UniProt ID: P53893 (1-1110) Mutated ribosome assembly protein 1 (R1086Q) with C-terminal tag (RSRSGSENLYFQGSHHHHHHHH)
|
|
|
|
| Sample: |
Mutated ribosome assembly protein 1 (R1086Q) with C-terminal tag (RSRSGSENLYFQGSHHHHHHHH) monomer, 127 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM HEPES, 300 mM NaCl, 5 mM MgCl2, 5% glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Oct 3
|
Hydroxyl radical footprinting modification reveals an intradomain communication pathway in EFL1 disrupted by a Shwachman-Diamond syndrome-associated mutation.
Protein Sci 35(4):e70504 (2026)
Zúñiga-Domínguez JA, Jain R, González-Andrade M, Farquhar ER, Chance MR, Gijsbers A, Sánchez-Puig N
|
| RgGuinier |
4.7 |
nm |
| Dmax |
15.2 |
nm |
| VolumePorod |
314 |
nm3 |
|
|
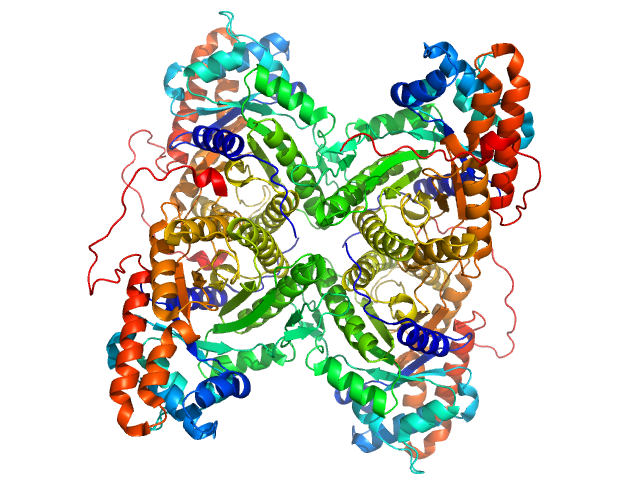
UniProt ID: P00883 (2-364) Fructose-bisphosphate aldolase A
|
|
|
|
| Sample: |
Fructose-bisphosphate aldolase A tetramer, 157 kDa Oryctolagus cuniculus protein
|
| Buffer: |
100 mM Tris/HCl 100 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2006 May 19
|
Accuracy of molecular mass determination of proteins in solution by small-angle X-ray scattering
Journal of Applied Crystallography 40(s1):s245-s249 (2007)
Mylonas E, Svergun D
|
| RgGuinier |
3.6 |
nm |
| Dmax |
10.5 |
nm |
|
|
UniProt ID: M4N8T6 (None-None) S-layer protein
|
|
|

|
| Sample: |
S-layer protein monomer, 116 kDa Lysinibacillus sphaericus protein
|
| Buffer: |
Water with Mg2+, pH: |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Jun 2
|
Analysis of self-assembly of S-layer protein slp-B53 from Lysinibacillus sphaericus.
Eur Biophys J 46(1):77-89 (2017)
Liu J, Falke S, Drobot B, Oberthuer D, Kikhney A, Guenther T, Fahmy K, Svergun D, Betzel C, Raff J
|
| RgGuinier |
6.6 |
nm |
| Dmax |
29.0 |
nm |
| VolumePorod |
597 |
nm3 |
|
|
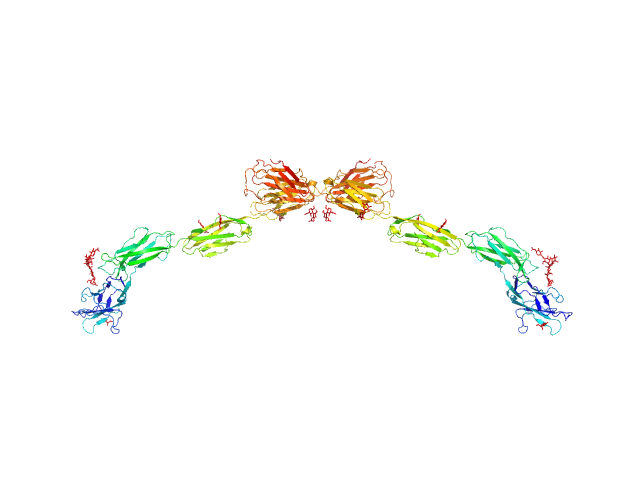
UniProt ID: P20917 (20-508) Myelin-associated glycoprotein (20-508; N406Q mutant)
|
|
|
|
| Sample: |
Myelin-associated glycoprotein (20-508; N406Q mutant) monomer, 54 kDa Mus musculus protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 5
|
Structural basis of myelin-associated glycoprotein adhesion and signalling.
