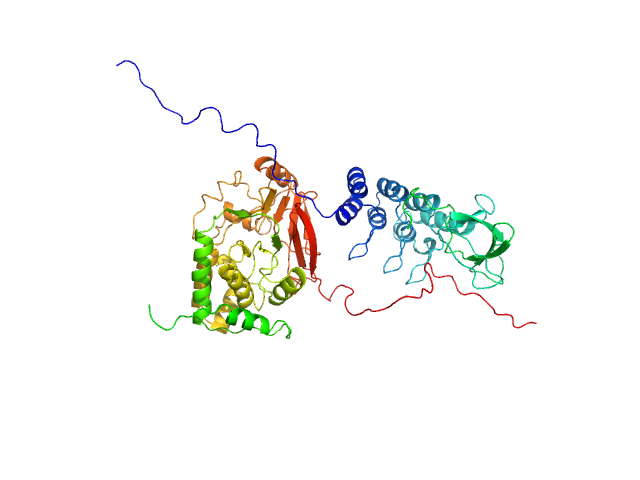
UniProt ID: P62136 (1-330) Serine/threonine-protein phosphatase PP1-alpha catalytic subunit
UniProt ID: Q8WUF5 (621-828) Inhibitor of apoptosis-stimulating protein of p53 (RelA-associated inhibitor)
|
|
|
|
| Sample: |
Serine/threonine-protein phosphatase PP1-alpha catalytic subunit monomer, 38 kDa Homo sapiens protein
Inhibitor of apoptosis-stimulating protein of p53 (RelA-associated inhibitor) monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2019 Apr 25
|
Flexible Tethering of ASPP Proteins Facilitates PP-1c Catalysis.
Structure 27(10):1485-1496.e4 (2019)
Zhou Y, Millott R, Kim HJ, Peng S, Edwards RA, Skene-Arnold T, Hammel M, Lees-Miller SP, Tainer JA, Holmes CFB, Glover JNM
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.9 |
nm |
| VolumePorod |
116 |
nm3 |
|
|
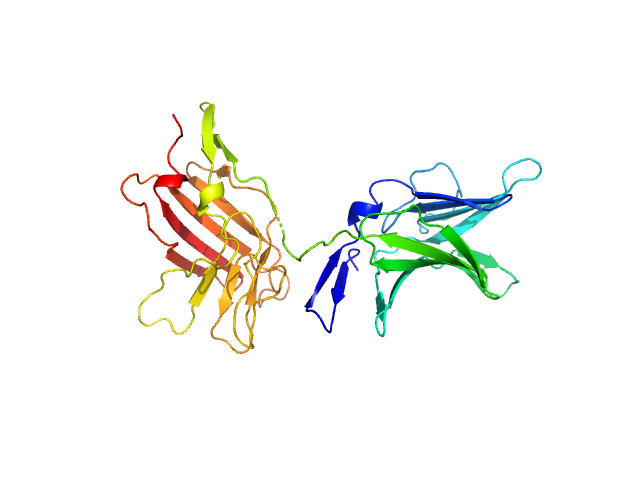
UniProt ID: P11717 (1222-1510) Cation-independent mannose-6-phosphate receptor
|
|
|
|
| Sample: |
Cation-independent mannose-6-phosphate receptor monomer, 34 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2020 Mar 3
|
Structure of the Human Cation-Independent Mannose 6-Phosphate/IGF2 Receptor Domains 7–11 Uncovers the Mannose 6-Phosphate Binding Site of Domain 9
Structure (2020)
Bochel A, Williams C, McCoy A, Hoppe H, Winter A, Nicholls R, Harlos K, Jones E, Berger I, Hassan A, Crump M
|
| RgGuinier |
2.5 |
nm |
| Dmax |
7.8 |
nm |
| VolumePorod |
48 |
nm3 |
|
|
UniProt ID: Q8TCU6 (38-499) Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1
|
|
|
|
| Sample: |
Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 monomer, 54 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, pH: 7 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Oct 19
|
The first DEP domain of the RhoGEF P-Rex1 autoinhibits activity andcontributes to membrane binding.
