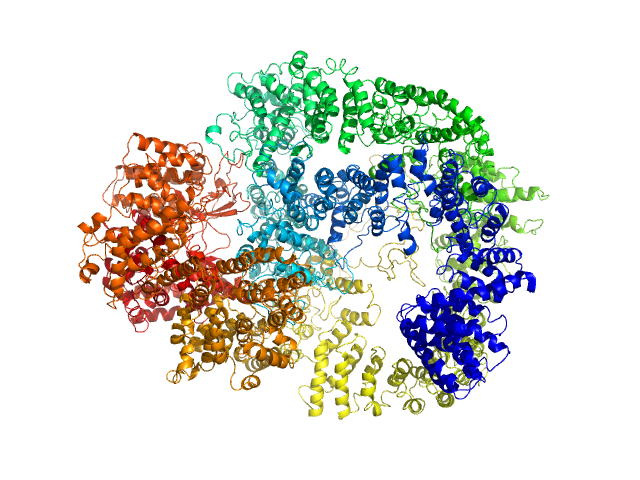
UniProt ID: P78527 (10-4128) DNA-dependent protein kinase catalytic subunit
|
|
|
|
| Sample: |
DNA-dependent protein kinase catalytic subunit monomer, 468 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 100 mM NaCl, 5% glycerol, 0.01% sodium azide, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2016 Dec 30
|
Visualizing functional dynamicity in the DNA-dependent protein kinase holoenzyme DNA-PK complex by integrating SAXS with cryo-EM.
Prog Biophys Mol Biol (2020)
Hammel M, Rosenberg DJ, Bierma J, Hura GL, Lees-Miller SP, Tainer JA
|
| RgGuinier |
5.6 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
1040 |
nm3 |
|
|
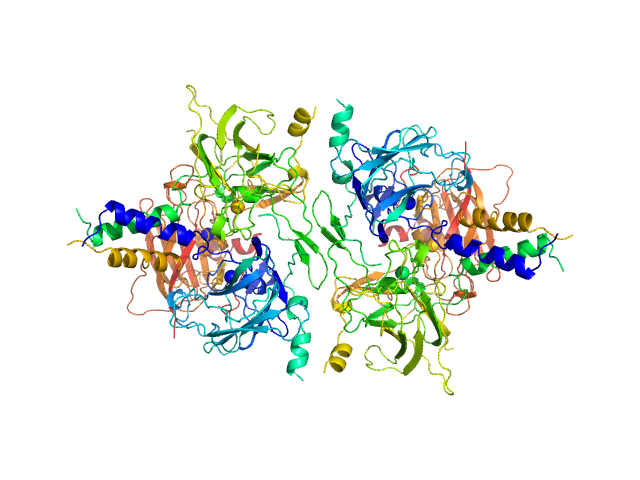
UniProt ID: O94901 (616-812) SUN domain-containing protein 1
UniProt ID: Q8NF91 (8769-8797) Nesprin-1
|
|
|
|
| Sample: |
SUN domain-containing protein 1 hexamer, 135 kDa Homo sapiens protein
Nesprin-1 hexamer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris pH 8.0, 150 mM KCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2017 Dec 17
|
A molecular mechanism for LINC complex branching by structurally diverse SUN-KASH 6:6 assemblies.
Elife 10 (2021)
Gurusaran M, Davies OR
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
254 |
nm3 |
|
|
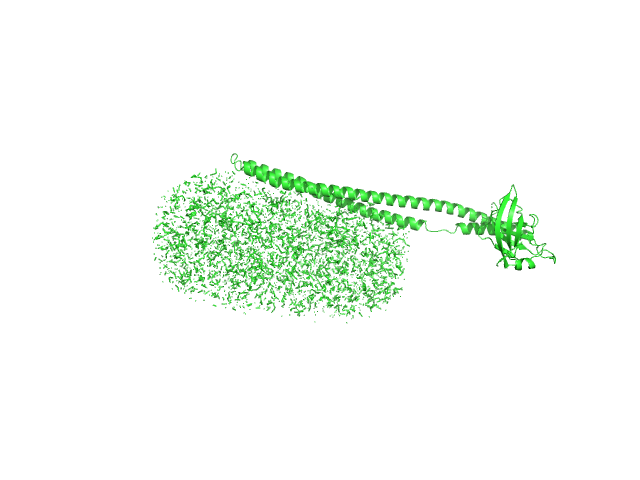
UniProt ID: Q79FZ9 (38-325) Mce-family protein Mce1A
UniProt ID: None (None-None) n-Dodecyl-β-D-Maltopyranoside
|
|
|

|
| Sample: |
Mce-family protein Mce1A monomer, 36 kDa Mycobacterium tuberculosis protein
N-Dodecyl-β-D-Maltopyranoside 0, 82 kDa
|
| Buffer: |
50 mM Tris, 500 mM NaCl, 10% Glycerol, 5 mM DDM, 1 mM β-ME, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 May 2
|
Structural insights into the substrate-binding proteins Mce1A and Mce4A from Mycobacterium tuberculosis
IUCrJ 8(5) (2021)
Asthana P, Singh D, Pedersen J, Hynönen M, Sulu R, Murthy A, Laitaoja M, Jänis J, Riley L, Venkatesan R
|
| RgGuinier |
4.6 |
nm |
| Dmax |
17.5 |
nm |
| VolumePorod |
151 |
nm3 |
|
|