Nat Commun 7:13584 (2016)
Pronker MF, Lemstra S, Snijder J, Heck AJ, Thies-Weesie DM, Pasterkamp RJ, Janssen BJ
|
| RgGuinier |
7.3 |
nm |
| Dmax |
25.6 |
nm |
| VolumePorod |
193 |
nm3 |
|
|
UniProt ID: Q15185 (1-117) Prostaglandin E synthase 3 (1-117)
|
|
|
|
| Sample: |
Prostaglandin E synthase 3 (1-117) monomer, 14 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris-HCl, 100 mM NaCl, 5 mM B-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS1 Beamline, Brazilian Synchrotron Light Laboratory on 2013 Jun 21
|
The C-terminal region of the human p23 chaperone modulates its structure and function.
Arch Biochem Biophys 565:57-67 (2015)
Seraphim TV, Gava LM, Mokry DZ, Cagliari TC, Barbosa LR, Ramos CH, Borges JC
|
| RgGuinier |
1.9 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
UniProt ID: P31483-2 (93-274) Nucleolysin TIA-1 isoform p40
UniProt ID: (None-None) DNA (TTTTTACTCC)
|
|
|
|
| Sample: |
Nucleolysin TIA-1 isoform p40 monomer, 21 kDa Homo sapiens protein
DNA (TTTTTACTCC) monomer, 1 kDa DNA
|
| Buffer: |
20 mM HEPES, 100 mM NaCl, 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2016 May 27
|
TIA-1 RRM23 binding and recognition of target oligonucleotides.
Nucleic Acids Res 45(8):4944-4957 (2017)
Waris S, García-Mauriño SM, Sivakumaran A, Beckham SA, Loughlin FE, Gorospe M, Díaz-Moreno I, Wilce MCJ, Wilce JA
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.1 |
nm |
| VolumePorod |
32 |
nm3 |
|
|
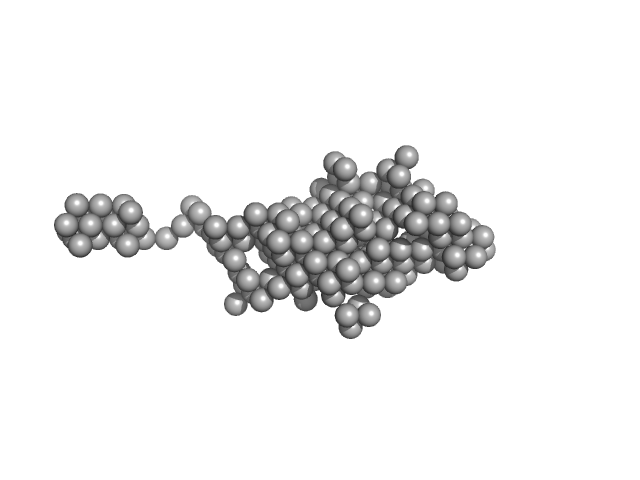
UniProt ID: Q96FI4 (None-None) Endonuclease 8-like 1
UniProt ID: (None-None) dsDNA
|
|
|
|
| Sample: |
Endonuclease 8-like 1 monomer, 45 kDa Homo sapiens protein
DsDNA monomer, 2 kDa DNA
|
| Buffer: |
25mM HEPES 100mM NaCl 1mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Jan 20
|
Destabilization of the PCNA trimer mediated by its interaction with the NEIL1 DNA glycosylase.
Nucleic Acids Res 45(5):2897-2909 (2017)
Prakash A, Moharana K, Wallace SS, Doublié S
|
| RgGuinier |
3.3 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
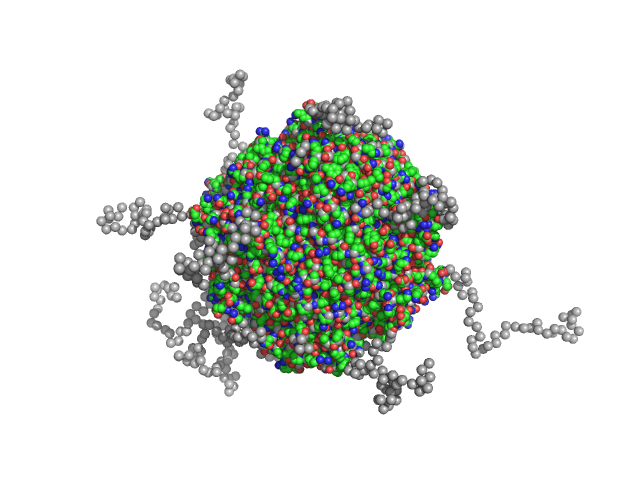
UniProt ID: Q9RZN1 (31-241) N-terminal truncated DNA protection during starvation protein 2
|
|
|
|
| Sample: |
N-terminal truncated DNA protection during starvation protein 2 dodecamer, 279 kDa Deinococcus radiodurans R1 protein
|
| Buffer: |
20 mM Tris-HCl, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2013 Nov 22
|
SAXS Structural Studies of Dps from Deinococcus radiodurans Highlights the Conformation of the Mobile N-Terminal Extensions.
