UniProt ID: O60885 (44-460) Bromodomain-containing protein 4
|
|
|
|
| Sample: |
Bromodomain-containing protein 4 monomer, 47 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 2% glycerol, 0.5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Jan 13
|
Interactome Rewiring Following Pharmacological Targeting of BET Bromodomains.
Mol Cell (2018)
Lambert JP, Picaud S, Fujisawa T, Hou H, Savitsky P, Uusküla-Reimand L, Gupta GD, Abdouni H, Lin ZY, Tucholska M, Knight JDR, Gonzalez-Badillo B, St-Denis N, Newman JA, Stucki M, Pelletier L, Bandeira N, Wilson MD, Filippakopoulos P, Gingras AC
|
| RgGuinier |
6.3 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
251 |
nm3 |
|
|
UniProt ID: P0A9Q8 (1-445) Aldehyde-alcohol dehydrogenase
|
|
|

|
| Sample: |
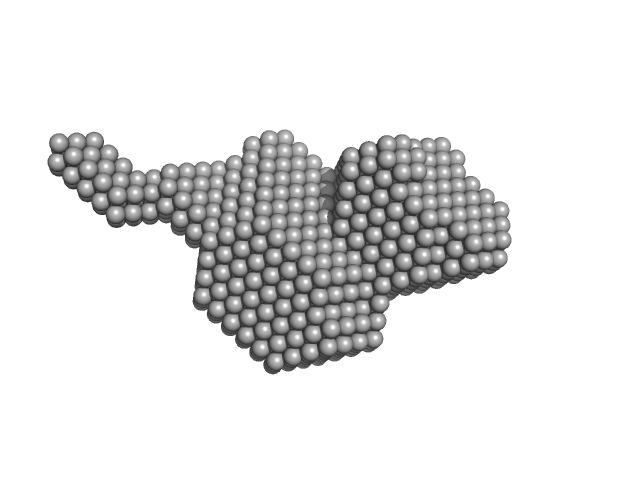
Aldehyde-alcohol dehydrogenase monomer, 48 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
30 mM HEPES, 150 mM NaCl, 5% (v/v) glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Jan 30
|
High-resolution structure of the alcohol dehydrogenase domain of the bifunctional bacterial enzyme AdhE.
Acta Crystallogr F Struct Biol Commun 76(Pt 9):414-421 (2020)
Azmi L, Bragginton EC, Cadby IT, Byron O, Roe AJ, Lovering AL, Gabrielsen M
|
| RgGuinier |
2.7 |
nm |
| Dmax |
11.1 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
UniProt ID: A0A0D7C2L1 (2-92) Plasmid stabilization protein ParE
UniProt ID: A0A0F6F6Q9 (13-63) Uncharacterized protein
|
|
|
|
| Sample: |
Plasmid stabilization protein ParE monomer, 13 kDa Escherichia coli protein
Uncharacterized protein monomer, 6 kDa Escherichia coli O157:H7 protein
|
| Buffer: |
50 mM Tris-HCl, 500 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2012 Feb 5
|
A unique hetero-hexadecameric architecture displayed by the Escherichia coli O157 PaaA2-ParE2 antitoxin-toxin complex.