J Biol Chem (2020)
Ravala SK, Hopkins JB, Plescia CB, Allgood SR, Kane MA, Cash JN, Stahelin RV, Tesmer JJG
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
75 |
nm3 |
|
|
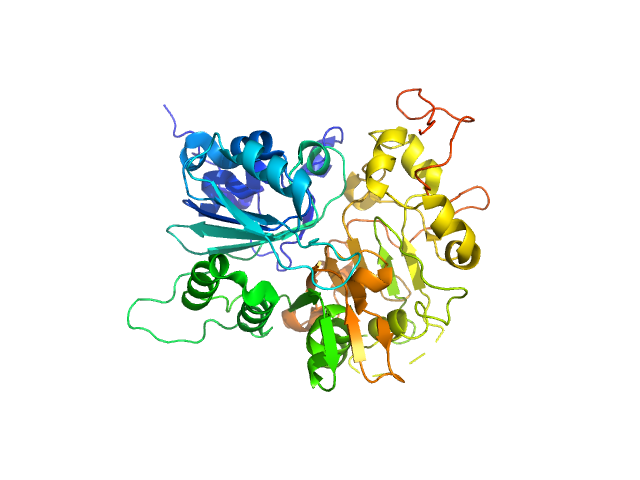
UniProt ID: Q9NUW8 (149-698) Tyrosyl-DNA phosphodiesterase 1 (149-608)
|
|
|
|
| Sample: |
Tyrosyl-DNA phosphodiesterase 1 (149-608) monomer, 53 kDa Homo sapiens protein
|
| Buffer: |
200 mM NaCl, 20 mM Tris-HCl, pH 7.5, 2% glycerol,, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Sep 18
|
Direct interaction of DNA repair protein tyrosyl DNA phosphodiesterase 1 and the DNA ligase III catalytic domain is regulated by phosphorylation of its flexible N-terminus.
J Biol Chem :100921 (2021)
Rashid I, Hammel M, Sverzhinsky A, Tsai MS, Pascal JM, Tainer JA, Tomkinson AE
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
80 |
nm3 |
|
|
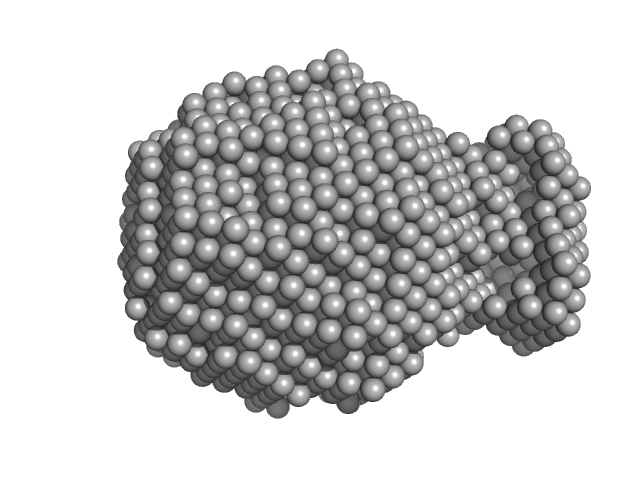
UniProt ID: P12111 (1634-1833) Collagen alpha-3(VI) chain, N2 domain
|
|
|

|
| Sample: |
Collagen alpha-3(VI) chain, N2 domain monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM TRIS, pH 7.4, 150mM NaCl 3% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 4
|
Structure of a collagen VI α3 chain VWA domain array: adaptability and functional implications of myopathy causing mutations
Journal of Biological Chemistry :jbc.RA120.014865 (2020)
Solomon-Degefa H, Gebauer J, Jeffries C, Freiburg C, Meckelburg P, Bird L, Baumann U, Svergun D, Owens R, Werner J, Behrmann E, Paulsson M, Wagener R
|
| RgGuinier |
1.8 |
nm |
| Dmax |
5.8 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
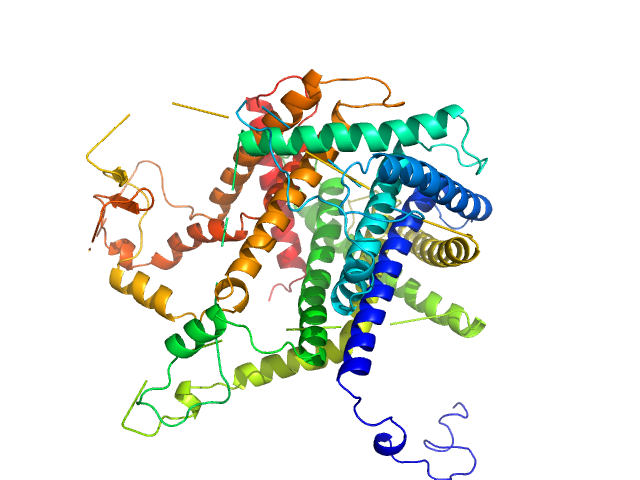
UniProt ID: P57744 (1-87) Mitochondrial import inner membrane translocase subunit TIM8
UniProt ID: P53299 (2-105) Mitochondrial import inner membrane translocase subunit TIM13
|
|
|
|
| Sample: |
Mitochondrial import inner membrane translocase subunit TIM8 trimer, 29 kDa Saccharomyces cerevisiae protein
Mitochondrial import inner membrane translocase subunit TIM13 trimer, 34 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
50 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Apr 21
|
Structural basis of client specificity in mitochondrial membrane-protein chaperones.
