UniProt ID: P51587 (1002-1551) Breast cancer type 2 susceptibility protein (BRC repeats 1-4)
UniProt ID: Q06609 (1-339) DNA repair protein RAD51 homolog 1 (F86E A89E)
|
|
|
|
| Sample: |
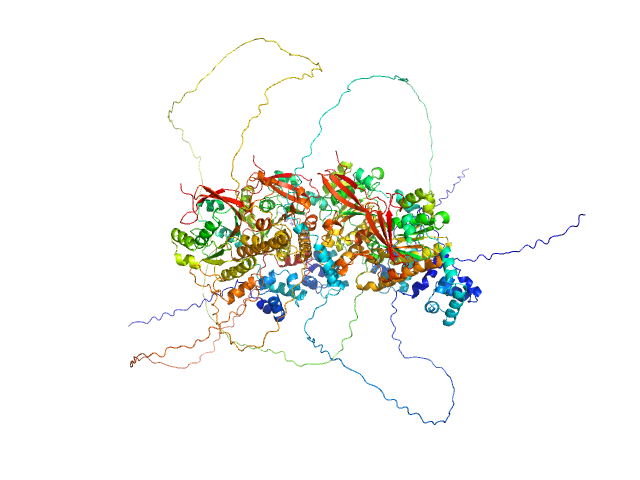
Breast cancer type 2 susceptibility protein (BRC repeats 1-4) monomer, 62 kDa Homo sapiens protein
DNA repair protein RAD51 homolog 1 (F86E A89E) tetramer, 148 kDa Homo sapiens protein
|
| Buffer: |
20 mM K₂HPO₄/KH₂PO₄, 100 mM NaCl, 100 mM Li₂SO₄, 1 mM DTT, 1% sucrose, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Dec 2
|
Breaking new ground into RAD51–BRC repeats interplay in Homologous Recombination
(2025)
Rinaldi F, Franco P, Veronesi M, Romeo E, Bresciani V, Varignani G, Catalano F, Bernetti M, Masetti M, Langer J, Girotto S, Cavalli A
|
| RgGuinier |
7.7 |
nm |
| Dmax |
34.0 |
nm |
| VolumePorod |
409 |
nm3 |
|
|
UniProt ID: P51587 (1002-1551) Breast cancer type 2 susceptibility protein (BRC repeats 1-4)
UniProt ID: Q06609 (1-339) DNA repair protein RAD51 homolog 1 (F86E A89E)
|
|
|
|
| Sample: |
Breast cancer type 2 susceptibility protein (BRC repeats 1-4) monomer, 62 kDa Homo sapiens protein
DNA repair protein RAD51 homolog 1 (F86E A89E) tetramer, 148 kDa Homo sapiens protein
|
| Buffer: |
20 mM K₂HPO₄/KH₂PO₄, 100 mM NaCl, 200 mM Li₂SO₄, 5% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Dec 2
|
Breaking new ground into RAD51–BRC repeats interplay in Homologous Recombination
(2025)
Rinaldi F, Franco P, Veronesi M, Romeo E, Bresciani V, Varignani G, Catalano F, Bernetti M, Masetti M, Langer J, Girotto S, Cavalli A
|
| RgGuinier |
8.1 |
nm |
| Dmax |
33.0 |
nm |
| VolumePorod |
422 |
nm3 |
|
|
UniProt ID: P51587 (1002-1551) Breast cancer type 2 susceptibility protein (BRC repeats 1-4)
UniProt ID: Q06609 (1-339) DNA repair protein RAD51 homolog 1 (F86E A89E)
|
|
|
|
| Sample: |
Breast cancer type 2 susceptibility protein (BRC repeats 1-4) monomer, 62 kDa Homo sapiens protein
DNA repair protein RAD51 homolog 1 (F86E A89E) tetramer, 148 kDa Homo sapiens protein
|
| Buffer: |
20 mM K₂HPO₄/KH₂PO₄, 100 mM NaCl, 200 mM Li₂SO₄, 5% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Dec 2
|
Breaking new ground into RAD51–BRC repeats interplay in Homologous Recombination
(2025)
Rinaldi F, Franco P, Veronesi M, Romeo E, Bresciani V, Varignani G, Catalano F, Bernetti M, Masetti M, Langer J, Girotto S, Cavalli A
|
| RgGuinier |
7.6 |
nm |
| Dmax |
33.3 |
nm |
| VolumePorod |
392 |
nm3 |
|
|
UniProt ID: P51587 (1002-1551) Breast cancer type 2 susceptibility protein (BRC repeats 1-4)
UniProt ID: Q06609 (1-339) DNA repair protein RAD51 homolog 1 (F86E A89E)
|
|
|
|
| Sample: |
Breast cancer type 2 susceptibility protein (BRC repeats 1-4) monomer, 62 kDa Homo sapiens protein
DNA repair protein RAD51 homolog 1 (F86E A89E) tetramer, 148 kDa Homo sapiens protein
|
| Buffer: |
20 mM K₂HPO₄/KH₂PO₄, 100 mM NaCl, 200 mM Li₂SO₄, 5% glycerol, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 Dec 2
|
Breaking new ground into RAD51–BRC repeats interplay in Homologous Recombination
(2025)
Rinaldi F, Franco P, Veronesi M, Romeo E, Bresciani V, Varignani G, Catalano F, Bernetti M, Masetti M, Langer J, Girotto S, Cavalli A
|
| RgGuinier |
7.7 |
nm |
| Dmax |
33.0 |
nm |
| VolumePorod |
417 |
nm3 |
|
|
UniProt ID: O95994 (41-171) Anterior gradient protein 2 homolog
|
|
|
|
| Sample: |
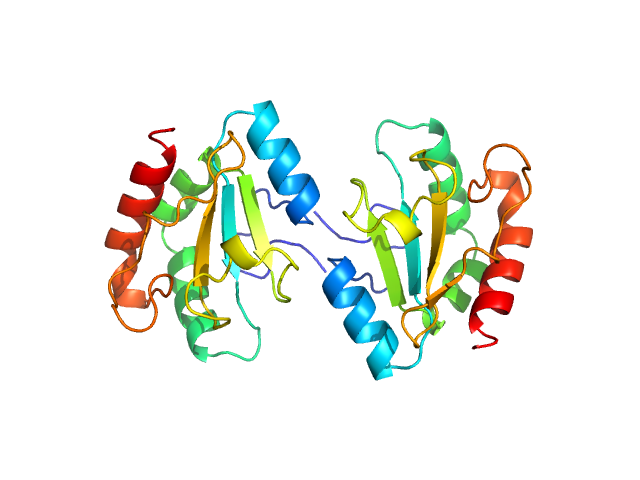
Anterior gradient protein 2 homolog dimer, 30 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH7.4, 150 mM NaCl, 1% glycerol, 0.2 mM TCEP, pH: 7.4 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2024 May 5
|
A structural basis for chaperone repression of stress signaling from the endoplasmic reticulum.
