UniProt ID: X5KGL2 (1-250) ABC transporter ATP-binding protein
UniProt ID: Q8DZX0 (1-651) ABC transporter, permease protein
|
|
|
|
| Sample: |
ABC transporter ATP-binding protein dimer, 62 kDa Streptococcus agalactiae protein
ABC transporter, permease protein monomer, 74 kDa Streptococcus agalactiae serotype … protein
|
| Buffer: |
50 mM Tris, 300 mM NaCl, 10 % Glycerol, 0.005 % LMNG,, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2023 Apr 25
|
The extracellular domain of SaNSrFP binds bacitracin and allows the identification of new members of the BceAB transporter family
Frontiers in Microbiology 16 (2025)
Mammen C, Gottstein J, Cea P, Tantsur K, Reiners J, Bonus M, Gohlke H, Smits S
|
| RgGuinier |
5.8 |
nm |
| Dmax |
20.5 |
nm |
| VolumePorod |
735 |
nm3 |
|
|
UniProt ID: Q7AL96 (None-None) Recombination directionality factor RdfS
UniProt ID: None (None-None) attP_8 40-mer DNA
|
|
|
|
| Sample: |
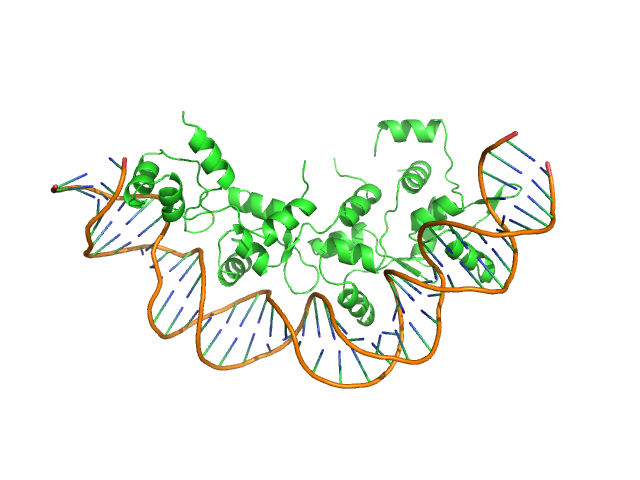
Recombination directionality factor RdfS tetramer, 52 kDa Mesorhizobium japonicum R7A protein
AttP_8 40-mer DNA dimer, 25 kDa DNA
|
| Buffer: |
150 mM Tris-HCl, 300 mM NaCl, 5% v/v glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2019 Jun 19
|
Structural basis for control of integrative and conjugative element excision and transfer by the oligomeric winged helix–turn–helix protein RdfS
Nucleic Acids Research 53(6) (2025)
Verdonk C, Agostino M, Eto K, Hall D, Bond C, Ramsay J
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.8 |
nm |
| VolumePorod |
134 |
nm3 |
|
|
UniProt ID: B5Z9C8 (1-117) Dihydroneopterin aldolase
|
|
|

|
| Sample: |
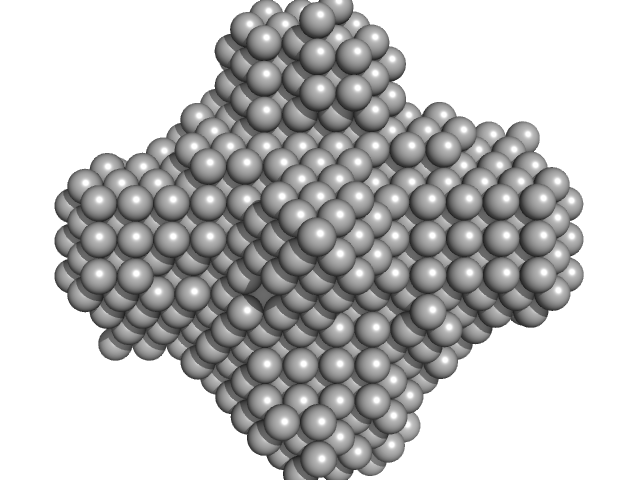
Dihydroneopterin aldolase tetramer, 55 kDa Helicobacter pylori (strain … protein
|
| Buffer: |
25 mM Tris-HCl pH 7.5 and 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2019 Apr 5
|
Structure of Helicobacter pylori dihydroneopterin aldolase suggests a fragment-based strategy for isozyme-specific inhibitor design.