Sci Adv 6(51) (2020)
Sučec I, Wang Y, Dakhlaoui O, Weinhäupl K, Jores T, Costa D, Hessel A, Brennich M, Rapaport D, Lindorff-Larsen K, Bersch B, Schanda P
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.9 |
nm |
| VolumePorod |
136 |
nm3 |
|
|
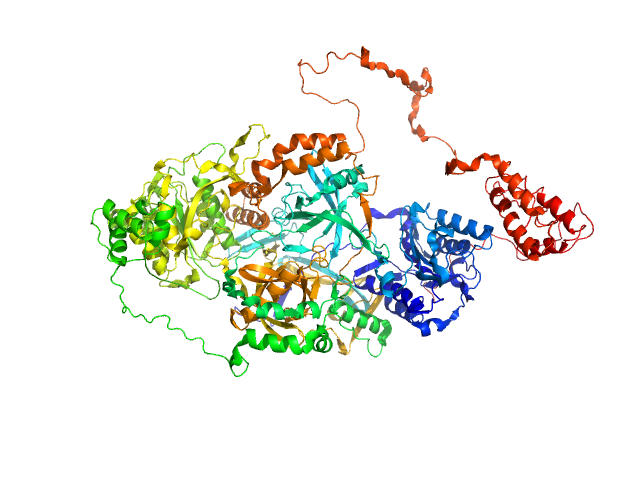
UniProt ID: P12956 (1-609) X-ray repair cross-complementing protein 6
UniProt ID: P13010 (1-732) X-ray repair cross-complementing protein 5
UniProt ID: None (None-None) Y-DNA
|
|
|
|
| Sample: |
X-ray repair cross-complementing protein 6 monomer, 70 kDa Homo sapiens protein
X-ray repair cross-complementing protein 5 monomer, 83 kDa Homo sapiens protein
Y-DNA monomer, 18 kDa DNA
|
| Buffer: |
50 mM Tris-HCl, 100 mM NaCl, 5% glycerol, 0.01% sodium azide, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2010 Jan 8
|
Visualizing functional dynamicity in the DNA-dependent protein kinase holoenzyme DNA-PK complex by integrating SAXS with cryo-EM.
