|
|
|
|
|
| Sample: |
Neurospora Varkud Satellite minimal trans ribozyme monomer, 33 kDa Neurospora crassa RNA
|
| Buffer: |
50 mM MES, 50 mM KCl, pH: 6.5
|
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, Département de Biochimie, Université de Montréal on 2019 Sep 19
|
An integrative NMR-SAXS approach for structural determination of large RNAs defines the substrate-free state of a trans
-cleaving Neurospora
Varkud Satellite ribozyme
Nucleic Acids Research (2021)
Dagenais P, Desjardins G, Legault P
|
| RgGuinier |
3.6 |
nm |
| Dmax |
12.4 |
nm |
| VolumePorod |
55 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
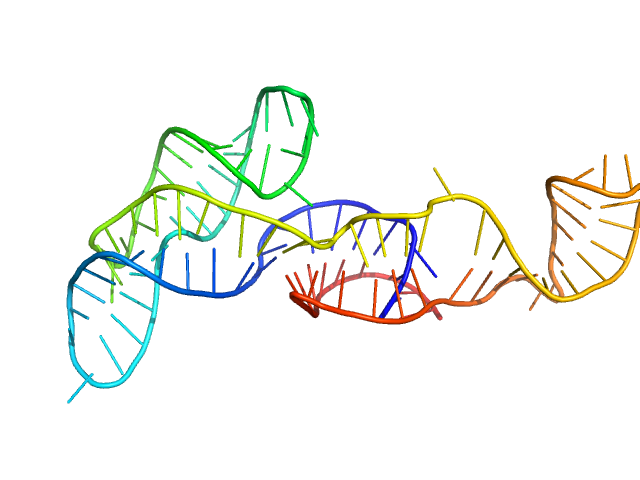
Neurospora Varkud Satellite minimal trans ribozyme monomer, 33 kDa Neurospora crassa RNA
|
| Buffer: |
50 mM MES, 50 mM KCl, 5 mM MgCl2,, pH: 6.5
|
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, Département de Biochimie, Université de Montréal on 2019 Sep 19
|
An integrative NMR-SAXS approach for structural determination of large RNAs defines the substrate-free state of a trans
-cleaving Neurospora
Varkud Satellite ribozyme
Nucleic Acids Research (2021)
Dagenais P, Desjardins G, Legault P
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Neurospora Varkud Satellite minimal trans ribozyme monomer, 33 kDa Neurospora crassa RNA
|
| Buffer: |
50 mM MES, 50 mM KCl, 20 mM MgCl2,, pH: 6.5
|
| Experiment: |
SAXS
data collected at Xenocs BioXolver L with MetalJet, Département de Biochimie, Université de Montréal on 2019 Sep 19
|
An integrative NMR-SAXS approach for structural determination of large RNAs defines the substrate-free state of a trans
-cleaving Neurospora
Varkud Satellite ribozyme
Nucleic Acids Research (2021)
Dagenais P, Desjardins G, Legault P
|
| RgGuinier |
3.5 |
nm |
| Dmax |
11.4 |
nm |
| VolumePorod |
61 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Microtubule-associated protein tau, Tau35 fragment monomer, 27 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Oct 8
|
The Disease Associated Tau35 Fragment has an Increased Propensity to Aggregate Compared to Full-Length Tau
Frontiers in Molecular Biosciences 8 (2021)
Lyu C, Da Vela S, Al-Hilaly Y, Marshall K, Thorogate R, Svergun D, Serpell L, Pastore A, Hanger D
|
| RgGuinier |
4.6 |
nm |
| Dmax |
16.2 |
nm |
| VolumePorod |
93 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Microtubule-associated protein tau, 2N3R isoform monomer, 43 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Oct 8
|
The Disease Associated Tau35 Fragment has an Increased Propensity to Aggregate Compared to Full-Length Tau
Frontiers in Molecular Biosciences 8 (2021)
Lyu C, Da Vela S, Al-Hilaly Y, Marshall K, Thorogate R, Svergun D, Serpell L, Pastore A, Hanger D
|
| RgGuinier |
6.3 |
nm |
| Dmax |
19.8 |
nm |
| VolumePorod |
211 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Microtubule-associated protein tau, 2N4R isoform monomer, 46 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4
|
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Oct 8
|
The Disease Associated Tau35 Fragment has an Increased Propensity to Aggregate Compared to Full-Length Tau
Frontiers in Molecular Biosciences 8 (2021)
Lyu C, Da Vela S, Al-Hilaly Y, Marshall K, Thorogate R, Svergun D, Serpell L, Pastore A, Hanger D
|
| RgGuinier |
6.7 |
nm |
| Dmax |
23.5 |
nm |
| VolumePorod |
254 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
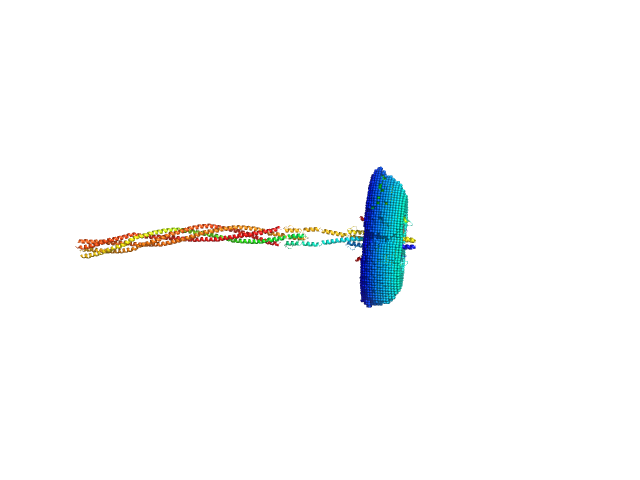
Sensory rhodopsin II from Natronbacterium pharaonis dimer, 53 kDa Natronomonas pharaonis protein
Sensory rhodopsin II transducer from Natronomonas pharaonis dimer, 116 kDa Natronomonas pharaonis protein
|
| Buffer: |
150 mM NaCl, 25 mM Na/Na-Pi, 1.0 mM EDTA, 0.05% DDM, pH: 8
|
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Dec 8
|
Ambiguities in and completeness of SAS data analysis of membrane proteins: the case of the sensory rhodopsin II–transducer complex
Acta Crystallographica Section D Structural Biology 77(11) (2021)
Ryzhykau Y, Vlasov A, Orekhov P, Rulev M, Rogachev A, Vlasova A, Kazantsev A, Verteletskiy D, Skoi V, Brennich M, Pernot P, Murugova T, Gordeliy V, Kuklin A
|
| RgGuinier |
7.6 |
nm |
| Dmax |
41.0 |
nm |
|
|
|
|
|
|

