UniProt ID: B1H241 (1-452) Resistance to inhibitors of cholinesterase 8 homolog A
|
|
|
|
| Sample: |
Resistance to inhibitors of cholinesterase 8 homolog A monomer, 51 kDa Rattus norvegicus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2018 Apr 24
|
Structure, Function, and Dynamics of the Gα Binding Domain of Ric-8A.
Structure (2019)
Zeng B, Mou TC, Doukov TI, Steiner A, Yu W, Papasergi-Scott M, Tall GG, Hagn F, Sprang SR
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
70 |
nm3 |
|
|
UniProt ID: Q9SU72 (None-None) Enhanced disease susceptibility
|
|
|
|
| Sample: |
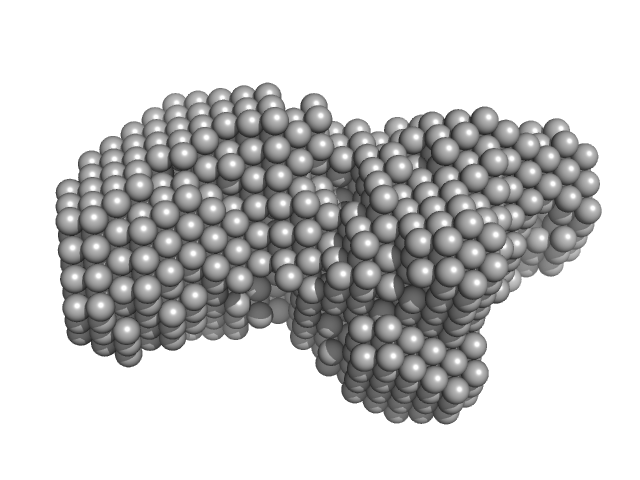
Enhanced disease susceptibility monomer, 72 kDa Arabidopsis thaliana protein
|
| Buffer: |
50 mM NaCl, 50 mM HEPES, 1% glyercol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2015 May 19
|
Arabidopsis immunity regulator EDS1 in a PAD4/SAG101-unbound form is a monomer with an inherently inactive conformation.
J Struct Biol :107390 (2019)
Voss M, Toelzer C, Bhandari DD, Parker JE, Niefind K
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
UniProt ID: P0ACT4 (2-208) Tetracycline repressor (class D)
|
|
|
|
| Sample: |
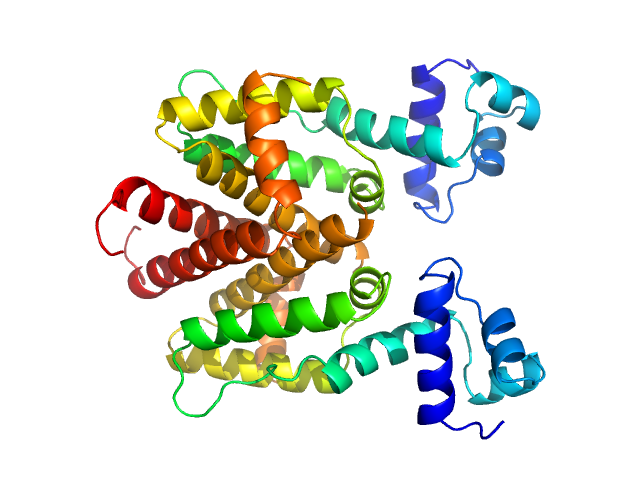
Tetracycline repressor (class D) dimer, 47 kDa Escherichia coli protein
|
| Buffer: |
50 mM Tris/HCl 150 mM NaCl 10 mM MgCl2, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Sep 23
|
Thermodynamics, cooperativity and stability of the tetracycline repressor (TetR) upon tetracycline binding.