Prog Biophys Mol Biol (2020)
Hammel M, Rosenberg DJ, Bierma J, Hura GL, Lees-Miller SP, Tainer JA
|
| RgGuinier |
4.1 |
nm |
| Dmax |
14.8 |
nm |
| VolumePorod |
280 |
nm3 |
|
|
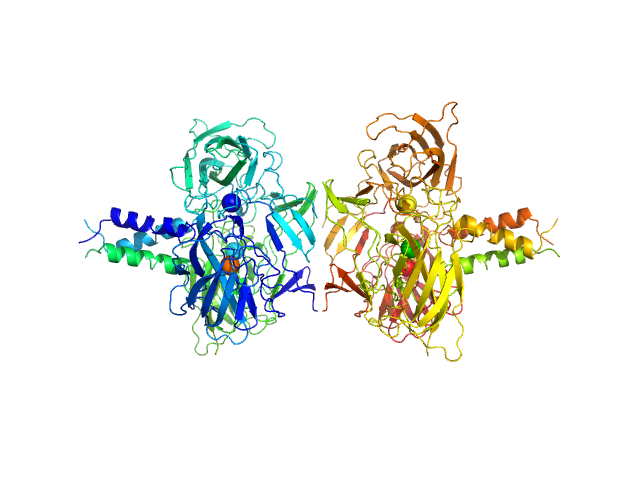
UniProt ID: O94901 (616-812) SUN domain-containing protein 1
UniProt ID: Q8N6L0 (542-562) Protein KASH5
|
|
|
|
| Sample: |
SUN domain-containing protein 1 hexamer, 135 kDa Homo sapiens protein
Protein KASH5 hexamer, 20 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8.0, 150 mM KCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Dec 17
|
A molecular mechanism for LINC complex branching by structurally diverse SUN-KASH 6:6 assemblies.
Elife 10 (2021)
Gurusaran M, Davies OR
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
244 |
nm3 |
|
|
UniProt ID: Q9I589 (26-389) Chitin-binding protein CbpD
|
|
|
|
| Sample: |
Chitin-binding protein CbpD monomer, 39 kDa Pseudomonas aeruginosa protein
|
| Buffer: |
15 mM Tris-HCl 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at Bruker Nanostar with InCoatec Cu microsource, RECX, University of Oslo on 2019 Aug 14
|
The lytic polysaccharide monooxygenase CbpD promotes Pseudomonas aeruginosa virulence in systemic infection.
Nat Commun 12(1):1230 (2021)
Askarian F, Uchiyama S, Masson H, Sørensen HV, Golten O, Bunæs AC, Mekasha S, Røhr ÅK, Kommedal E, Ludviksen JA, Arntzen MØ, Schmidt B, Zurich RH, van Sorge NM, Eijsink VGH, Krengel U, Mollnes TE, Lewis NE, Nizet V, Vaaje-Kolstad G
|
| RgGuinier |
3.5 |
nm |
| Dmax |
15.0 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
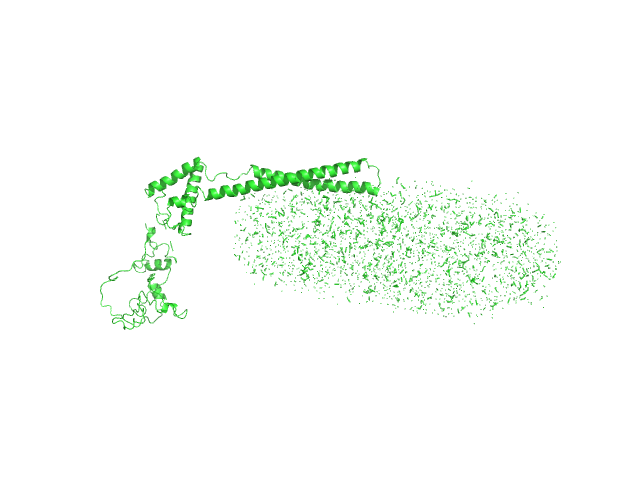
UniProt ID: Q79FZ9 (126-454) Mce-family protein Mce1A
UniProt ID: None (None-None) n-Dodecyl-β-D-Maltopyranoside
|
|
|

|
| Sample: |
Mce-family protein Mce1A monomer, 39 kDa Mycobacterium tuberculosis protein
N-Dodecyl-β-D-Maltopyranoside 0, 123 kDa
|
| Buffer: |
50 mM Tris, 350 mM NaCl, 10% Glycerol, 5mM DDM, 1 mM β-ME, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 Nov 28
|
Structural insights into the substrate-binding proteins Mce1A and Mce4A from Mycobacterium tuberculosis
IUCrJ 8(5) (2021)
Asthana P, Singh D, Pedersen J, Hynönen M, Sulu R, Murthy A, Laitaoja M, Jänis J, Riley L, Venkatesan R
|
| RgGuinier |
5.5 |
nm |
| Dmax |
22.0 |
nm |
| VolumePorod |
216 |
nm3 |
|
|