UniProt ID: P20273 (None-None) CD22 extracellular domain
|
|
|

|
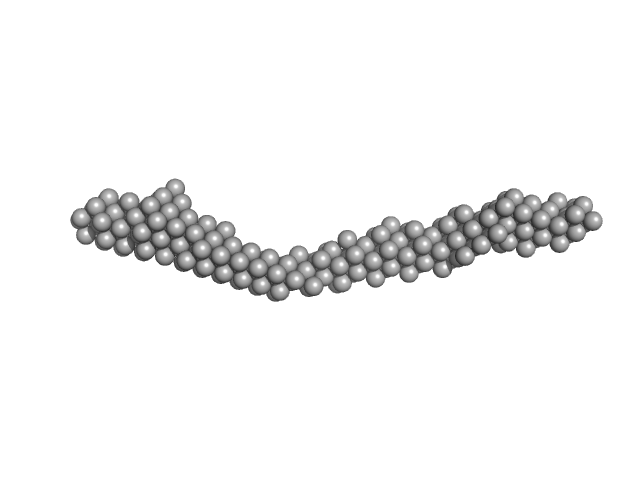
| Sample: |
CD22 extracellular domain monomer, 90 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris 150 mM NaCl, pH: 9 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2016 Apr 21
|
Molecular basis of human CD22 function and therapeutic targeting.
Nat Commun 8(1):764 (2017)
Ereño-Orbea J, Sicard T, Cui H, Mazhab-Jafari MT, Benlekbir S, Guarné A, Rubinstein JL, Julien JP
|
| RgGuinier |
8.0 |
nm |
| Dmax |
30.6 |
nm |
|
|
UniProt ID: P20273 (None-None) CD22 extracellular domain
UniProt ID: (None-None) alpha(2,6)-Sialyllactose
|
|
|

|
| Sample: |
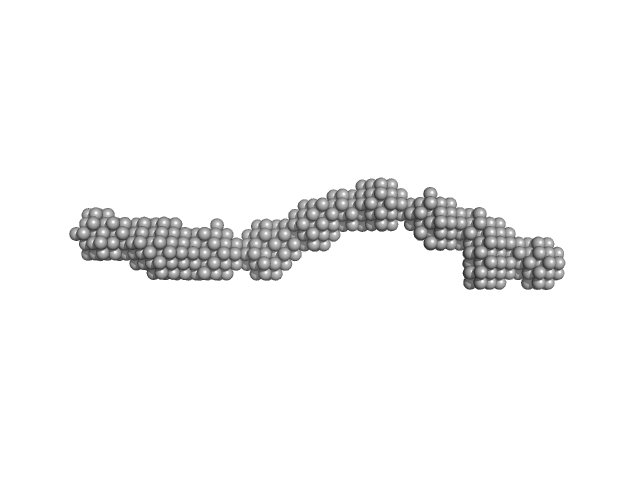
CD22 extracellular domain monomer, 90 kDa Homo sapiens protein
Alpha(2,6)-Sialyllactose, 1 kDa
|
| Buffer: |
20 mM Tris 150 mM NaCl, pH: 9 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2016 Apr 21
|
Molecular basis of human CD22 function and therapeutic targeting.
Nat Commun 8(1):764 (2017)
Ereño-Orbea J, Sicard T, Cui H, Mazhab-Jafari MT, Benlekbir S, Guarné A, Rubinstein JL, Julien JP
|
| RgGuinier |
8.1 |
nm |
| Dmax |
29.8 |
nm |
|
|
UniProt ID: A1U4R9 (None-None) Diguanylate cyclase with PAS/PAC sensor
|
|
|

|
| Sample: |
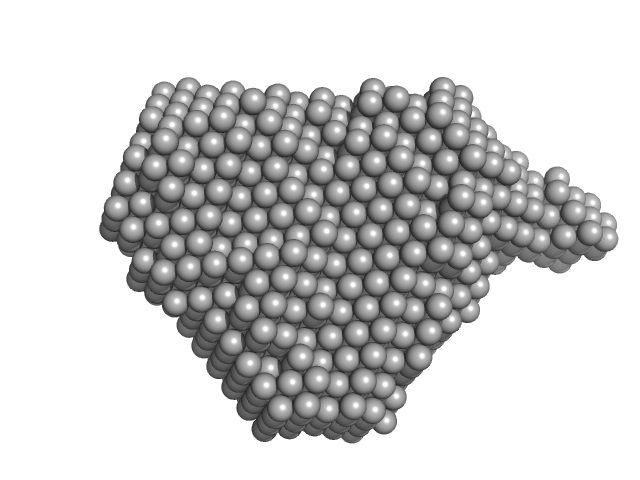
Diguanylate cyclase with PAS/PAC sensor dimer, 27 kDa Marinobacter hydrocarbonoclasticus protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.7 |
nm |
| VolumePorod |
41 |
nm3 |
|
|
UniProt ID: Q24NB0 (None-None) Uncharacterized protein
|
|
|

