|
|
|
|
|
| Sample: |
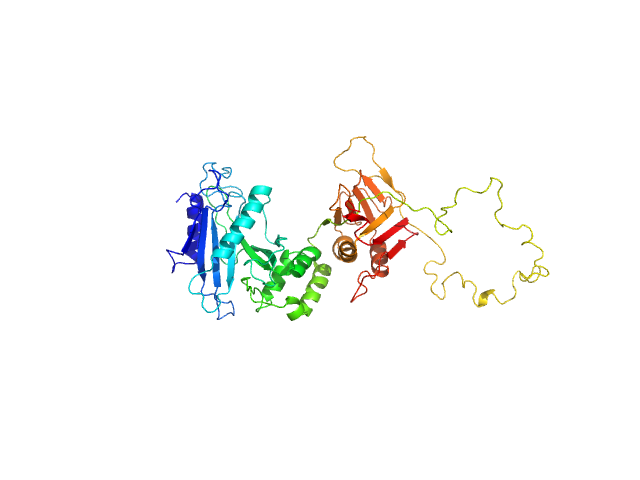
Suppressor of fused homolog monomer, 51 kDa Homo sapiens protein
|
| Buffer: |
50 mM bis-TRIS pH 5.5, 200 mM NaCl, 10% glycerol, pH: 5.5
|
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2018 Jun 30
|
Suppressor of Fused
Valerie Biou
|
| RgGuinier |
3.2 |
nm |
| Dmax |
14.0 |
nm |
|
|
|
|
|
|

|
| Sample: |
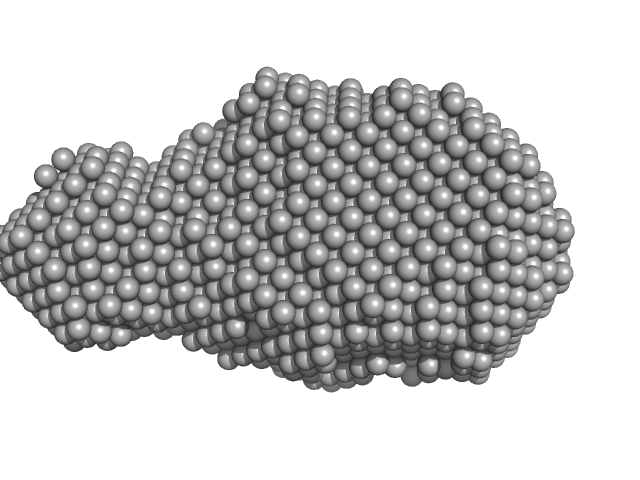
Putative DNA binding protein dimer, 37 kDa Bacillus subtilis subsp. … protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2016 Aug 17
|
YeeF, a polymorphic toxin from Bacillus subtilis, is a ribosomal RNase with promiscuous DNA binding property
Krishan Gopal
|
| RgGuinier |
2.7 |
nm |
| Dmax |
9.5 |
nm |
| VolumePorod |
55 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
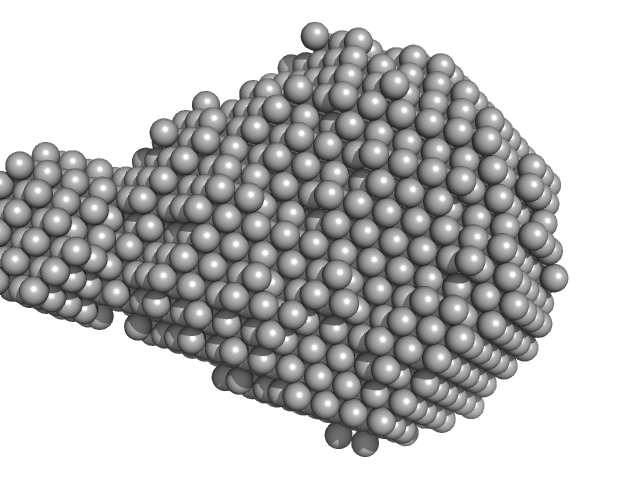
YezG, cognate immunity protein of YeeF monomer, 19 kDa Bacillus subtilis subsp. … protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2016 Aug 17
|
YeeF, a polymorphic toxin from Bacillus subtilis, is a ribosomal RNase with promiscuous DNA binding property
Krishan Gopal
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.1 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
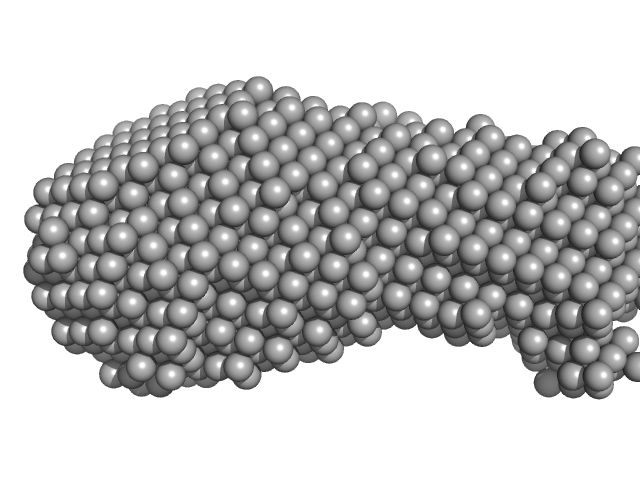
Putative DNA binding protein dimer, 37 kDa Bacillus subtilis subsp. … protein
YezG, cognate immunity protein of YeeF monomer, 19 kDa Bacillus subtilis subsp. … protein
|
| Buffer: |
20 mM HEPES 150 mM NaCl, pH: 7.5
|
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR - Institute of Microbial Technology (IMTech) on 2016 Aug 18
|
YeeF, a polymorphic toxin from Bacillus subtilis, is a ribosomal RNase with promiscuous DNA binding property
Krishan Gopal
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
115 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
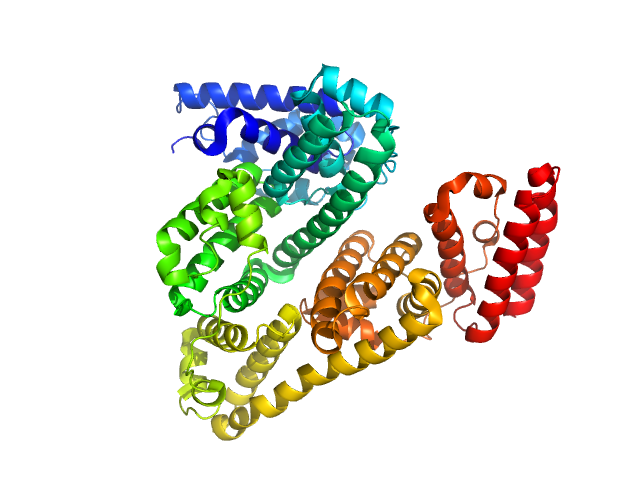
Bovine serum albumin monomer, 66 kDa Bos taurus protein
|
| Buffer: |