Biochim Biophys Acta Proteins Proteom :140404 (2020)
Palm GJ, Buchholz I, Werten S, Girbardt B, Berndt L, Delcea M, Hinrichs W
|
| RgGuinier |
2.6 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
UniProt ID: P0ACT4 (2-208) Tetracycline repressor (class D)
UniProt ID: None (None-None) 5a,6-anhydrotetracycline
|
|
|
|
| Sample: |
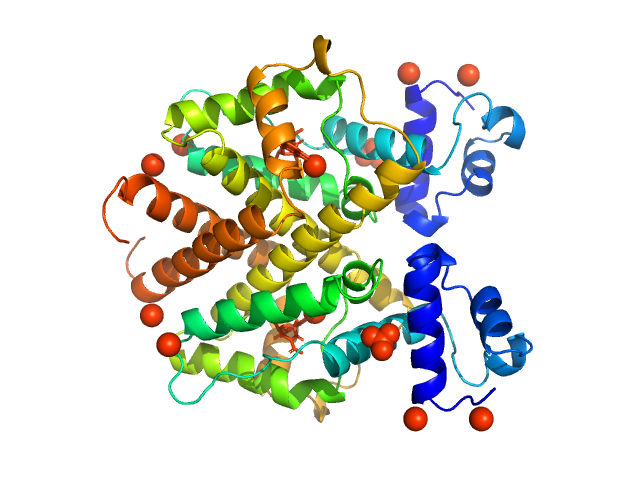
Tetracycline repressor (class D) dimer, 47 kDa Escherichia coli protein
5a,6-anhydrotetracycline dimer, 1 kDa
|
| Buffer: |
50 mM Tris/HCl 150 mM NaCl 10 mM MgCl2, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Sep 23
|
Thermodynamics, cooperativity and stability of the tetracycline repressor (TetR) upon tetracycline binding.
Biochim Biophys Acta Proteins Proteom :140404 (2020)
Palm GJ, Buchholz I, Werten S, Girbardt B, Berndt L, Delcea M, Hinrichs W
|
| RgGuinier |
2.6 |
nm |
| Dmax |
6.8 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: Q8ILC1 (None-None) STI1-like protein
|
|
|

|
| Sample: |
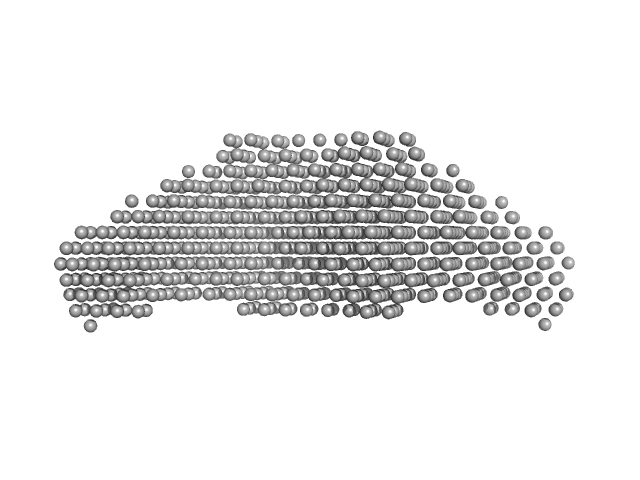
STI1-like protein dimer, 136 kDa Plasmodium falciparum protein
|
| Buffer: |
25 mM Tris-HCl, 100 mM NaCl, 1 mM EDTA, 1 mM β-mercaptoethanol, pH: 8 |
| Experiment: |
SAXS
data collected at SAXS1 Beamline, Brazilian Synchrotron Light Laboratory on 2016 Aug 3
|
Structural studies of the Hsp70/Hsp90 organizing protein of Plasmodium falciparum and its modulation of Hsp70 and Hsp90 ATPase activities.