|
| Sample: |
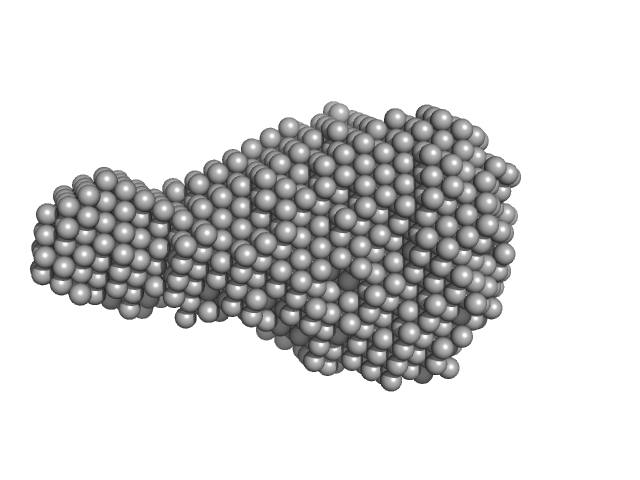
Uncharacterized protein monomer, 9 kDa Desulfitobacterium hafniense protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
1.5 |
nm |
| Dmax |
5.3 |
nm |
| VolumePorod |
13 |
nm3 |
|
|
UniProt ID: Q2Y879 (None-None) Uncharacterized protein
|
|
|

