UniProt ID: P61823 (None-None) Ribonuclease pancreatic
|
|
|
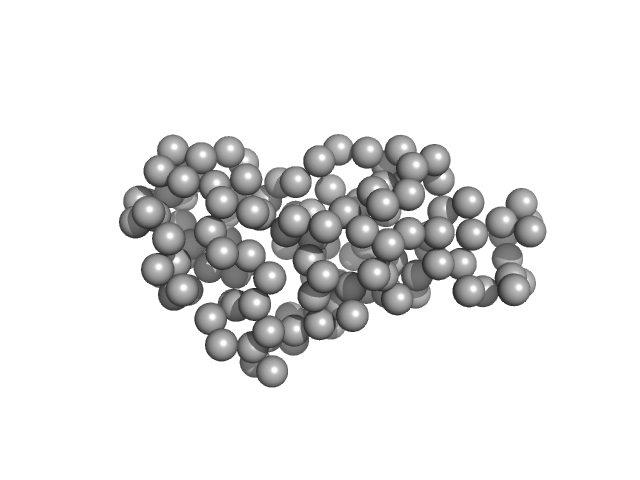
|
| Sample: |
Ribonuclease pancreatic monomer, 16 kDa Bos taurus protein
|
| Buffer: |
PBS, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Sep 18
|
Standard proteins
Darja Ruskule
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.0 |
nm |
| VolumePorod |
15 |
nm3 |
|
|
UniProt ID: P61823 (None-None) Ribonuclease pancreatic
|
|
|

|
| Sample: |
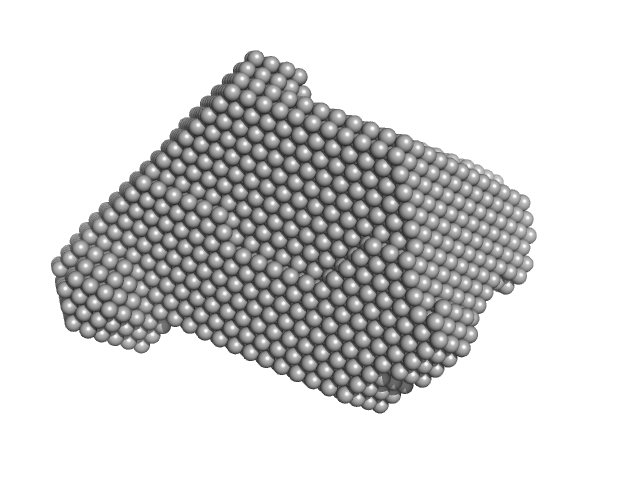
Ribonuclease pancreatic monomer, 16 kDa Bos taurus protein
|
| Buffer: |
PBS, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Sep 18
|
Standard proteins
Darja Ruskule
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.0 |
nm |
| VolumePorod |
17 |
nm3 |
|
|
UniProt ID: P61823 (None-None) Ribonuclease pancreatic
|
|
|
|
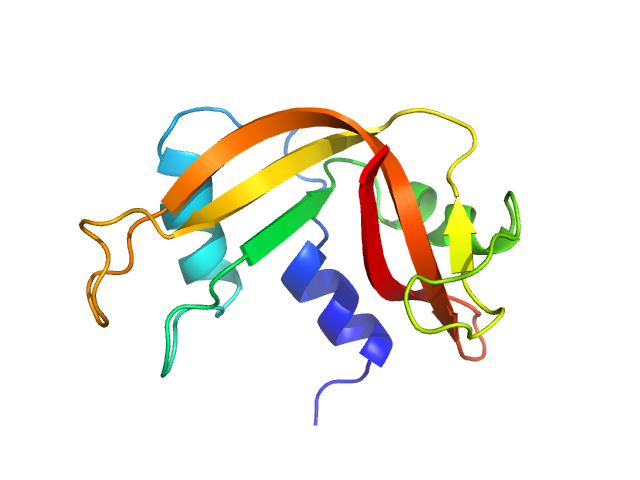
| Sample: |
Ribonuclease pancreatic monomer, 16 kDa Bos taurus protein
|
| Buffer: |
phosphate buffered saline (PBS), pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Jul 29
|
Machine Learning Methods for X-Ray Scattering Data Analysis from Biomacromolecular Solutions.