J Mol Biol 429(5):667-687 (2017)
Santos SP, Cuypers MG, Round A, Finet S, Narayanan T, Mitchell EP, Romão CV
|
| RgGuinier |
4.2 |
nm |
| Dmax |
12.7 |
nm |
| VolumePorod |
445 |
nm3 |
|
|
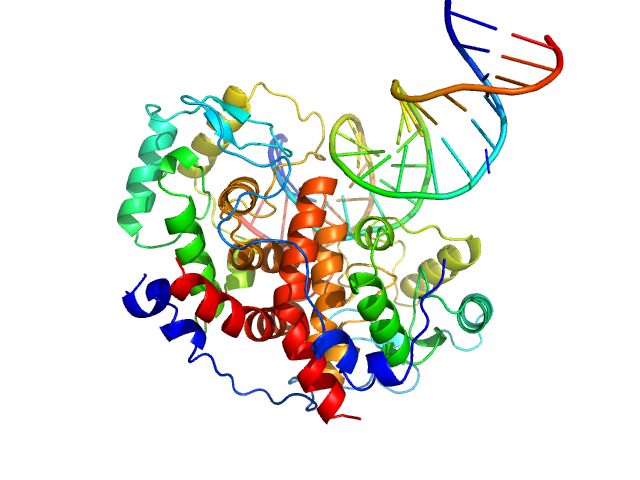
UniProt ID: P08392 (258-487) Major viral transcription factor ICP4
UniProt ID: (None-None) Immediate Early 3
|
|
|
|
| Sample: |
Major viral transcription factor ICP4 dimer, 49 kDa Human alphaherpesvirus 1 protein
Immediate Early 3 dimer, 12 kDa DNA
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2015 Jul 27
|
The herpes viral transcription factor ICP4 forms a novel DNA recognition complex.
Nucleic Acids Res 45(13):8064-8078 (2017)
Tunnicliffe RB, Lockhart-Cairns MP, Levy C, Mould AP, Jowitt TA, Sito H, Baldock C, Sandri-Goldin RM, Golovanov AP
|
| RgGuinier |
2.5 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
90 |
nm3 |
|
|
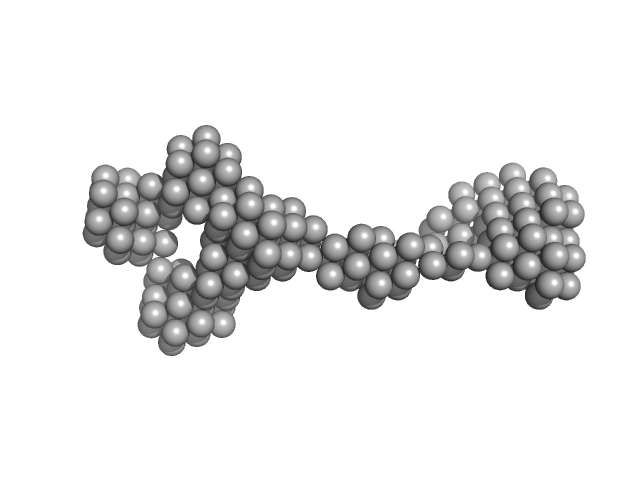
UniProt ID: P39685 (None-None) Nucleoporin POM152
|
|
|
|
| Sample: |
Nucleoporin POM152 monomer, 12 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
10mM HEPES, 150mM NaCl, 10%(v/v) glycerol, 5mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2015 Apr 12
|
Molecular Architecture of the Major Membrane Ring Component of the Nuclear Pore Complex.
Structure 25(3):434-445 (2017)
Upla P, Kim SJ, Sampathkumar P, Dutta K, Cahill SM, Chemmama IE, Williams R, Bonanno JB, Rice WJ, Stokes DL, Cowburn D, Almo SC, Sali A, Rout MP, Fernandez-Martinez J
|
| RgGuinier |
2.8 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
57 |
nm3 |
|
|