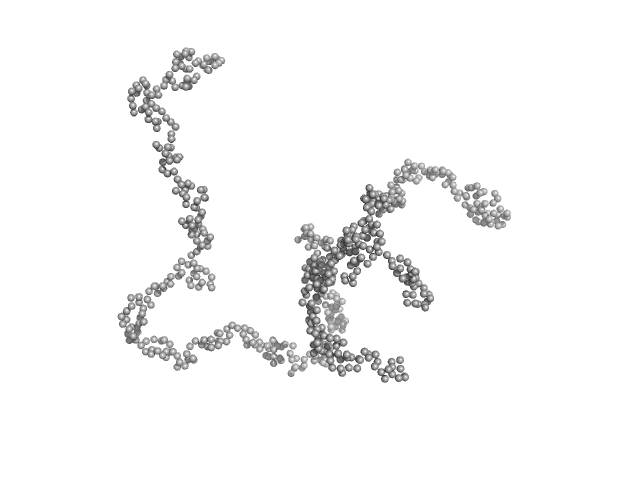
UniProt ID: P72509 (1-162) C-phycocyanin alpha subunit
UniProt ID: P72508 (1-172) C-phycocyanin beta subunit
|
|
|

|
| Sample: |
C-phycocyanin alpha subunit monomer, 18 kDa Arthrospira platensis protein
C-phycocyanin beta subunit monomer, 18 kDa Arthrospira platensis protein
|
| Buffer: |
8 M urea, 30 mM βME, pH: 7 |
| Experiment: |
SAXS
data collected at Rigaku MicroMax 007-HF, Moscow Institute of Physics and Technology (MIPT) on 2022 Apr 14
|
Anti-Stokes fluorescence excitation reveals conformational mobility of the C-phycocyanin chromophores
Structural Dynamics 9(5):054701 (2022)
Tsoraev G, Protasova E, Klimanova E, Ryzhykau Y, Kuklin A, Semenov Y, Ge B, Li W, Qin S, Friedrich T, Sluchanko N, Maksimov E
|
| RgGuinier |
6.6 |
nm |
| Dmax |
23.5 |
nm |
|
|
UniProt ID: P0DP23 (1-149) Calmodulin-1
|
|
|
|
| Sample: |
Calmodulin-1 monomer, 17 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES pH 7.5, 150 mM NaCl, 5 mM CaCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Jun 4
|
New insights into P2X7 receptor regulation: Ca2+-calmodulin and GDP bind to the soluble P2X7 ballast domain.
J Biol Chem :102495 (2022)
Sander S, Müller I, Alai MG, Nicke A, Tidow H
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.9 |
nm |
| VolumePorod |
23 |
nm3 |
|
|
UniProt ID: None (None-None) L-methionine gamma-lyase from Clostridium sporogenes fused with VGF S3 domain
|
|
|
|
| Sample: |
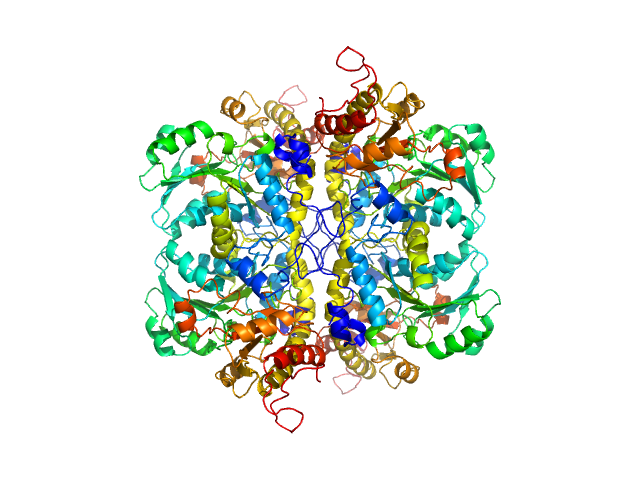
L-methionine gamma-lyase from Clostridium sporogenes fused with VGF S3 domain tetramer, 183 kDa Clostridium sporogenes protein
|
| Buffer: |
PBS-D2O: 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4 (D2O buffer), pH: 7.4 |
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 May 19
|
Methionine gamma lyase fused with S3 domain VGF forms octamers and adheres to tumor cells via binding EGFR
Biochemical and Biophysical Research Communications :149319 (2023)
Bondarev N, Bagaeva D, Bazhenov S, Buben M, Bulushova N, Ryzhykau Y, Okhrimenko I, Zagryadskaya Y, Maslov I, Anisimova N, Sokolova D, Kuklin A, Pokrovsky V, Manukhov I
|
| RgGuinier |
5.2 |
nm |
| Dmax |
17.3 |
nm |
| VolumePorod |
303 |
nm3 |
|
|