UniProt ID: Q8U3D2 (5-770) Piwi domain-containing protein
|
|
|
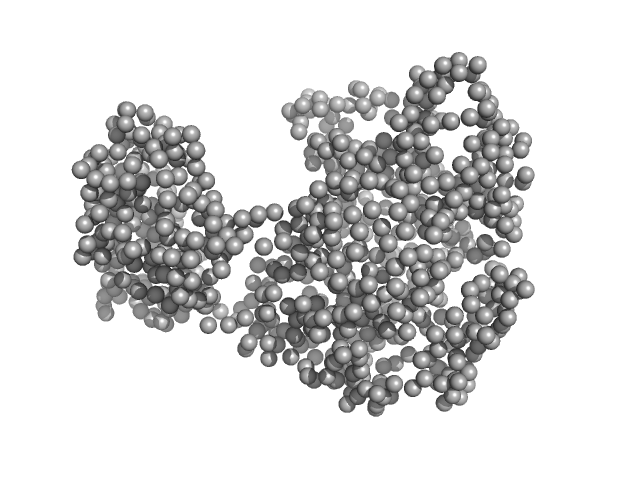
|
| Sample: |
Piwi domain-containing protein monomer, 91 kDa Pyrococcus furiosus (strain … protein
|
| Buffer: |
20 mM Tris–HCl, 250 mM NaCl, 2mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2021 May 7
|
Argonaute protein SAXS investigation
lirong zheng
|
| RgGuinier |
3.2 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
187 |
nm3 |
|
|
UniProt ID: None (719-1706) Mono-acylated variant of a genetic fusion comprising the C-terminal folding scaffold of RTX block V (residues 1562–1681 of CyaA) and the CyaA residues 719–1294
|
|
|
|
| Sample: |
Mono-acylated variant of a genetic fusion comprising the C-terminal folding scaffold of RTX block V (residues 1562–1681 of CyaA) and the CyaA residues 719–1294 monomer, 78 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM NaCl, 3 mM CaCl2, 20 mM Tris, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2022 Feb 17
|
Acyl chains stabilize the acylated domain and determine the receptor-mediated interaction of the Bordetella adenylate cyclase toxin with cell membrane.
J Biol Chem :110392 (2025)
Espinosa-Vinals C, Stransky J, Osicka R, Osickova A, Jurnecka D, Sebo P, Bumba L
|
| RgGuinier |
5.3 |
nm |
| Dmax |
26.3 |
nm |
| VolumePorod |
187 |
nm3 |
|
|
UniProt ID: P0DP23 (1-149) Calmodulin-1
UniProt ID: Q99572 (395-595) P2X purinoceptor 7
|
|
|
|
| Sample: |
Calmodulin-1 monomer, 17 kDa Homo sapiens protein
P2X purinoceptor 7 monomer, 25 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES pH 7.5, 150 mM NaCl, 5 mM CaCl2, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Jun 4
|
New insights into P2X7 receptor regulation: Ca2+-calmodulin and GDP bind to the soluble P2X7 ballast domain.
J Biol Chem :102495 (2022)
Sander S, Müller I, Alai MG, Nicke A, Tidow H
|
| RgGuinier |
3.2 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
73 |
nm3 |
|
|
UniProt ID: A0A1L3NFS6 (1-400) L-methionine gamma-lyase (K272S)
|
|
|
|
| Sample: |
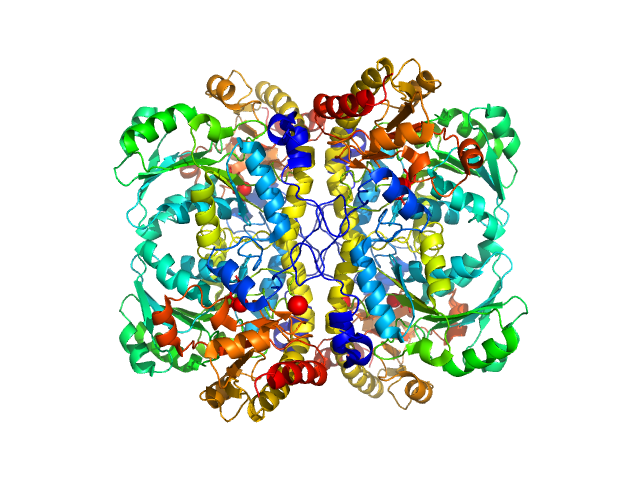
L-methionine gamma-lyase (K272S) tetramer, 174 kDa Clostridium sporogenes protein
|
| Buffer: |
PBS-D2O: 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4 (D2O buffer), pH: 7.4 |
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 May 19
|
Methionine gamma lyase fused with S3 domain VGF forms octamers and adheres to tumor cells via binding EGFR
Biochemical and Biophysical Research Communications :149319 (2023)
Bondarev N, Bagaeva D, Bazhenov S, Buben M, Bulushova N, Ryzhykau Y, Okhrimenko I, Zagryadskaya Y, Maslov I, Anisimova N, Sokolova D, Kuklin A, Pokrovsky V, Manukhov I
|
| RgGuinier |
3.7 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
211 |
nm3 |
|
|
UniProt ID: P9WJD9 (2-348) ESX-1 secretion-associated protein EspB
|
|
|
|
| Sample: |
ESX-1 secretion-associated protein EspB heptamer, 261 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM Tris-HCl, 300 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Sep 12
|
The crystal structure of the EspB-EspK virulence factor-chaperone complex suggests an additional type VII secretion mechanism in M. tuberculosis.