Biophys J 114(11):2485-2492 (2018)
Franke D, Jeffries CM, Svergun DI
|
| RgGuinier |
1.6 |
nm |
| Dmax |
5.6 |
nm |
| VolumePorod |
16 |
nm3 |
|
|
UniProt ID: P61823 (None-None) Ribonuclease pancreatic
|
|
|
|
| Sample: |
Ribonuclease pancreatic monomer, 16 kDa Bos taurus protein
|
| Buffer: |
10 mM HCl, pH: 1 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Jul 29
|
Machine Learning Methods for X-Ray Scattering Data Analysis from Biomacromolecular Solutions.
Biophys J 114(11):2485-2492 (2018)
Franke D, Jeffries CM, Svergun DI
|
| RgGuinier |
2.3 |
nm |
| Dmax |
9.0 |
nm |
|
|
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|
|
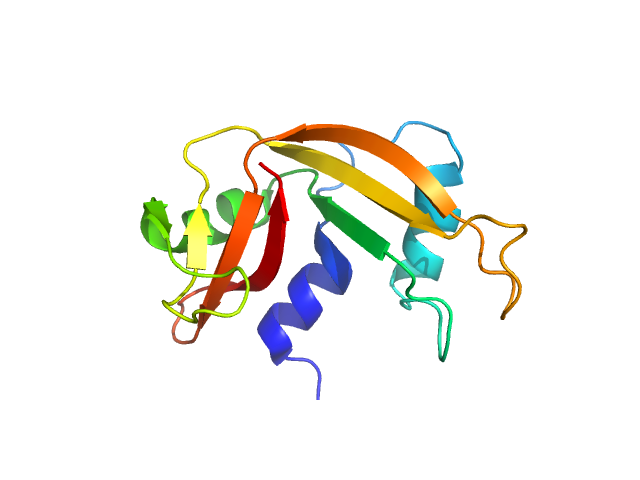
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
PBS, pH: 7.4 |
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, University of Copenhagen, Department of Drug Design and Pharmacology on 2018 Jan 23
|
Size-exclusion chromatography small-angle X-ray scattering of water soluble proteins on a laboratory instrument.
J Appl Crystallogr 51(Pt 6):1623-1632 (2018)
Bucciarelli S, Midtgaard SR, Nors Pedersen M, Skou S, Arleth L, Vestergaard B
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.8 |
nm |
| VolumePorod |
10 |
nm3 |
|
|
UniProt ID: P05307 (1-510) Protein disulfide-isomerase
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|

|
| Sample: |
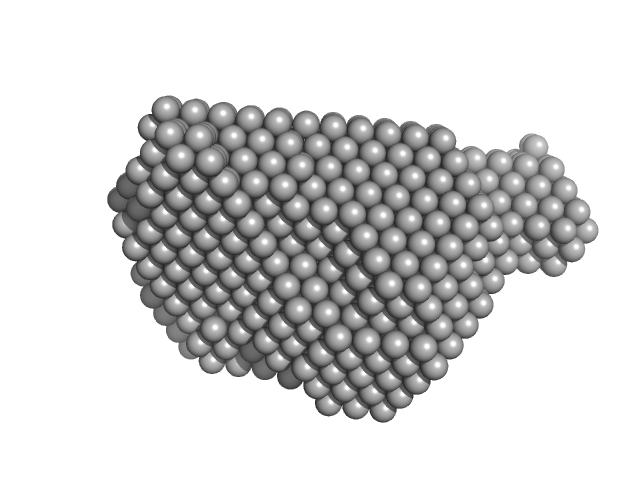
Protein disulfide-isomerase monomer, 57 kDa Bos taurus protein
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris-HCL, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2015 Nov 4
