|
|
|
|
|
| Sample: |
Polyubiquitin-B monomer, 33 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02 % NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 28
|
Polyubiquitin ligand-induced phase transitions are optimized by spacing between ubiquitin units
Proceedings of the National Academy of Sciences 120(42) (2023)
Galagedera S, Dao T, Enos S, Chaudhuri A, Schmit J, Castañeda C
|
| RgGuinier |
3.3 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
41 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Polyubiquitin-B monomer, 34 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02 % NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 28
|
Polyubiquitin ligand-induced phase transitions are optimized by spacing between ubiquitin units
Proceedings of the National Academy of Sciences 120(42) (2023)
Galagedera S, Dao T, Enos S, Chaudhuri A, Schmit J, Castañeda C
|
| RgGuinier |
3.2 |
nm |
| Dmax |
12.2 |
nm |
| VolumePorod |
43 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Tetraubiquitin, M1(1-76)-PS(GS)4-Ub4 monomer, 36 kDa synthetic construct protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02 % NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 28
|
Polyubiquitin ligand-induced phase transitions are optimized by spacing between ubiquitin units
Proceedings of the National Academy of Sciences 120(42) (2023)
Galagedera S, Dao T, Enos S, Chaudhuri A, Schmit J, Castañeda C
|
| RgGuinier |
3.4 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Tetraubiquitin, M1(1-74)-A(EA3K)3A-Ub monomer, 39 kDa synthetic construct protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02 % NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 28
|
Polyubiquitin ligand-induced phase transitions are optimized by spacing between ubiquitin units
Proceedings of the National Academy of Sciences 120(42) (2023)
Galagedera S, Dao T, Enos S, Chaudhuri A, Schmit J, Castañeda C
|
| RgGuinier |
4.2 |
nm |
| Dmax |
18.2 |
nm |
| VolumePorod |
62 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Tetraubiquitin, M1(1-74)-A(EA3K)6A-Ub monomer, 43 kDa synthetic construct protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02 % NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2022 Nov 28
|
Polyubiquitin ligand-induced phase transitions are optimized by spacing between ubiquitin units
Proceedings of the National Academy of Sciences 120(42) (2023)
Galagedera S, Dao T, Enos S, Chaudhuri A, Schmit J, Castañeda C
|
| RgGuinier |
5.2 |
nm |
| Dmax |
21.2 |
nm |
| VolumePorod |
108 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
HOTag-(PA)25-Ubiquitin tetramer, 68 kDa synthetic construct protein
|
| Buffer: |
20 mM sodium phosphate, 0.5 mM EDTA, 0.02 % NaN3, pH: 6.8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2023 Mar 22
|
Polyubiquitin ligand-induced phase transitions are optimized by spacing between ubiquitin units
Proceedings of the National Academy of Sciences 120(42) (2023)
Galagedera S, Dao T, Enos S, Chaudhuri A, Schmit J, Castañeda C
|
| RgGuinier |
6.3 |
nm |
| Dmax |
24.9 |
nm |
| VolumePorod |
180 |
nm3 |
|
|
|
|
|
|
|
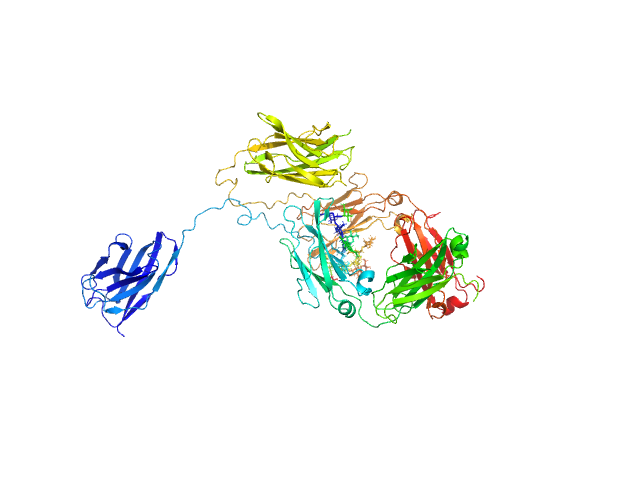
| Sample: |
Immunoglobulin heavy constant gamma 1 monomer, 78 kDa Homo sapiens protein
|
| Buffer: |
20 mM L-histidine, 138 mM NaCl, 2.6 mM KCl buffer, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Sep 28
|
The solution structure of the heavy chain-only C5-Fc nanobody reveals exposed variable regions that are optimal for COVID-19 antigen interactions.