Curr Res Struct Biol 5:100095 (2023)
Shaw GX, Fan L, Cherry S, Shi G, Tropea JE, Ji X
|
| RgGuinier |
2.5 |
nm |
| Dmax |
7.3 |
nm |
| VolumePorod |
77 |
nm3 |
|
|
UniProt ID: E9BL28 (None-None) Dynamin central region family protein
|
|
|
|
| Sample: |
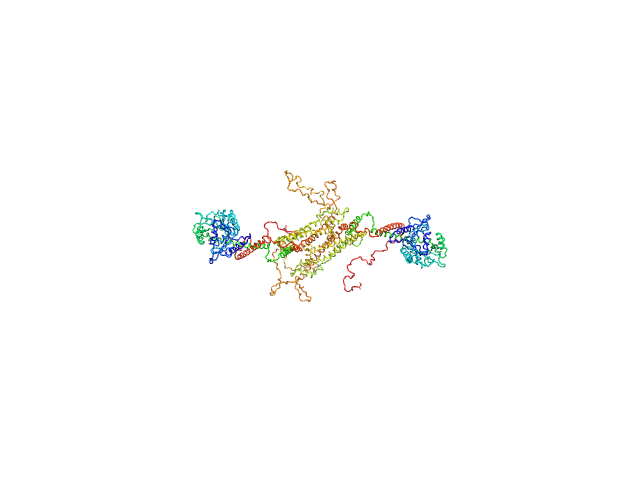
Dynamin central region family protein dimer, 165 kDa Leishmania donovani protein
|
| Buffer: |
20 mM HEPES, 200 mM NaCl, 3% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2024 Apr 4
|
Biophysical analysis of an oligomerization-attenuated variant of the Leishmania donovani dynamin-1-like protein.
Mol Biochem Parasitol 263:111691 (2025)
Wuyts E, Sundaramoorthy R, Tulloch L, Monsieurs P, Eadsforth TC, Pintelon I, Timmermans JP, Dujardin JC, Domagalska MA, Caljon G, De Rycker M, Postis VLG, Wyllie S, Sterckx YG
|
| RgGuinier |
6.6 |
nm |
| Dmax |
25.0 |
nm |
| VolumePorod |
340 |
nm3 |
|
|
UniProt ID: P03372 (310-549) Estrogen receptor alpha
|
|
|

|
| Sample: |
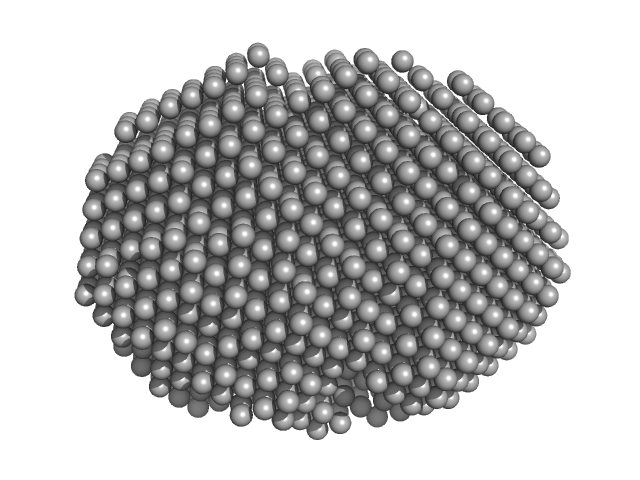
Estrogen receptor alpha dimer, 55 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris-HCl pH 7.5, 150 mM NaCl, 5% glycerol, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2024 Nov 9
|
A ternary switch model governing ERα ligand binding domain conformation.