J Mol Biol 428(8):1589-603 (2016)
Sterckx YG, Jové T, Shkumatov AV, Garcia-Pino A, Geerts L, De Kerpel M, Lah J, De Greve H, Van Melderen L, Loris R
|
| RgGuinier |
2.2 |
nm |
| Dmax |
9.3 |
nm |
| VolumePorod |
36 |
nm3 |
|
|
UniProt ID: Q9X721 (787-1118) Collagenase ColG segement s2s3as3b
|
|
|

|
| Sample: |
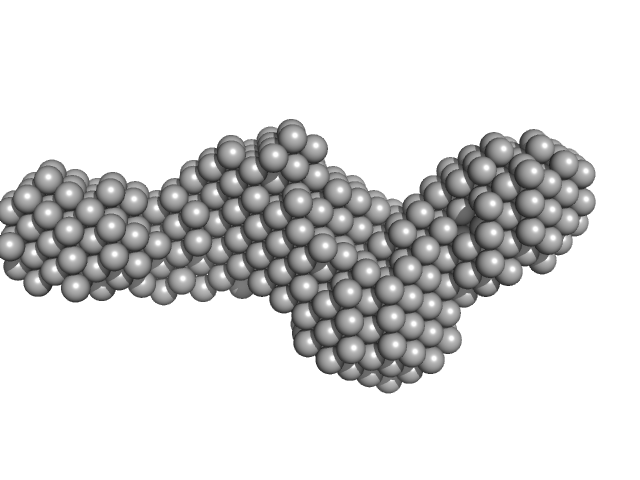
Collagenase ColG segement s2s3as3b monomer, 37 kDa Hathewaya histolytica protein
|
| Buffer: |
10mM HEPES 100mM NaCl 0.2mM EGTA, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Oct 12
|
Ca2+ - Induced Structural Change of Multi-Domain Collagen Binding Segments of Collagenases ColG and ColH from Hathewaya histolytica
University of Arkansas Dissertation - (2018)
Christopher E Ruth
|
| RgGuinier |
3.3 |
nm |
| Dmax |
11.5 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
UniProt ID: P9WNV1 (None-None) DNA ligase A
UniProt ID: None (None-None) Nicked DNA
|
|
|

|
| Sample: |
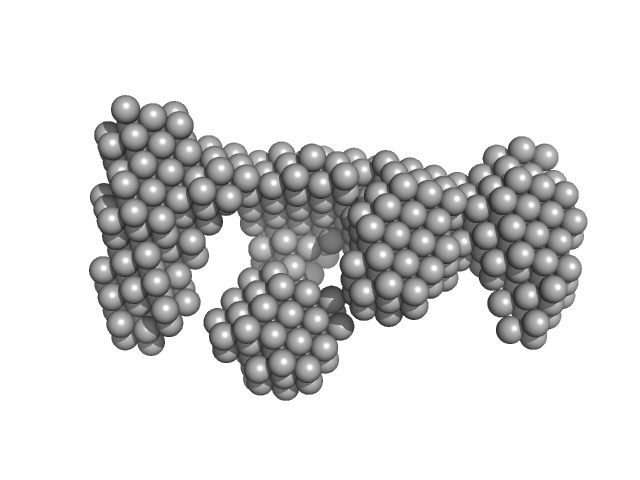
DNA ligase A monomer, 76 kDa Mycobacterium tuberculosis protein
Nicked DNA dimer, 16 kDa DNA
|
| Buffer: |
50 mM Tris-HCl, 200 mM NaCl, 2 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 May 13
|
M. tuberculosis class II apurinic/ apyrimidinic-endonuclease/3'-5' exonuclease (XthA) engages with NAD+-dependent DNA ligase A (LigA) to counter futile cleavage and ligation cycles in base excision repair.
Nucleic Acids Res (2020)
Khanam T, Afsar M, Shukla A, Alam F, Kumar S, Soyar H, Dolma K, Pasupuleti M, Srivastava KK, Ampapathi RS, Ramachandran R
|
| RgGuinier |
4.4 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
262 |
nm3 |
|
|
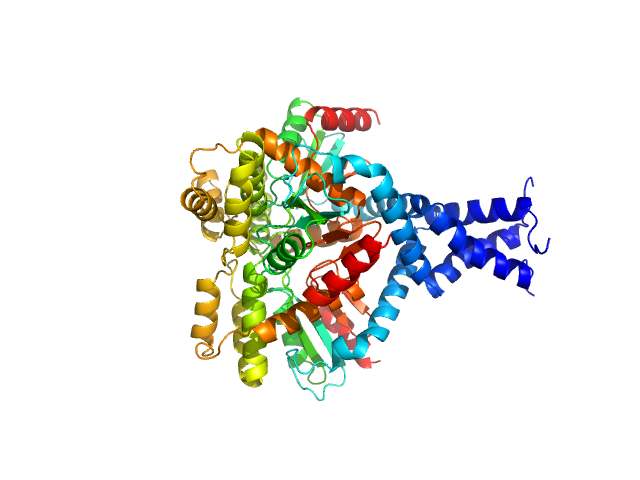
UniProt ID: P10408 (1-832) Protein translocase subunit SecA
|
|
|
|
| Sample: |
Protein translocase subunit SecA dimer, 189 kDa Escherichia coli protein
|
| Buffer: |
20mM HEPES, 100mM NaCl, 1mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Jul 18
|
The C-terminal tail of the bacterial translocation ATPase SecA modulates its activity.