J Biol Chem :105337 (2023)
Gao X, Thrush JW, Gor J, Naismith JH, Owens RJ, Perkins SJ
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.2 |
nm |
| VolumePorod |
157 |
nm3 |
|
|
|
|
|
|
|
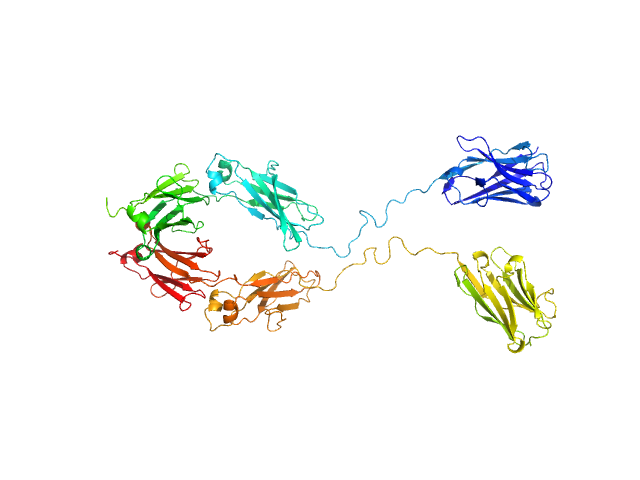
| Sample: |
Immunoglobulin heavy constant gamma 1 monomer, 78 kDa Homo sapiens protein
|
| Buffer: |
20 mM L-histidine, 138 mM NaCl, 2.6 mM KCl buffer, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2022 Sep 28
|
The solution structure of the heavy chain-only C5-Fc nanobody reveals exposed variable regions that are optimal for COVID-19 antigen interactions.
J Biol Chem :105337 (2023)
Gao X, Thrush JW, Gor J, Naismith JH, Owens RJ, Perkins SJ
|
| RgGuinier |
4.0 |
nm |
| Dmax |
13.1 |
nm |
| VolumePorod |
175 |
nm3 |
|
|
|
|
|
|
|
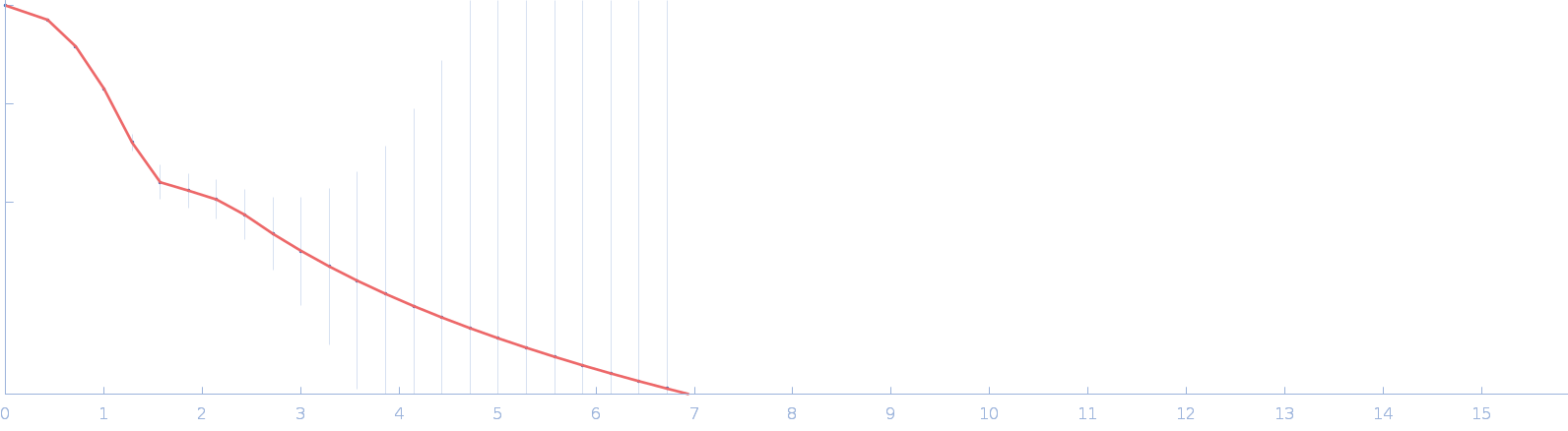
| Sample: |
DUF4374 domain-containing protein monomer, 46 kDa Bacteroides thetaiotaomicron (strain … protein
|
| Buffer: |
20 mM Tris 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 May 9
|
Iron acquisition by a commensal bacterium modifies host nutritional immunity during Salmonella infection.
