|
|
|
|

|
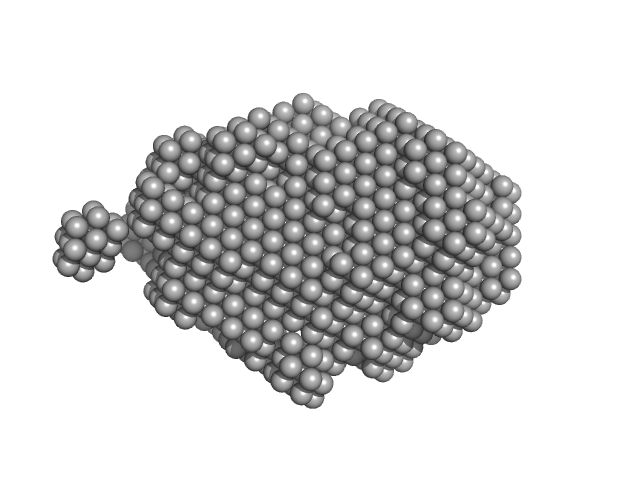
| Sample: |
Trehalose transferase (Trehalose phosphorylase/synthase) monomer, 47 kDa Thermoproteus uzoniensis protein
|
| Buffer: |
50 mM HEPES, 100 NaCl, 4 mM DTT, 20 mM MgCl2, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Sep 27
|
Anomeric Selectivity of Trehalose Transferase with Rare L-Sugars
ACS Catalysis (2020)
Mestrom L, Marsden S, van der Eijk H, Laustsen J, Jeffries C, Svergun D, Hagedoorn P, Bento I, Hanefeld U
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|

|
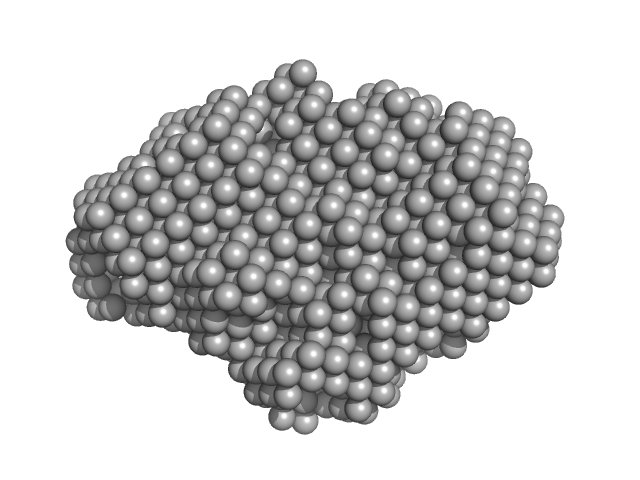
| Sample: |
Trehalose transferase (Trehalose phosphorylase/synthase) monomer, 47 kDa Thermoproteus uzoniensis protein
|
| Buffer: |
50 mM HEPES, 100 NaCl, 4 mM DTT, 20 mM MgCl2, 1 mM trehalose, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Sep 27
|
Anomeric Selectivity of Trehalose Transferase with Rare L-Sugars
ACS Catalysis (2020)
Mestrom L, Marsden S, van der Eijk H, Laustsen J, Jeffries C, Svergun D, Hagedoorn P, Bento I, Hanefeld U
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.3 |
nm |
| VolumePorod |
76 |
nm3 |
|
|
|
|
|
|

|
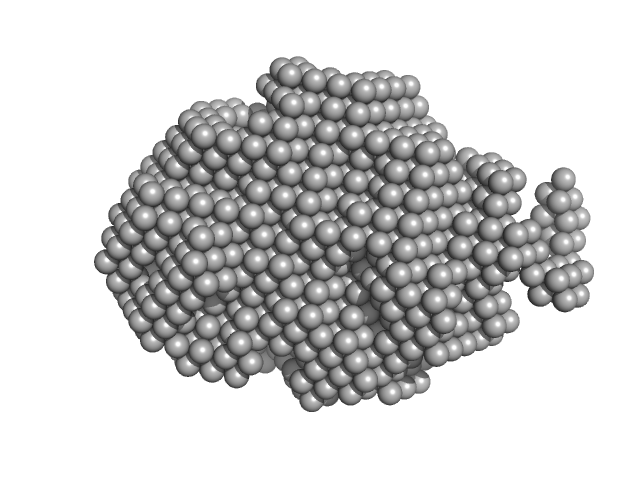
| Sample: |
Trehalose transferase (Trehalose phosphorylase/synthase) monomer, 47 kDa Thermoproteus uzoniensis protein
|
| Buffer: |
50 mM HEPES, 100 NaCl, 4 mM DTT, 20 mM MgCl2, 1 mM UDP-glucose, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Sep 27
|
Anomeric Selectivity of Trehalose Transferase with Rare L-Sugars
ACS Catalysis (2020)