|
| Sample: |
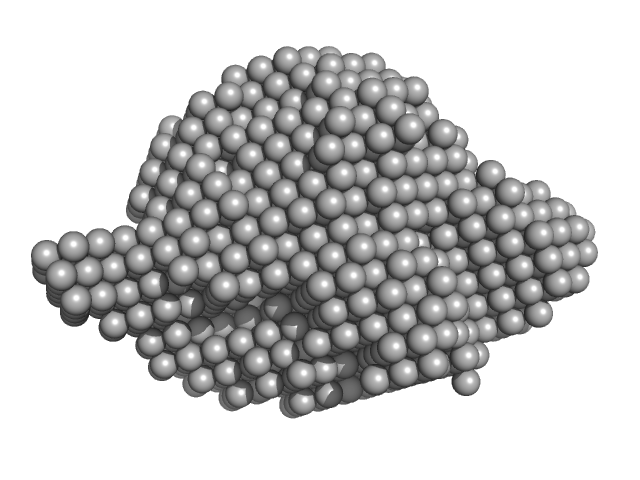
Uncharacterized protein tetramer, 55 kDa Nitrosospira multiformis protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.5 |
nm |
| VolumePorod |
83 |
nm3 |
|
|
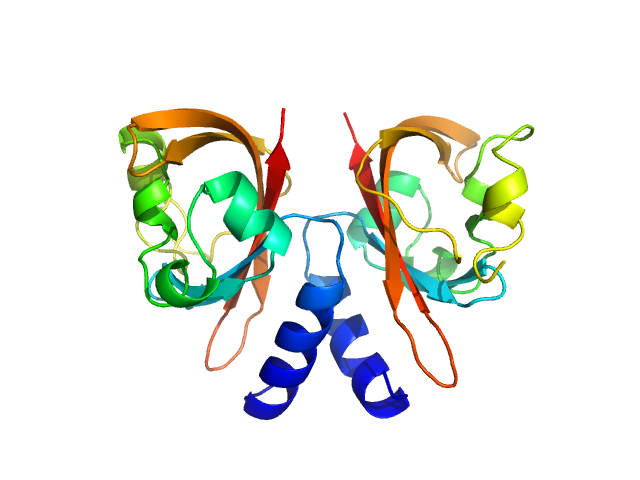
UniProt ID: Q8EGB0 (None-None) Diguanylate cyclase with PAS sensory domain
|
|
|
|
| Sample: |
Diguanylate cyclase with PAS sensory domain dimer, 29 kDa Shewanella oneidensis protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.4 |
nm |
| VolumePorod |
48 |
nm3 |
|
|
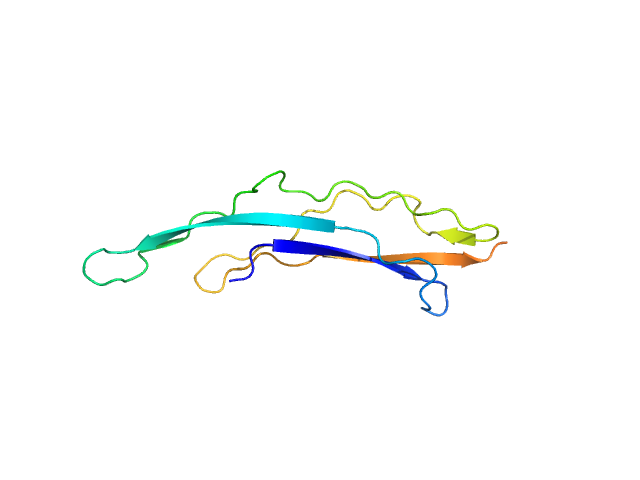
UniProt ID: Q03HU7 (None-None) Adhesion exoprotein
|
|
|
|
| Sample: |
Adhesion exoprotein monomer, 14 kDa Pediococcus pentosaceus protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.2 |
nm |
| VolumePorod |
18 |
nm3 |
|
|
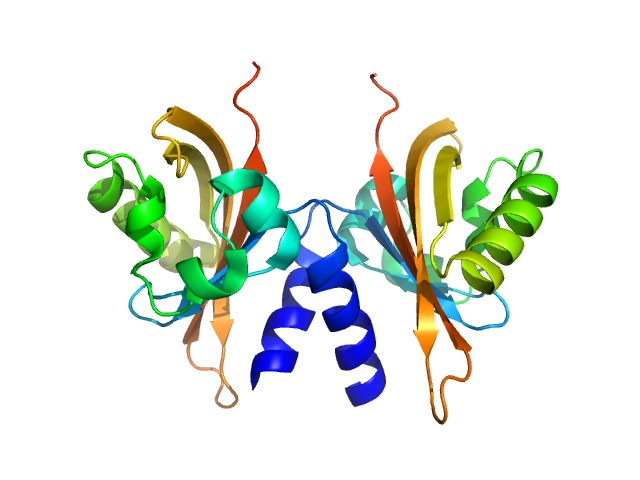
UniProt ID: Q486I1 (None-None) Sensory box/GGDEF domain protein
|
|
|
|
| Sample: |
Sensory box/GGDEF domain protein dimer, 30 kDa Colwellia psychrerythraea protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
52 |
nm3 |
|
|
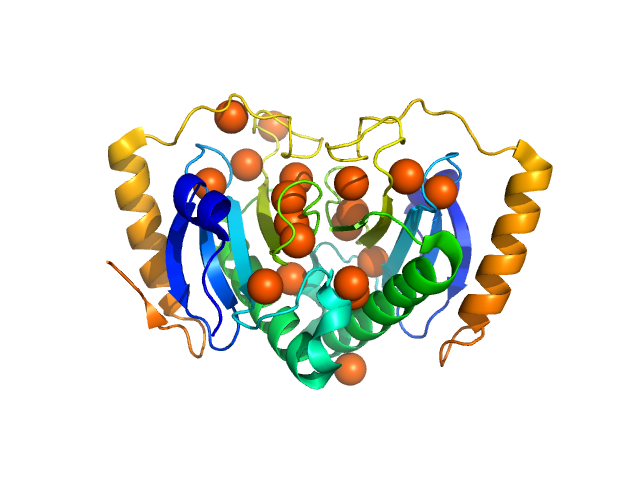
UniProt ID: Q5E3V1 (None-None) HIT family hydrolase
|
|
|
|
| Sample: |
HIT family hydrolase dimer, 34 kDa Aliivibrio fischeri protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.1 |
nm |
| Dmax |
7.2 |
nm |
| VolumePorod |
58 |
nm3 |
|
|
UniProt ID: Q60BX6 (None-None) EAL/GGDEF domain protein
|
|
|
|
| Sample: |
EAL/GGDEF domain protein monomer, 19 kDa Methylococcus capsulatus protein
|
| Buffer: |
5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at BL4-2, Stanford Synchrotron Radiation Lightsource (SSRL) on 2010 Feb 12
|
Small angle X-ray scattering as a complementary tool for high-throughput structural studies.
Biopolymers 95(8):517-30 (2011)
Grant TD, Luft JR, Wolfley JR, Tsuruta H, Martel A, Montelione GT, Snell EH
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.6 |
nm |
| VolumePorod |
34 |
nm3 |
|
|