Biochim Biophys Acta Proteins Proteom :140282 (2019)
Silva NSM, Bertolino-Reis DE, Dores-Silva PR, Anneta FB, Seraphim TV, Barbosa LRS, Borges JC
|
| RgGuinier |
6.3 |
nm |
| Dmax |
24.0 |
nm |
| VolumePorod |
557 |
nm3 |
|
|
UniProt ID: Q83V25 (1-334) 2-amino-3-carboxymuconate 6-semialdehyde decarboxylase
|
|
|
|
| Sample: |
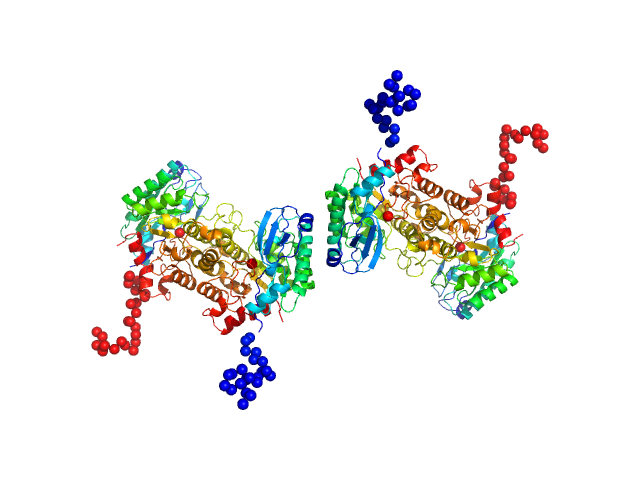
2-amino-3-carboxymuconate 6-semialdehyde decarboxylase tetramer, 159 kDa Pseudomonas fluorescens protein
|
| Buffer: |
50 mM Tris, 5 mM DTT, pH: 8.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2018 Jul 15
|
Quaternary structure of α-amino-β-carboxymuconate-ϵ-semialdehyde decarboxylase (ACMSD) controls its activity.
J Biol Chem 294(30):11609-11621 (2019)
Yang Y, Davis I, Matsui T, Rubalcava I, Liu A
|
| RgGuinier |
5.2 |
nm |
| Dmax |
19.0 |
nm |
| VolumePorod |
238 |
nm3 |
|
|
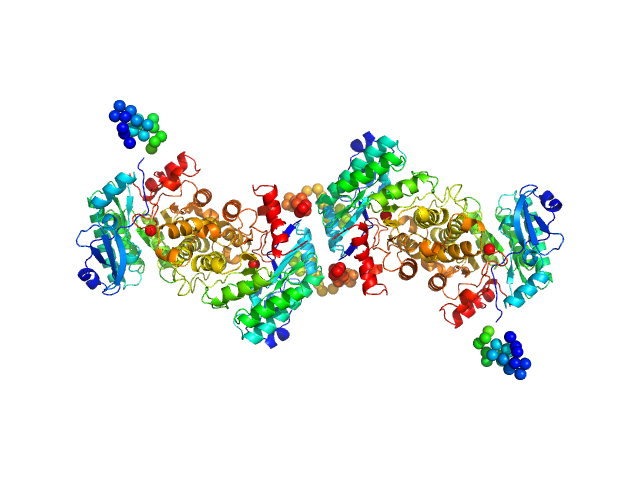
UniProt ID: Q83V25 (1-334) 2-amino-3-carboxymuconate 6-semialdehyde decarboxylase
|
|
|
|
| Sample: |
2-amino-3-carboxymuconate 6-semialdehyde decarboxylase tetramer, 159 kDa Pseudomonas fluorescens protein
|
| Buffer: |
25 mM HEPES, 5 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2018 Jan 10
|
Quaternary structure of α-amino-β-carboxymuconate-ϵ-semialdehyde decarboxylase (ACMSD) controls its activity.
J Biol Chem 294(30):11609-11621 (2019)
Yang Y, Davis I, Matsui T, Rubalcava I, Liu A
|
| RgGuinier |
4.7 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
195 |
nm3 |
|
|
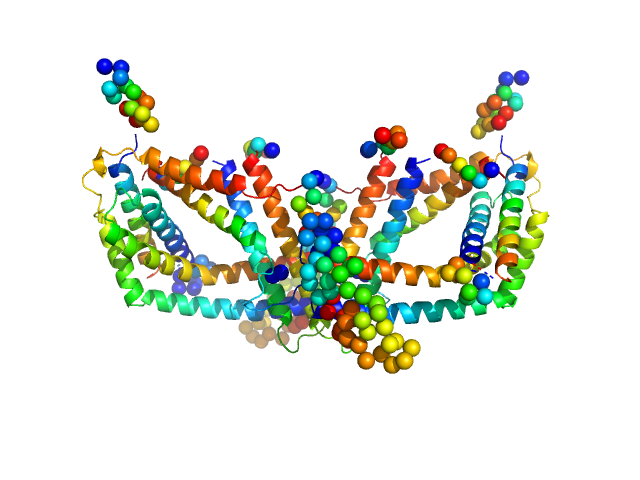
UniProt ID: A0A2N9BBV4 (1-374) Polyphosphate-targeting protein A
|
|
|
|
| Sample: |
Polyphosphate-targeting protein A dimer, 79 kDa Streptomyces chartreusis protein
|
| Buffer: |
20 mM Tris-HCl 400 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Nov 24
|
Structural and biochemical analysis of a phosin from Streptomyces chartreusis reveals a combined polyphosphate- and metal-binding fold.