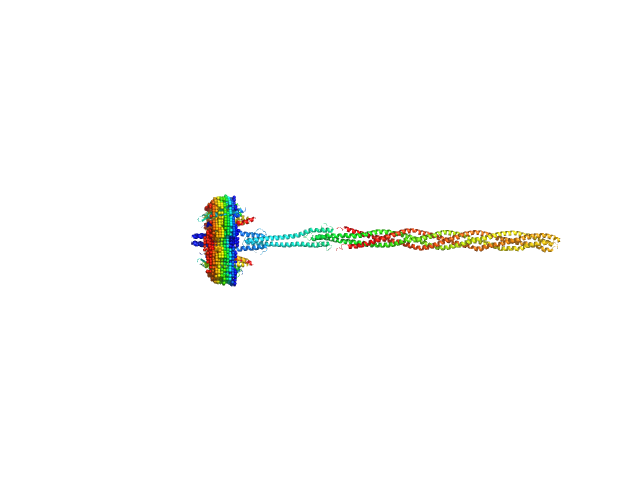
UniProt ID: P42196 (1-239) Sensory rhodopsin II from Natronbacterium pharaonis
UniProt ID: P42259 (3-534) Sensory rhodopsin II transducer from Natronomonas pharaonis
|
|
|

|
| Sample: |
Sensory rhodopsin II from Natronbacterium pharaonis dimer, 53 kDa Natronomonas pharaonis protein
Sensory rhodopsin II transducer from Natronomonas pharaonis dimer, 116 kDa Natronomonas pharaonis protein
|
| Buffer: |
1400 mM NaCl, 49.4 mM Na/Na-Pi, 1.0 mM EDTA, 0.05% DDM (D2O buffer), pH: 8 |
| Experiment: |
SANS
data collected at YuMO SANS TOF spectrometer, IBR-2, Frank Laboratory of Neutron Physics, Joint Institute for Nuclear Research on 2019 Feb 10
|
Molecular model of a sensor of two-component signaling system
Scientific Reports 11(1) (2021)
Ryzhykau Y, Orekhov P, Rulev M, Vlasov A, Melnikov I, Volkov D, Nikolaev M, Zabelskii D, Murugova T, Chupin V, Rogachev A, Gruzinov A, Svergun D, Brennich M, Gushchin I, Soler-Lopez M, Bothe A, Büldt G, Leonard G, Engelhard M, Kuklin A, Gordeliy V
|
| RgGuinier |
9.9 |
nm |
| Dmax |
36.5 |
nm |
|
|
UniProt ID: P25440 (71-455) Bromodomain-containing protein 2
|
|
|
|
| Sample: |
Bromodomain-containing protein 2 monomer, 43 kDa Homo sapiens protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 2% glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Sep 25
|
Multivalent nucleosome scaffolding by bromodomain and extraterminal domain tandem bromodomains.
J Biol Chem :108289 (2025)
Olp MD, Bursch KL, Wynia-Smith SL, Nuñez R, Goetz CJ, Jackson V, Smith BC
|
| RgGuinier |
4.7 |
nm |
| Dmax |
18.8 |
nm |
| VolumePorod |
80 |
nm3 |
|
|
UniProt ID: B7UM99 (388-550) Translocated intimin receptor Tir
|
|
|
|
| Sample: |
Translocated intimin receptor Tir monomer, 18 kDa Escherichia coli O127:H6 … protein
|
| Buffer: |
20 mM Sodium Phosphate, 150 mM NaCl, 1 mM EDTA, pH: 6.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Sep 15
|
The pathogen-encoded signalling receptor Tir exploits host-like intrinsic disorder for infection.