TRIS 50mM, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Dec 21
|
A high flux setup for millisecond-scale small-angle X-ray scattering studies on macromolecular solutions
Clement Blanchet
|
|
|
|
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
100 mM HEPES, 1 mM MgCl2, pH: 7
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Dec 21
|
A high flux setup for millisecond-scale small-angle X-ray scattering studies on macromolecular solutions
Clement Blanchet
|
|
|
|
|
|
|
|
| Sample: |
Cytochrome C monomer, 12 kDa Bos taurus protein
|
| Buffer: |
TRIS 50mM, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Dec 21
|
A high flux setup for millisecond-scale small-angle X-ray scattering studies on macromolecular solutions
Clement Blanchet
|
|
|
|
|
|
|
|
| Sample: |
PAS fold family dimer, 78 kDa Coleofasciculus chthonoplastes PCC … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5 mM MgCl2, 5 % w/v Glycerol, pH: 7.5
|
| Experiment: |
SAXS
data collected at cSAXS, Swiss Light Source on 2015 Mar 11
|
mPAC Delta132
Robert Lindner
|
| RgGuinier |
4.4 |
nm |
| Dmax |
15.9 |
nm |
| VolumePorod |
132 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
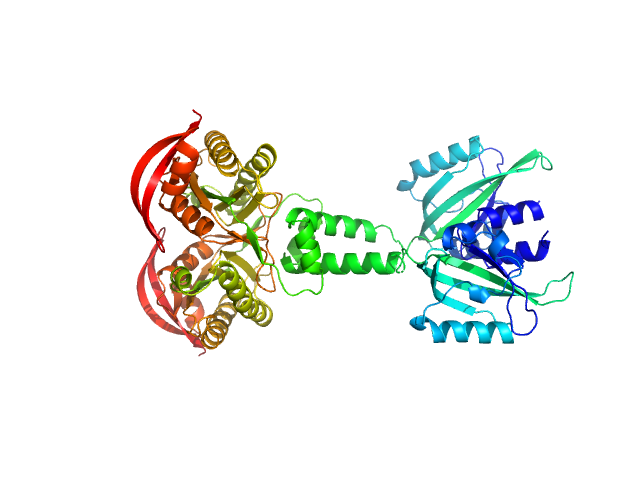
PAS fold family dimer, 78 kDa Coleofasciculus chthonoplastes PCC … protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 5 mM MgCl2, 5 % w/v Glycerol, 1 mM ApCpp, pH: 7.5
|
| Experiment: |
SAXS
data collected at cSAXS, Swiss Light Source on 2015 Mar 11
|
mPAC Delta132
Robert Lindner
|
| RgGuinier |
4.0 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
99 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
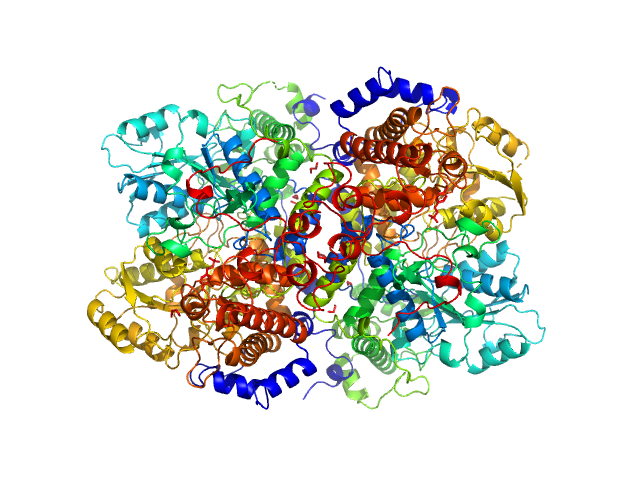
Glycine decarboxylase dimer, 219 kDa Homo sapiens protein
|
| Buffer: |
10 mM TRIS pH 7.5, 100 mM NaCl, 1 mM DTT, pH: 7.5
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Nov 12
|
Glycine decarboxylase (P-protein of glycine cleavage system)
Bart Van Laer
|
| RgGuinier |
3.9 |
nm |
| VolumePorod |
304 |
nm3 |
|
|