|
Insights on the dynamic behavior of protein disulfide isomerase in the solution environment through the SAXS technique
In Silico Pharmacology 12(1) (2024)
Sanyasi C, Balakrishnan S, Chinnasamy T, Venugopalan N, Kandavelu P, Batra-Safferling R, Muthuvel S
|
| RgGuinier |
3.0 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
85 |
nm3 |
|
|
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|

|
| Sample: |
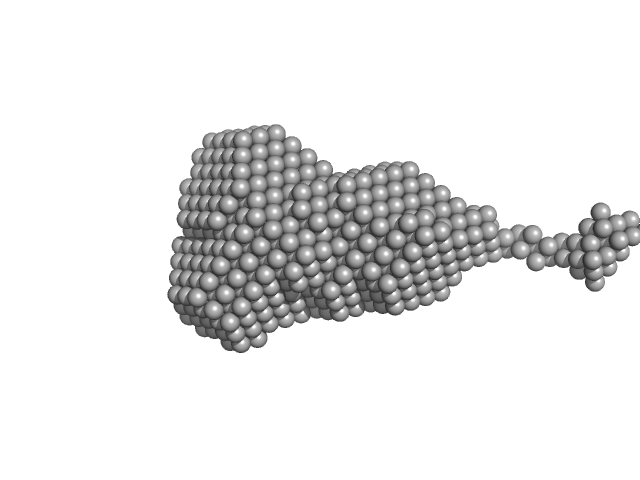
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris-HCL, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2015 Nov 4
|
Insights on the dynamic behavior of protein disulfide isomerase in the solution environment through the SAXS technique
In Silico Pharmacology 12(1) (2024)
Sanyasi C, Balakrishnan S, Chinnasamy T, Venugopalan N, Kandavelu P, Batra-Safferling R, Muthuvel S
|
| RgGuinier |
1.9 |
nm |
| Dmax |
7.2 |
nm |
| VolumePorod |
20 |
nm3 |
|
|
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|

|
| Sample: |
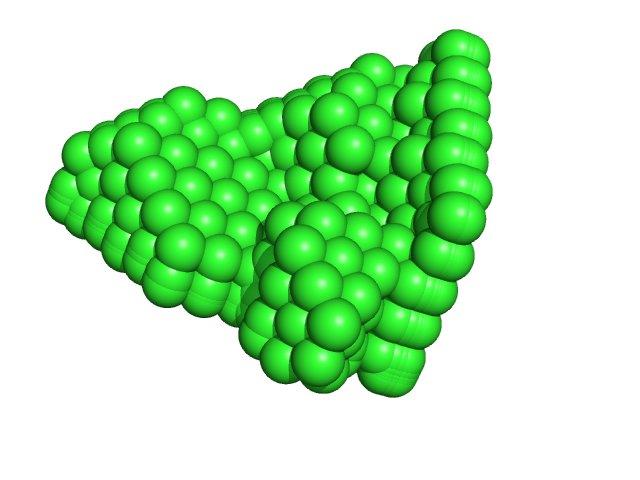
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.9 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|
|
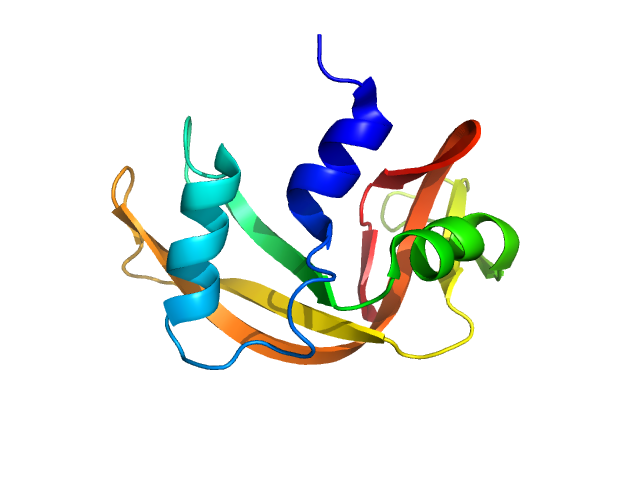
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.4 |
nm |
| Dmax |
4.4 |
nm |
|
|
UniProt ID: P61823 (27-150) Ribonuclease pancreatic
|
|
|
|
| Sample: |
Ribonuclease pancreatic monomer, 14 kDa Bos taurus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 22
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
1.5 |
nm |
| Dmax |
4.1 |
nm |
|
|