Commun Biol 7(1):179 (2024)
Vieira MFM, Hernandez G, Zhong Q, Arbesú M, Veloso T, Gomes T, Martins ML, Monteiro H, Frazão C, Frankel G, Zanzoni A, Cordeiro TN
|
| RgGuinier |
3.8 |
nm |
| Dmax |
12.8 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
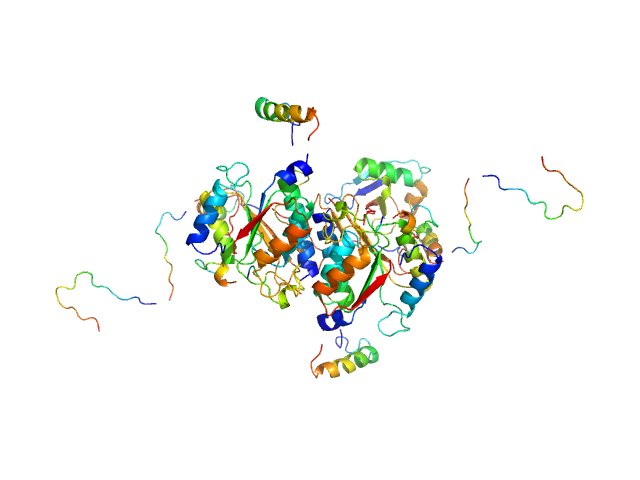
UniProt ID: Q9GZZ9 (57-346) Ubiquitin-like modifier-activating enzyme 5
UniProt ID: P61960 (1-83) Ubiquitin fold modifer 1
|
|
|
|
| Sample: |
Ubiquitin-like modifier-activating enzyme 5 dimer, 68 kDa Homo sapiens protein
Ubiquitin fold modifer 1 monomer, 9 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, 2 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Oct 29
|
Structure and dynamics of UBA5-UFM1 complex formation showing new insights in the UBA5 activation mechanism
Journal of Structural Biology :107796 (2021)
Fuchs S, Kikhney A, Schubert R, Kaiser C, Liebau E, Svergun D, Betzel C, Perbandt M
|
| RgGuinier |
3.1 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
100 |
nm3 |
|
|
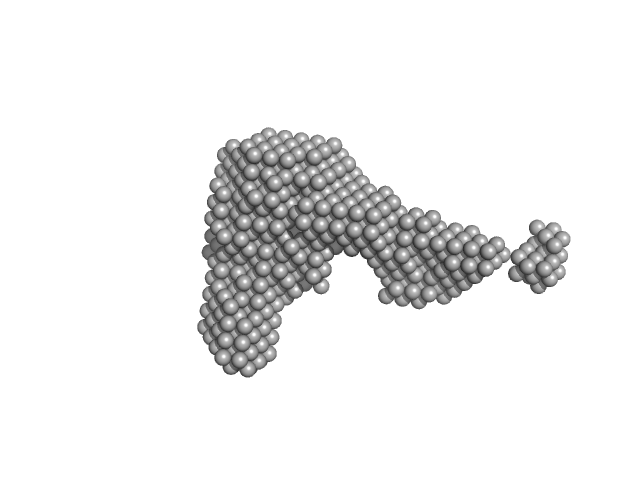
UniProt ID: Q14626 (23-319) Interleukin-11 receptor subunit alpha
UniProt ID: None (None-None) Interleukin 11 Mutein
UniProt ID: P40189 (22-324) Interleukin-6 receptor subunit beta
|
|
|

|
| Sample: |
Interleukin-11 receptor subunit alpha monomer, 32 kDa Homo sapiens protein
Interleukin 11 Mutein monomer, 18 kDa Homo sapiens protein
Interleukin-6 receptor subunit beta monomer, 35 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, 0.2% sodium azide, pH: 8.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2019 Jun 8
|
Structures of the interleukin 11 signalling complex reveal gp130 dynamics and the inhibitory mechanism of a cytokine variant
Nature Communications 14(1) (2023)
Metcalfe R, Hanssen E, Fung K, Aizel K, Kosasih C, Zlatic C, Doughty L, Morton C, Leis A, Parker M, Gooley P, Putoczki T, Griffin M
|
| RgGuinier |
4.4 |
nm |
| Dmax |
15.6 |
nm |
| VolumePorod |
160 |
nm3 |
|
|
UniProt ID: P02768 (None-None) Albumin
|
|
|
|
| Sample: |
Albumin monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 150 mM KCl, 2% glycerol, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2020 Dec 1
|
Albumin in patients with liver disease shows an altered conformation.
Commun Biol 4(1):731 (2021)
Paar M, Fengler VH, Rosenberg DJ, Krebs A, Stauber RE, Oettl K, Hammel M
|
| RgGuinier |
2.8 |
nm |
| Dmax |
8.1 |
nm |
|
|
UniProt ID: Q45589 (97-273) Cyclic di-AMP synthase CdaA
|
|
|
|
| Sample: |
Cyclic di-AMP synthase CdaA dimer, 44 kDa Bacillus subtilis (strain … protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Jul 21
|
Structural basis for the inhibition of the Bacillus subtilis c-di-AMP cyclase CdaA by the phosphoglucomutase GlmM
Journal of Biological Chemistry :101317 (2021)
Pathania M, Tosi T, Millership C, Hoshiga F, Morgan R, Freemont P, Gründling A
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
60 |
nm3 |
|
|