|
| Sample: |
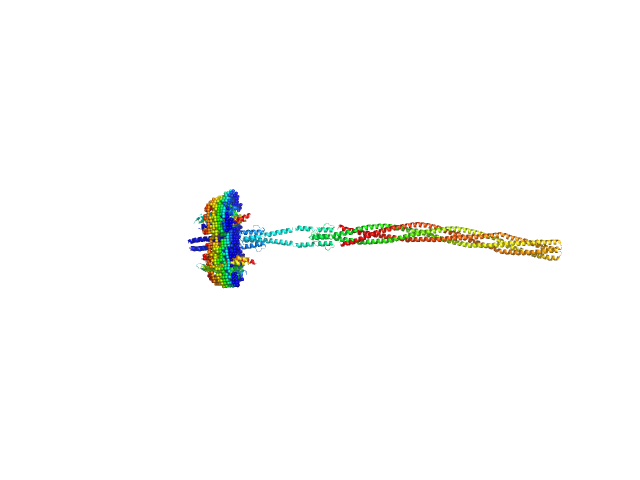
Sensory rhodopsin II from Natronbacterium pharaonis dimer, 53 kDa Natronomonas pharaonis protein
Sensory rhodopsin II transducer from Natronomonas pharaonis dimer, 116 kDa Natronomonas pharaonis protein
|
| Buffer: |
150 mM NaCl, 25 mM Na/Na-Pi, 1.0 mM EDTA, 0.05% DDM (D2O buffer), pH: 8
|
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 Jan 25
|
Ambiguities in and completeness of SAS data analysis of membrane proteins: the case of the sensory rhodopsin II–transducer complex
Acta Crystallographica Section D Structural Biology 77(11) (2021)
Ryzhykau Y, Vlasov A, Orekhov P, Rulev M, Rogachev A, Vlasova A, Kazantsev A, Verteletskiy D, Skoi V, Brennich M, Pernot P, Murugova T, Gordeliy V, Kuklin A
|
| RgGuinier |
8.9 |
nm |
| Dmax |
39.0 |
nm |
|
|
|
|
|
|

|
| Sample: |
Sensory rhodopsin II from Natronbacterium pharaonis dimer, 53 kDa Natronomonas pharaonis protein
Sensory rhodopsin II transducer from Natronomonas pharaonis dimer, 116 kDa Natronomonas pharaonis protein
|
| Buffer: |
1400 mM NaCl, 49.4 mM Na/Na-Pi, 1.0 mM EDTA, 0.05% DDM (D2O buffer), pH: 8
|
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 Feb 10
|
Ambiguities in and completeness of SAS data analysis of membrane proteins: the case of the sensory rhodopsin II–transducer complex
Acta Crystallographica Section D Structural Biology 77(11) (2021)
Ryzhykau Y, Vlasov A, Orekhov P, Rulev M, Rogachev A, Vlasova A, Kazantsev A, Verteletskiy D, Skoi V, Brennich M, Pernot P, Murugova T, Gordeliy V, Kuklin A
|
| RgGuinier |
8.6 |
nm |
| Dmax |
39.0 |
nm |
|
|
|
|
|
|

|
| Sample: |
Sensory rhodopsin II from Natronbacterium pharaonis dimer, 53 kDa Natronomonas pharaonis protein
Sensory rhodopsin II transducer from Natronomonas pharaonis dimer, 116 kDa Natronomonas pharaonis protein
|
| Buffer: |
2800 mM NaCl, 76.6 mM Na/Na-Pi, 1.0 mM EDTA, 0.05% DDM (D2O buffer), pH: 8
|
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 Feb 10
|
Ambiguities in and completeness of SAS data analysis of membrane proteins: the case of the sensory rhodopsin II–transducer complex
Acta Crystallographica Section D Structural Biology 77(11) (2021)
Ryzhykau Y, Vlasov A, Orekhov P, Rulev M, Rogachev A, Vlasova A, Kazantsev A, Verteletskiy D, Skoi V, Brennich M, Pernot P, Murugova T, Gordeliy V, Kuklin A
|
| RgGuinier |
9.0 |
nm |
| Dmax |
39.0 |
nm |
|
|