Mol Cell (2025)
Neidhardt L, Tung J, Kuchersky M, Milczarek J, Kargas V, Stott K, Rosenzweig R, Ron D, Yan Y
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.3 |
nm |
| VolumePorod |
30 |
nm3 |
|
|
UniProt ID: P02791 (2-175) Ferritin light chain
|
|
|
|
| Sample: |
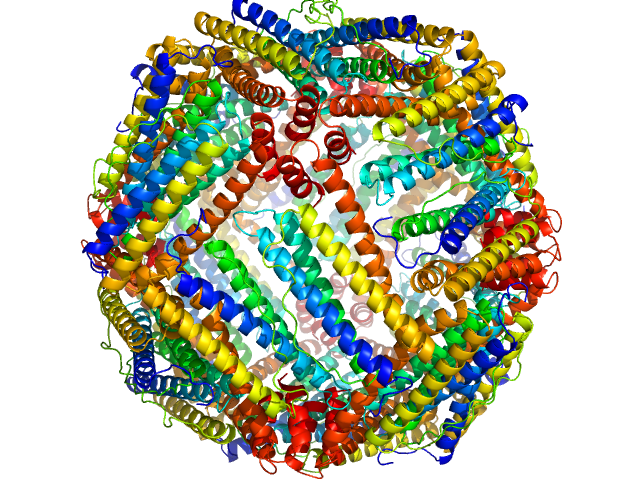
Ferritin light chain 24-mer, 476 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
5.4 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
651 |
nm3 |
|
|
UniProt ID: P02791 (2-175) Ferritin light chain
|
|
|
|
| Sample: |
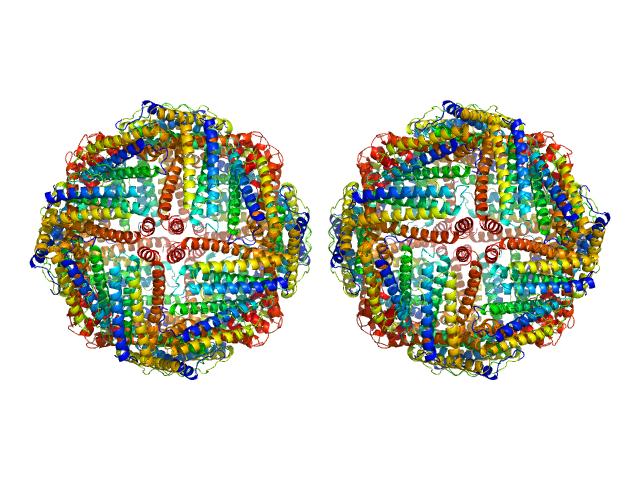
Ferritin light chain, 953 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
7.7 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
1751 |
nm3 |
|
|
UniProt ID: P02791 (2-175) Ferritin light chain
|
|
|
|
| Sample: |
Ferritin light chain, 953 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
8.0 |
nm |
| Dmax |
26.8 |
nm |
| VolumePorod |
1369 |
nm3 |
|
|
UniProt ID: P02791 (2-175) Ferritin light chain
|
|
|
|
| Sample: |
Ferritin light chain, 953 kDa Equus caballus protein
|
| Buffer: |
137 mM NaCl, 2.7 mM KCl, 10 mM phosphate buffer, 0.9 mM CaCl2, 0.5 mM MgCl2, pH: 7.4 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2024 May 24
|
Structural insights into the nature of static and dynamic apoferritin dimers
Aleksei Tsarenko
|
| RgGuinier |
8.1 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
1677 |
nm3 |
|
|
UniProt ID: Q8DZX0 (311-412) ABC transporter, permease protein, extracellular domain
|
|
|
|
| Sample: |
ABC transporter, permease protein, extracellular domain monomer, 23 kDa Streptococcus agalactiae serotype … protein
|
| Buffer: |
25 mM MES, 500 mM NaCl, pH: 6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2020 Dec 3
|
The extracellular domain of SaNSrFP binds bacitracin and allows the identification of new members of the BceAB transporter family
Frontiers in Microbiology 16 (2025)
Mammen C, Gottstein J, Cea P, Tantsur K, Reiners J, Bonus M, Gohlke H, Smits S
|
| RgGuinier |
2.4 |
nm |
| Dmax |
9.0 |
nm |
| VolumePorod |
50 |
nm3 |
|
|