Nat Commun 16(1):10363 (2025)
McDougal DP, Pederick JL, Novick SJ, Jovcevski B, Warrender AK, Pascal BD, Griffin PR, Bruning JB
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
74 |
nm3 |
|
|
UniProt ID: D6N7U3 (310-553) Estrogen receptor alpha
|
|
|

|
| Sample: |
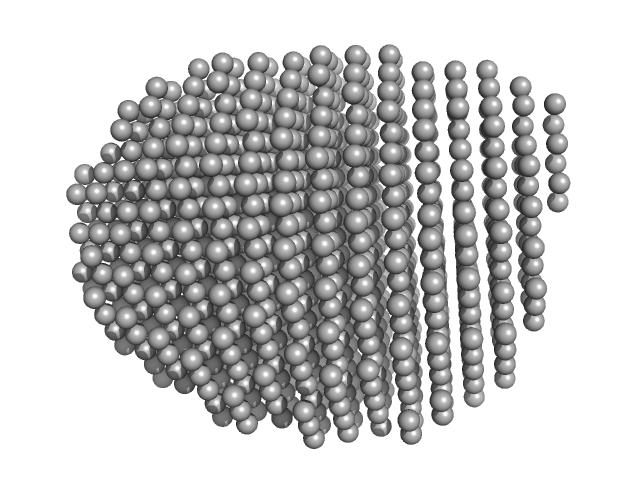
Estrogen receptor alpha dimer, 57 kDa Melanotaenia fluviatilis protein
|
| Buffer: |
20 mM Tris-HCl pH 7.5, 150 mM NaCl, 5% glycerol, 2 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at BioSAXS, Australian Synchrotron on 2024 Nov 9
|
A ternary switch model governing ERα ligand binding domain conformation.
Nat Commun 16(1):10363 (2025)
McDougal DP, Pederick JL, Novick SJ, Jovcevski B, Warrender AK, Pascal BD, Griffin PR, Bruning JB
|
| RgGuinier |
2.5 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
72 |
nm3 |
|
|
UniProt ID: O53943 (None-None) ESX-5 secretion-associated protein EspG5
UniProt ID: P9WPI1 (1-278) ESX-5 secretion system protein EccA5
|
|
|
|
| Sample: |
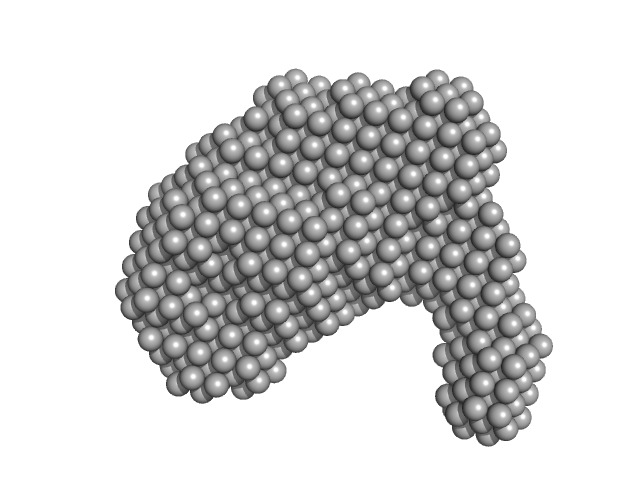
ESX-5 secretion-associated protein EspG5 monomer, 32 kDa Mycobacterium tuberculosis (strain … protein
ESX-5 secretion system protein EccA5 monomer, 31 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM HEPES, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2025 Jan 28
|
The crystal structure of the TPR domain of the EccA5 ATPase and demonstration of its interaction with EspG5 from the mycobacterial ESX-5 pathway.
FEBS Lett (2026)
Sharma VK, Vishwakarma J, Kabrambam R, Kumar S, Ramachandran R
|
| RgGuinier |
3.5 |
nm |
| Dmax |
9.1 |
nm |
| VolumePorod |
83 |
nm3 |
|
|
UniProt ID: O53943 (None-None) ESX-5 secretion-associated protein EspG5
UniProt ID: P9WPI1 (1-278) ESX-5 secretion system protein EccA5 N-terminal domain
|
|
|

|
| Sample: |
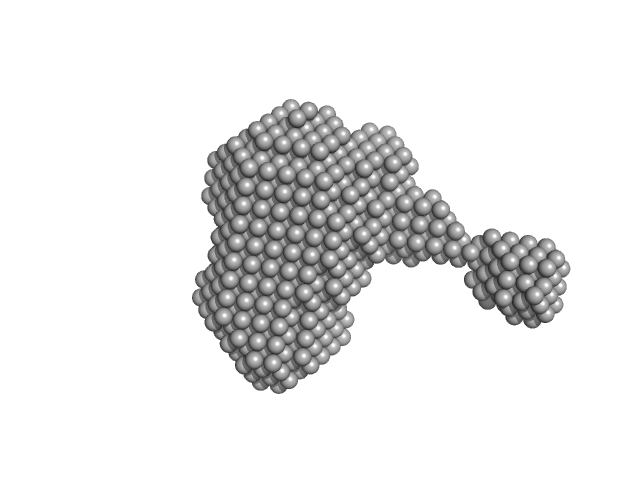
ESX-5 secretion-associated protein EspG5 monomer, 32 kDa Mycobacterium tuberculosis (strain … protein