FEBS Lett (2019)
Werten S, Rustmeier NH, Gemmer M, Virolle MJ, Hinrichs W
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.8 |
nm |
| VolumePorod |
124 |
nm3 |
|
|
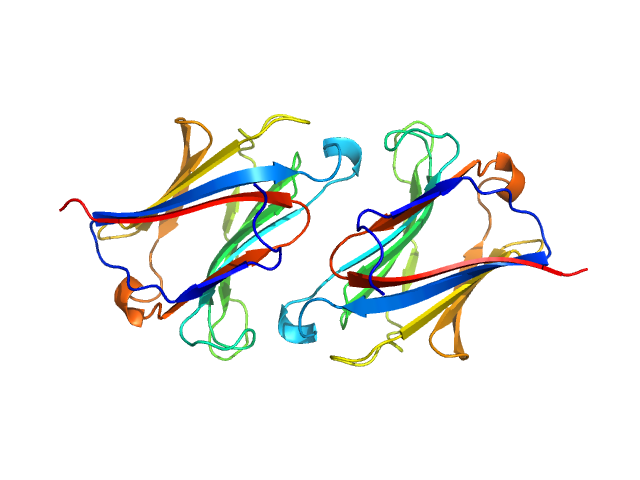
UniProt ID: Q05315 (None-None) Galectin-10 Tyr69Glu
|
|
|
|
| Sample: |
Galectin-10 Tyr69Glu dimer, 33 kDa Homo sapiens protein
|
| Buffer: |
20 mM Hepes 150 NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Feb 4
|
Protein crystallization promotes type 2 immunity and is reversible by antibody treatment.
Science 364(6442) (2019)
Persson EK, Verstraete K, Heyndrickx I, Gevaert E, Aegerter H, Percier JM, Deswarte K, Verschueren KHG, Dansercoer A, Gras D, Chanez P, Bachert C, Gonçalves A, Van Gorp H, De Haard H, Blanchetot C, Saunders M, Hammad H, Savvides SN, Lambrecht BN
|
| RgGuinier |
2.1 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
UniProt ID: O88998-2 (17-153) Noelin
|
|
|

|
| Sample: |
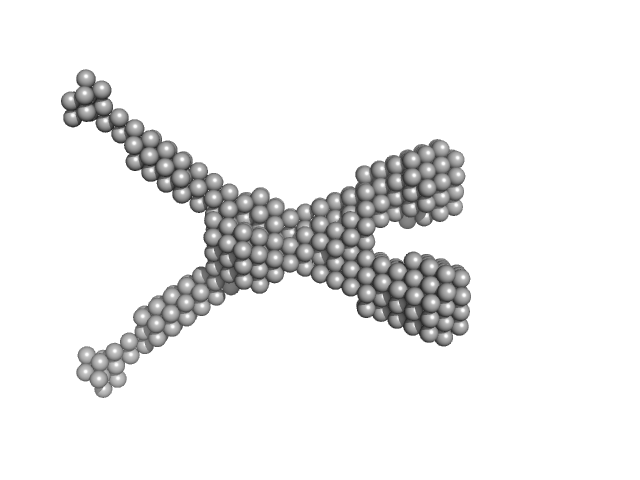
Noelin tetramer, 72 kDa Mus musculus protein
|
| Buffer: |
150 mM NaCl, 20 mM HEPES, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Feb 5
|
Design and structural characterisation of olfactomedin-1 variants as tools for functional studies.
BMC Mol Cell Biol 20(1):50 (2019)
Pronker MF, van den Hoek H, Janssen BJC
|
| RgGuinier |
5.4 |
nm |
| Dmax |
16.3 |
nm |
| VolumePorod |
160 |
nm3 |
|
|