J Biol Chem :102761 (2022)
Gijsbers A, Eymery M, Gao Y, Menart I, Vinciauskaite V, Siliqi D, Peters PJ, McCarthy A, Ravelli RBG
|
| RgGuinier |
5.9 |
nm |
| Dmax |
18.8 |
nm |
| VolumePorod |
911 |
nm3 |
|
|
UniProt ID: None (None-None) Custom 28 base pair double stranded DNA
UniProt ID: P0DW33 (2-416) DNA-guanine transglycosylase - D95A mutant
|
|
|
|
| Sample: |
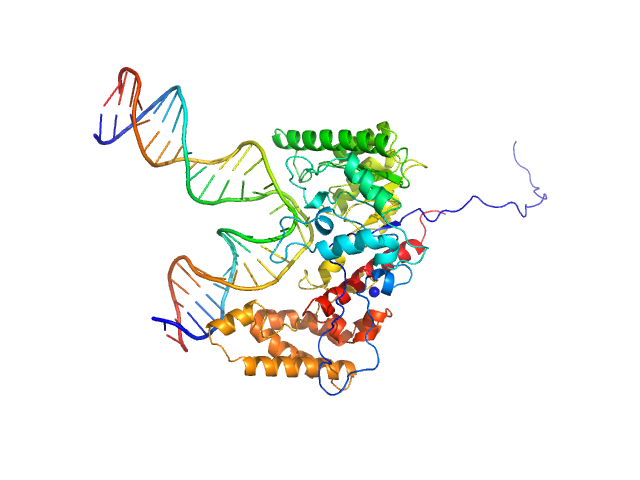
Custom 28 base pair double stranded DNA monomer, 17 kDa Synthetic, purchased from … DNA
DNA-guanine transglycosylase - D95A mutant monomer, 50 kDa Salmonella enterica subsp. … protein
|
| Buffer: |
100 mM KCl, 50 mM Tris pH 7.0, 1 mM DTT, pH: 7 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Dec 22
|
7-Deazaguanines in DNA: functional and structural elucidation of a DNA modification system.
Nucleic Acids Res (2023)
Gedara SH, Wood E, Gustafson A, Liang C, Hung SH, Savage J, Phan P, Luthra A, de Crécy-Lagard V, Dedon P, Swairjo MA, Iwata-Reuyl D
|
| RgGuinier |
3.4 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
104 |
nm3 |
|
|
UniProt ID: P22363 (53-297) Phosphoprotein (Isoform P3; K214A, R260A, D289N, C297S)
|
|
|
|
| Sample: |
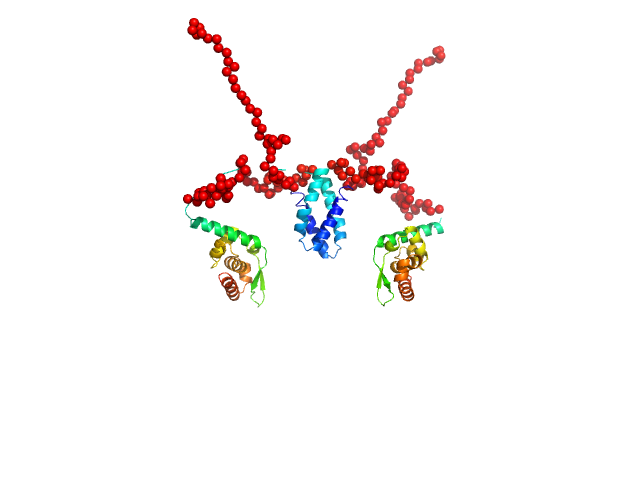
Phosphoprotein (Isoform P3; K214A, R260A, D289N, C297S) dimer, 55 kDa Rabies virus (strain … protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 1 mM TCEP, pH: 7.4 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2021 Mar 7
|
Conformational dynamics, RNA binding, and phase separation regulate the multifunctionality of rabies virus P protein.