Elife 8 (2019)
Jamshad M, Knowles TJ, White SA, Ward DG, Mohammed F, Rahman KF, Wynne M, Hughes GW, Kramer G, Bukau B, Huber D
|
| RgGuinier |
4.5 |
nm |
| Dmax |
15.7 |
nm |
| VolumePorod |
398 |
nm3 |
|
|
UniProt ID: Q9LM76 (None-None) Senescence-associated E3 ubiquitin ligase 1
|
|
|
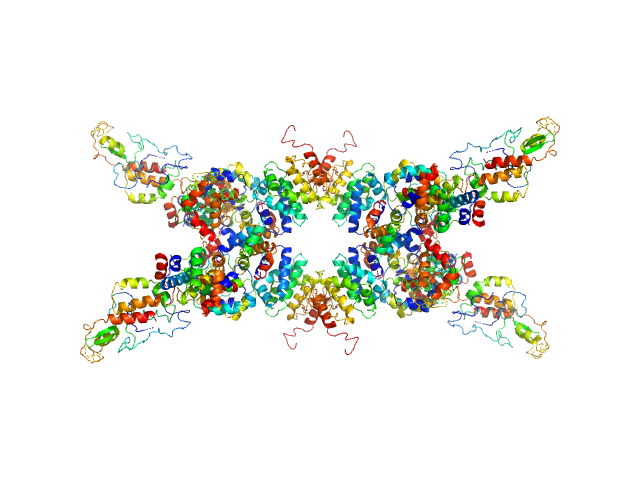
|
| Sample: |
Senescence-associated E3 ubiquitin ligase 1 tetramer, 355 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM Tris, 250 mM NaCl, pH: 9 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Jun 2
|
Senescence-associated ubiquitin ligase 1 (SAUL1)
Haifa El Kilani,
Al Kikhney
|
| RgGuinier |
5.9 |
nm |
| Dmax |
23.0 |
nm |
|
|
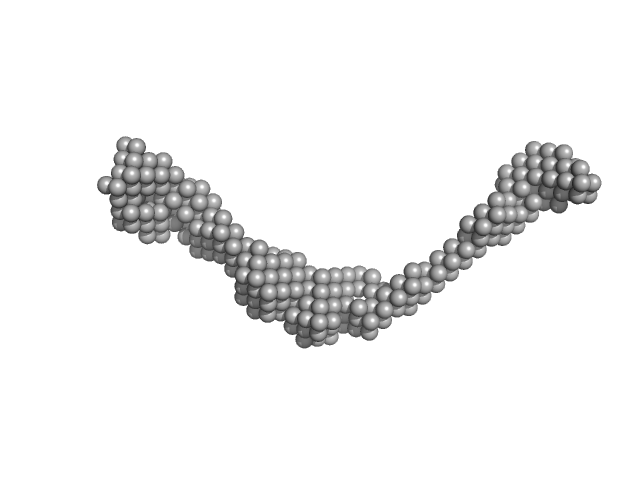
UniProt ID: Q911W0 (478-884) Tegument protein UL37
|
|
|
|
| Sample: |
Tegument protein UL37 monomer, 43 kDa Suid alphaherpesvirus 1 protein
|
| Buffer: |
100 mM HEPES 150 mM NaCl 5% glycerol 0.1 mM tris(2-carboxyethyl)phosphine (TCEP), pH: 7.5 |
| Experiment: |
SAXS
data collected at G1, Cornell High Energy Synchrotron Source (CHESS) on 2017 Jun 3
|
The dynamic nature of the conserved tegument protein UL37 of herpesviruses.