Mestrom L, Marsden S, van der Eijk H, Laustsen J, Jeffries C, Svergun D, Hagedoorn P, Bento I, Hanefeld U
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.5 |
nm |
| VolumePorod |
73 |
nm3 |
|
|
|
|
|
|

|
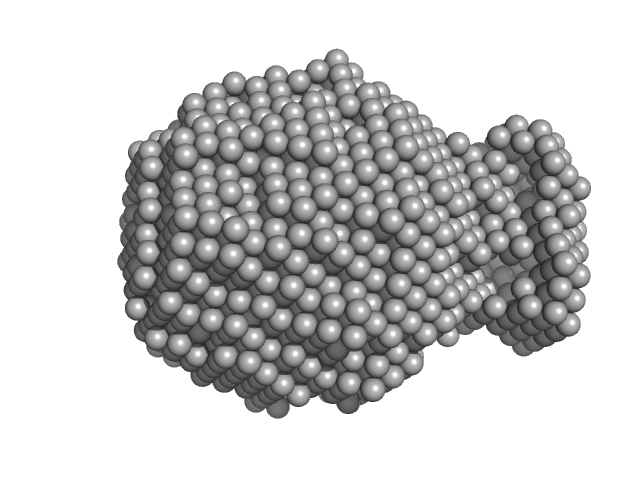
| Sample: |
Collagen alpha-3(VI) chain, N2 domain monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM TRIS, pH 7.4, 150mM NaCl 3% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 May 4
|
Structure of a collagen VI α3 chain VWA domain array: adaptability and functional implications of myopathy causing mutations
Journal of Biological Chemistry :jbc.RA120.014865 (2020)
Solomon-Degefa H, Gebauer J, Jeffries C, Freiburg C, Meckelburg P, Bird L, Baumann U, Svergun D, Owens R, Werner J, Behrmann E, Paulsson M, Wagener R
|
| RgGuinier |
1.8 |
nm |
| Dmax |
5.8 |
nm |
| VolumePorod |
40 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
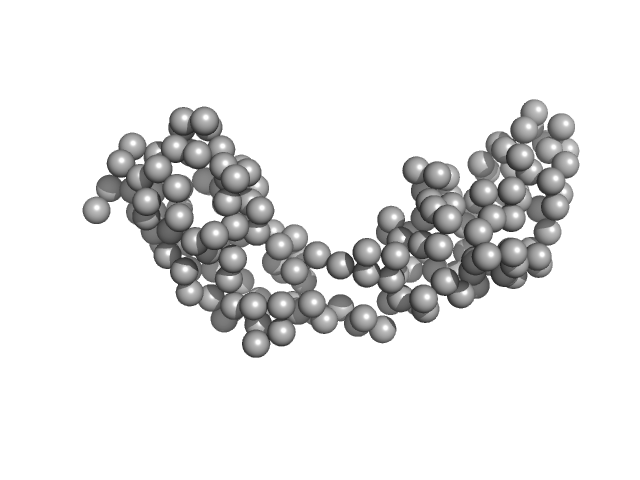
Minimal hepatocyte growth factor mimic K1K1 monomer, 19 kDa synthetic construct protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2016 Nov 3
|
Dimerization of kringle 1 domain from hepatocyte growth factor/scatter factor provides a potent MET receptor agonist
Life Science Alliance 5(12):e202201424 (2022)
de Nola G, Leclercq B, Mougel A, Taront S, Simonneau C, Forneris F, Adriaenssens E, Drobecq H, Iamele L, Dubuquoy L, Melnyk O, Gherardi E, de Jonge H, Vicogne J
|
| RgGuinier |
2.2 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
22 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
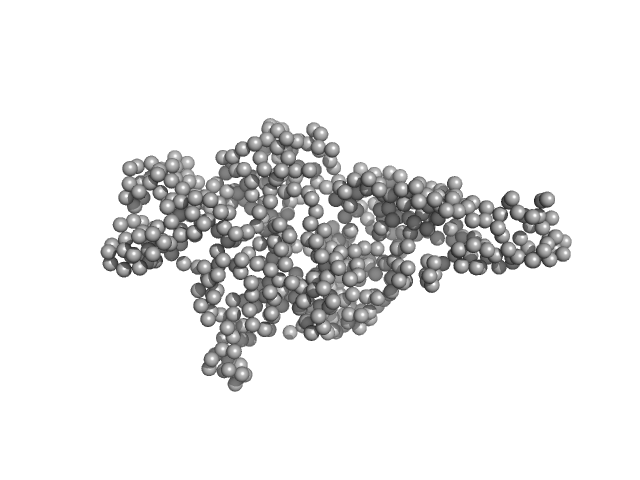