Nat Commun 16(1):9491 (2025)
Rawlinson SM, Chakraborty S, Sethi A, David CT, Harrison AR, Bird LE, Rozario AM, Uthishtran S, Ardipradja K, Zhao T, Oksayan S, Jans DA, Ang CS, Lu ZH, Yan F, Williamson NA, Arumugam S, Sundaramoorthy V, Bell TDM, Gooley PR, Moseley GW
|
| RgGuinier |
4.6 |
nm |
| Dmax |
22.5 |
nm |
| VolumePorod |
180 |
nm3 |
|
|
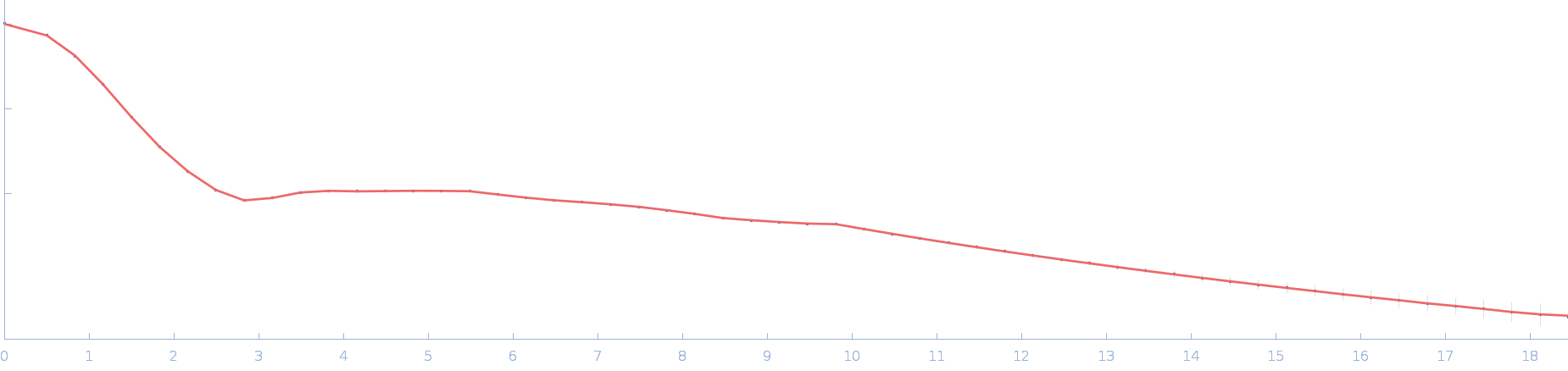
UniProt ID: P73129 (1-96) Ssr1698 protein
|
|
|
|
| Sample: |
Ssr1698 protein dimer, 23 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Hepes, 200 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2022 Sep 27
|
A hemoprotein with a zinc-mirror heme site ties heme availability to carbon metabolism in cyanobacteria.
Nat Commun 15(1):3167 (2024)
Grosjean N, Yee EF, Kumaran D, Chopra K, Abernathy M, Biswas S, Byrnes J, Kreitler DF, Cheng JF, Ghosh A, Almo SC, Iwai M, Niyogi KK, Pakrasi HB, Sarangi R, van Dam H, Yang L, Blaby IK, Blaby-Haas CE
|
| RgGuinier |
2.0 |
nm |
| Dmax |
6.3 |
nm |
| VolumePorod |
29 |
nm3 |
|
|
UniProt ID: B7UM99 (76-180) Translocated intimin receptor Tir
|
|
|
|
| Sample: |
Translocated intimin receptor Tir monomer, 12 kDa Escherichia coli O127:H6 … protein
|
| Buffer: |
100 mM Tris-HCl, 150 mM NaCl, 1 mM EDTA, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Nov 21
|
The pathogen-encoded signalling receptor Tir exploits host-like intrinsic disorder for infection.
Commun Biol 7(1):179 (2024)
Vieira MFM, Hernandez G, Zhong Q, Arbesú M, Veloso T, Gomes T, Martins ML, Monteiro H, Frazão C, Frankel G, Zanzoni A, Cordeiro TN
|
| RgGuinier |
1.7 |
nm |
| Dmax |
7.0 |
nm |
| VolumePorod |
21 |
nm3 |
|
|
UniProt ID: Q9NRY4 (1083-1111) Rho GTPase-activating protein 35
UniProt ID: P20936 (174-444) Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S)
|
|
|
|
| Sample: |
Rho GTPase-activating protein 35 monomer, 3 kDa Homo sapiens protein
Ras GTPase-activating protein 1 (C236S, C261S, C372S, C402S) monomer, 31 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8, 350 mM NaCl, 1 mM DTT, pH: 8 |
| Experiment: |
SAXS
data collected at BioCAT 18ID, Advanced Photon Source (APS), Argonne National Laboratory on 2020 Dec 12
|
Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP and Dok1
Journal of Biological Chemistry :105098 (2023)
Vish K, Stiegler A, Boggon T
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
44 |
nm3 |
|
|