J Biol Chem 293(41):15827-15839 (2018)
Koenigsberg AL, Heldwein EE
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.5 |
nm |
|
|
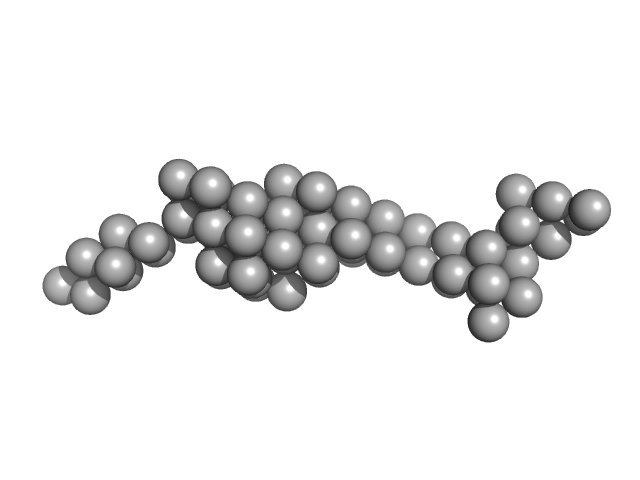
UniProt ID: A0A1Z1SYD5 (22-243) Deletion mutant of PmScsC
|
|
|
|
| Sample: |
Deletion mutant of PmScsC, 23 kDa Proteus mirabilis protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2018 Apr 5
|
Engineered variants provide new insight into the structural properties important for activity of the highly dynamic, trimeric protein disulfide isomerase ScsC from Proteus mirabilis.
Acta Crystallogr D Struct Biol 75(Pt 3):296-307 (2019)
Furlong EJ, Kurth F, Premkumar L, Whitten AE, Martin JL
|
| RgGuinier |
2.6 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
54 |
nm3 |
|
|
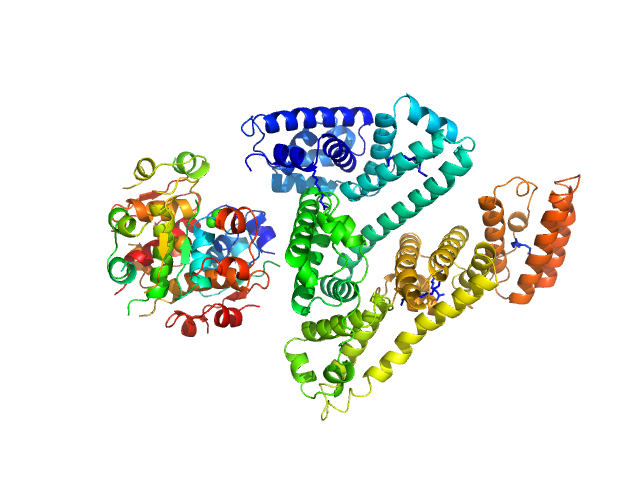
UniProt ID: P01308 (25-110) Insulin detemir (Levemir(R), Novo Nordisk A/S)
UniProt ID: P02768 (25-609) Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.)
|
|
|
|
| Sample: |
Insulin detemir (Levemir(R), Novo Nordisk A/S) hexamer, 35 kDa protein
Human Albumin (Recombumin(R) Alpha, Albumedix Ltd.) monomer, 66 kDa protein
|
| Buffer: |
6.9 mM Na2HPO4, 11.9 mM m-cresol, 13.7 mM phenol, 157.3 mM glycerol, 38.5 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at I911-4, MAX IV on 2015 Sep 25
|
Solution structures of long-acting insulin analogues and their complexes with albumin.
Acta Crystallogr D Struct Biol 75(Pt 3):272-282 (2019)
Ryberg LA, Sønderby P, Barrientos F, Bukrinski JT, Peters GHJ, Harris P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
162 |
nm3 |
|
|