Hepatocyte growth factor receptor monomer, 62 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jan 27
|
Dimerization of kringle 1 domain from hepatocyte growth factor/scatter factor provides a potent MET receptor agonist
Life Science Alliance 5(12):e202201424 (2022)
de Nola G, Leclercq B, Mougel A, Taront S, Simonneau C, Forneris F, Adriaenssens E, Drobecq H, Iamele L, Dubuquoy L, Melnyk O, Gherardi E, de Jonge H, Vicogne J
|
| RgGuinier |
3.2 |
nm |
| Dmax |
11.7 |
nm |
| VolumePorod |
126 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
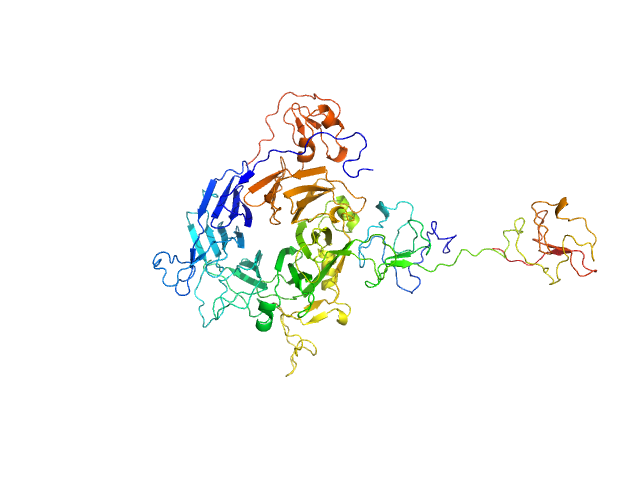
Minimal hepatocyte growth factor mimic K1K1 monomer, 19 kDa synthetic construct protein
Hepatocyte growth factor receptor monomer, 62 kDa Homo sapiens protein
|
| Buffer: |
25 mM Tris, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Jan 23
|
Dimerization of kringle 1 domain from hepatocyte growth factor/scatter factor provides a potent MET receptor agonist
Life Science Alliance 5(12):e202201424 (2022)
de Nola G, Leclercq B, Mougel A, Taront S, Simonneau C, Forneris F, Adriaenssens E, Drobecq H, Iamele L, Dubuquoy L, Melnyk O, Gherardi E, de Jonge H, Vicogne J
|
| RgGuinier |
3.7 |
nm |
| Dmax |
14.3 |
nm |
| VolumePorod |
136 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
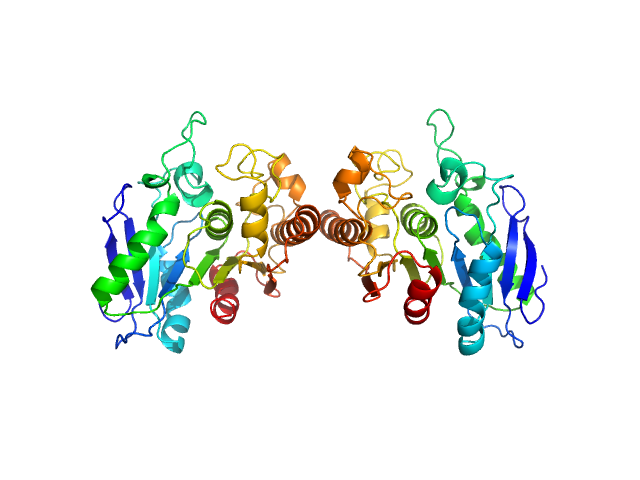
Poly(Aspartic acid) hydrolase-1 dimer, 65 kDa Sphingomonas sp. KT-1 protein
|
| Buffer: |
20 mM Tris pH 7.0, 100 mM NaCl, 1 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Nov 27
|
Structural Characterization of Sphingomonas