Cell Host Microbe 31(10):1639-1654.e10 (2023)
Spiga L, Fansler RT, Perera YR, Shealy NG, Munneke MJ, David HE, Torres TP, Lemoff A, Ran X, Richardson KL, Pudlo N, Martens EC, Folta-Stogniew E, Yang ZJ, Skaar EP, Byndloss MX, Chazin WJ, Zhu W
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
|
|
|
|
|
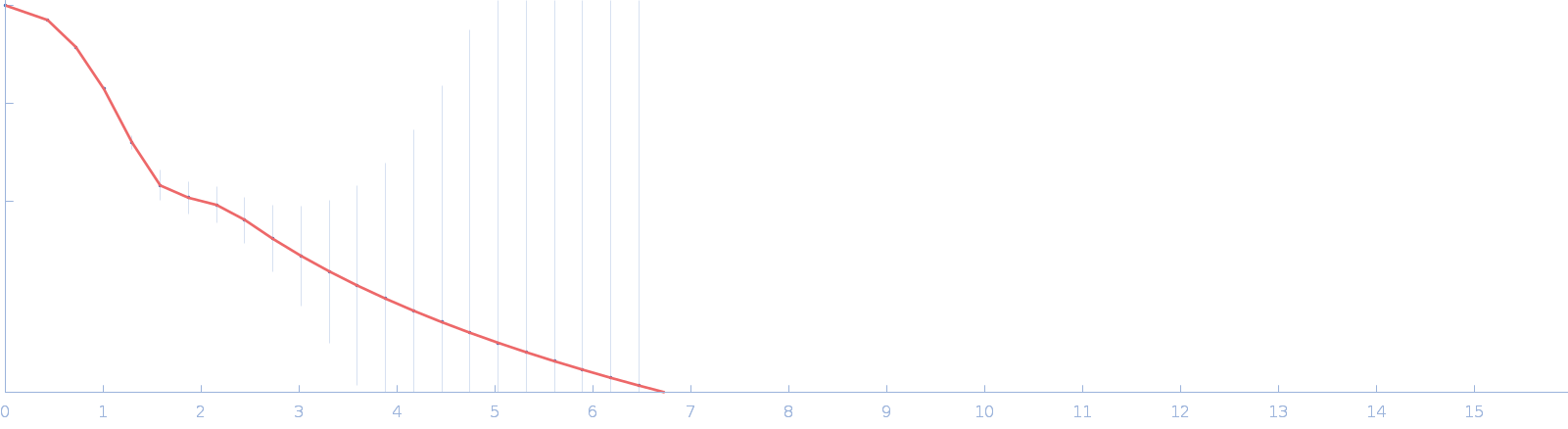
| Sample: |
DUF4374 domain-containing protein monomer, 46 kDa Bacteroides thetaiotaomicron (strain … protein
|
| Buffer: |
20 mM Tris 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 12-ID-B, Advanced Photon Source (APS), Argonne National Laboratory on 2021 May 9
|
Iron acquisition by a commensal bacterium modifies host nutritional immunity during Salmonella infection.
Cell Host Microbe 31(10):1639-1654.e10 (2023)
Spiga L, Fansler RT, Perera YR, Shealy NG, Munneke MJ, David HE, Torres TP, Lemoff A, Ran X, Richardson KL, Pudlo N, Martens EC, Folta-Stogniew E, Yang ZJ, Skaar EP, Byndloss MX, Chazin WJ, Zhu W
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
77 |
nm3 |
|
|