ESX-5 secretion system protein EccA5 N-terminal domain monomer, 31 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50mM Hepes 200mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpace, CSIR-Central Drug Research Institute on 2025 Jan 28
|
SAXS analysis of EccA5 N-terminal domain and EspG5 of ESX-5 secretion pathway of Mycobacterium tuberculosis
Rajlakshmi K
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.7 |
nm |
| VolumePorod |
84 |
nm3 |
|
|
UniProt ID: P35755 (23-332) Iron-utilization periplasmic protein
|
|
|

|
| Sample: |
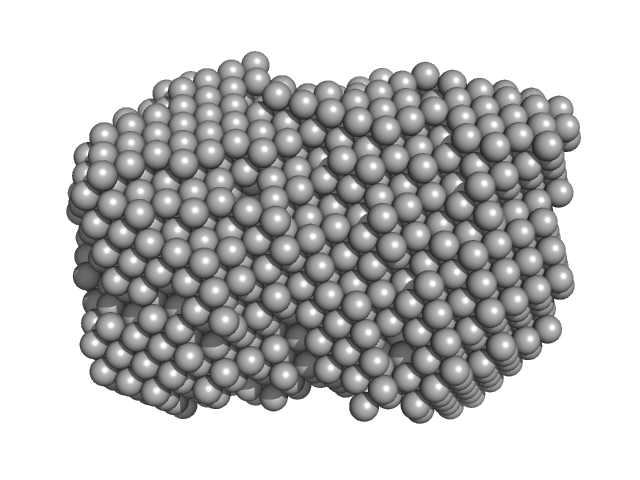
Iron-utilization periplasmic protein monomer, 34 kDa Haemophilus influenzae protein
|
| Buffer: |
1 mM Na2HPO4.7H2O, 0.18 mM KH2PO4, 13.7 mM NaCl, 0.27 mM KCl, 5%v/v Glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Aug 20
|
Conformational multiplicity of bacterial ferric binding protein revealed by small angle x-ray scattering and molecular dynamics calculations
The Journal of Chemical Physics 158(8) (2023)
Liu G, Ekmen E, Jalalypour F, Mertens H, Jeffries C, Svergun D, Atilgan A, Atilgan C, Sayers Z
|
| RgGuinier |
2.1 |
nm |
| Dmax |
6.3 |
nm |
| VolumePorod |
28 |
nm3 |
|
|
UniProt ID: P35755 (23-332) Iron-utilization periplasmic protein
|
|
|
|
| Sample: |
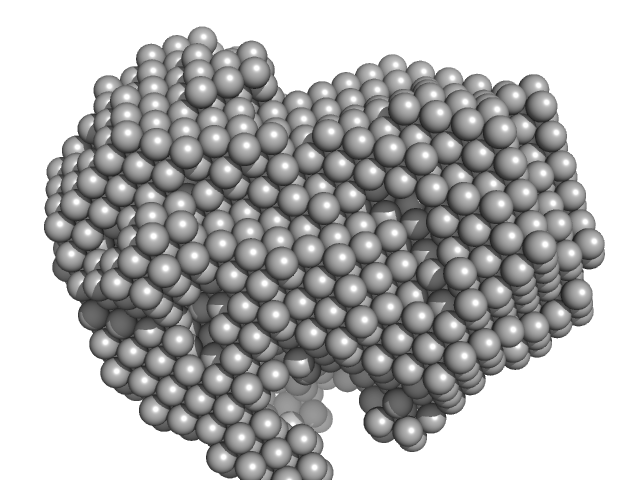
Iron-utilization periplasmic protein monomer, 34 kDa Haemophilus influenzae protein
|
| Buffer: |
10 mM Na2HPO4 . 7H2O 1.8 mM KH2PO4 137 mM NaCl 2.7 mM KCl 5% v/v Glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Aug 20
|
Conformational multiplicity of bacterial ferric binding protein revealed by small angle x-ray scattering and molecular dynamics calculations
The Journal of Chemical Physics 158(8) (2023)
Liu G, Ekmen E, Jalalypour F, Mertens H, Jeffries C, Svergun D, Atilgan A, Atilgan C, Sayers Z
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.2 |
nm |
| VolumePorod |
34 |
nm3 |
|
|