sp. KT-1 PahZ1-Catalyzed Biodegradation of Thermally Synthesized Poly(aspartic acid)
ACS Sustainable Chemistry & Engineering (2020)
Brambley C, Bolay A, Salvo H, Jansch A, Yared T, Miller J, Wallen J, Weiland M
|
| RgGuinier |
2.7 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
|
|
|
|
|
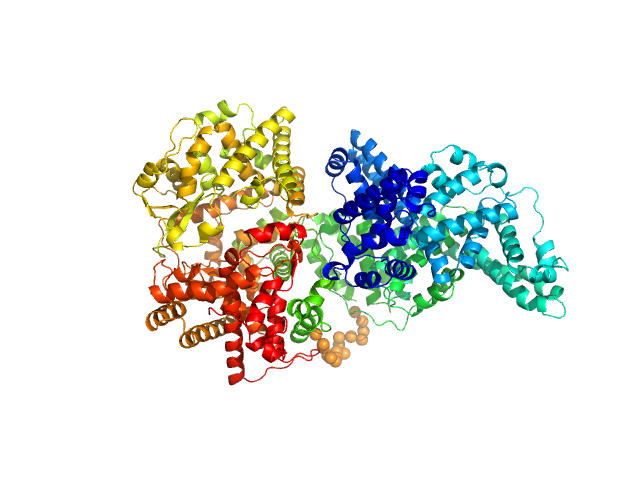
| Sample: |
Neprilysin - G400V mutant monomer, 80 kDa Homo sapiens protein
Human serum albumin - C58S mutant monomer, 66 kDa Homo sapiens protein
|
| Buffer: |
10 mM histidine, pH: 5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Dec 21
|
Albumin-neprilysin fusion protein: understanding stability using small angle X-ray scattering and molecular dynamic simulations.
Sci Rep 10(1):10089 (2020)
Kulakova A, Indrakumar S, Sønderby Tuelung P, Mahapatra S, Streicher WW, Peters GHJ, Harris P
|
| RgGuinier |
4.9 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
267 |
nm3 |
|
|
|
|
|
|
|
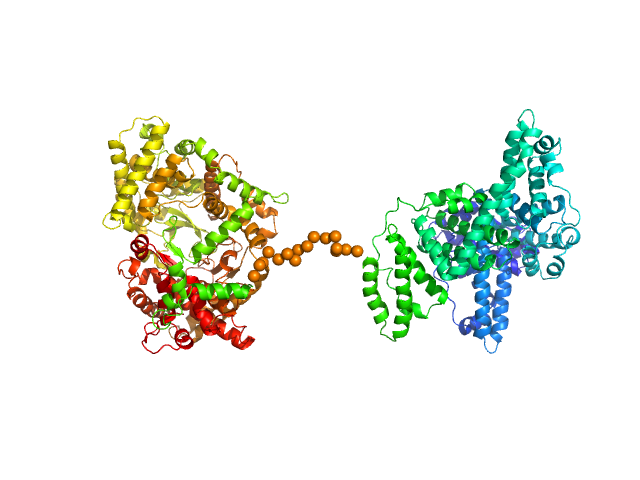
| Sample: |
Neprilysin - G400V mutant monomer, 80 kDa Homo sapiens protein
Human serum albumin - C58S mutant monomer, 66 kDa Homo sapiens protein
|
| Buffer: |
10 mM histidine, pH: 5.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2017 Dec 21
|
Albumin-neprilysin fusion protein: understanding stability using small angle X-ray scattering and molecular dynamic simulations.
Sci Rep 10(1):10089 (2020)
Kulakova A, Indrakumar S, Sønderby Tuelung P, Mahapatra S, Streicher WW, Peters GHJ, Harris P
|
| RgGuinier |
4.9 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
274 |
nm3 |
|
|