|
|
|
|
|
| Sample: |
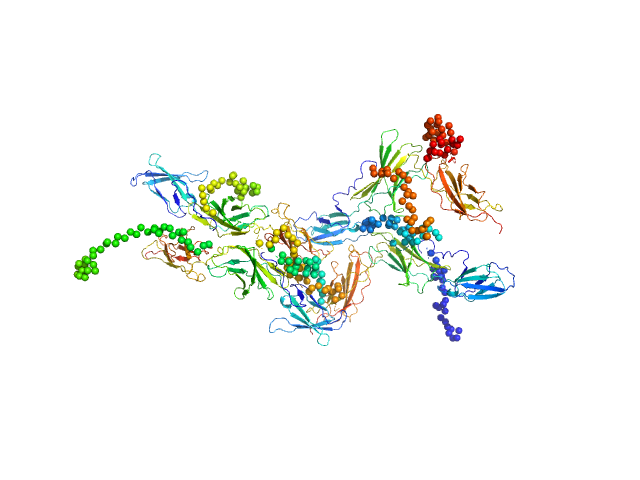
Interleukin-6 receptor subunit alpha tetramer, 164 kDa Homo sapiens protein
Interleukin-6 receptor subunit alpha dimer, 82 kDa Homo sapiens protein
|
| Buffer: |
water, pH: 7.5
|
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2014 Oct 1
|
Structure and target interaction of a G-quadruplex RNA-aptamer.
RNA Biol 13(10):973-987 (2016)
...Valentini E, Magbanua E, Kwiatkowski M, Chauvot de Beauchêne I, Krichel B, Schamoni K, Uetrecht C, Svergun DI, Schlüter H, Zacharias M, Hahn U
|
| RgGuinier |
4.9 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
240 |
nm3 |
|