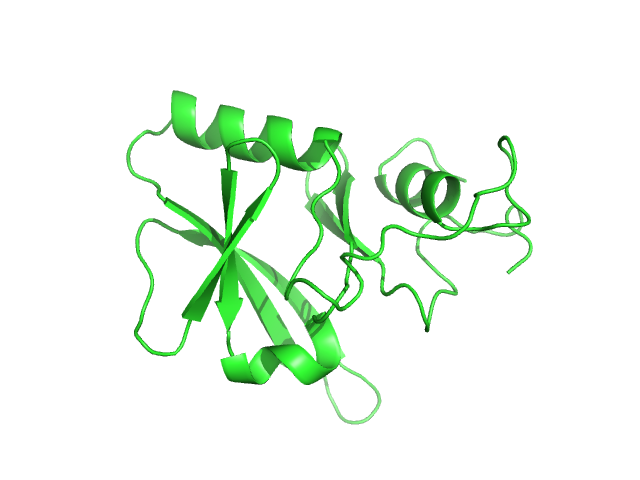
UniProt ID: P0DN75 (2-132) Iron-sulfur cluster assembly 1 homolog, mitochondrial
|
|
|
|
| Sample: |
Iron-sulfur cluster assembly 1 homolog, mitochondrial, 15 kDa Columba livia protein
|
| Buffer: |
20 mM Tris-HCl, 0.15 M NaCl, 10 mM 3-mercapto-1,2-propanediol, pH: 8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2020 Feb 25
|
Magnetic field effects on the structure and molecular behavior of pigeon iron–sulfur protein
Protein Science 31(6) (2022)
Arai S, Shimizu R, Adachi M, Hirai M
|
|
|
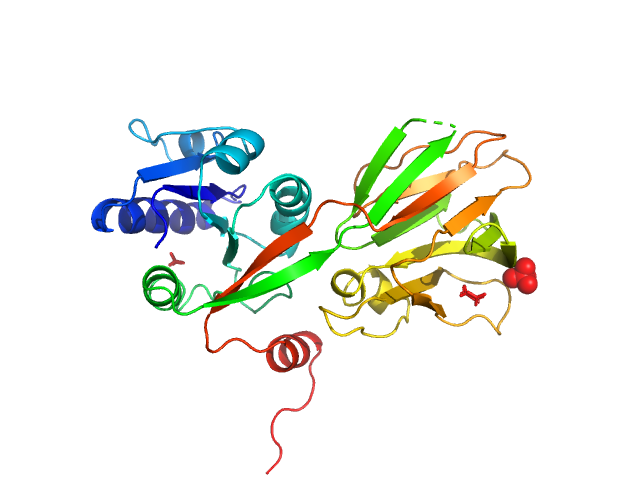
UniProt ID: P0A7B3 (1-292) NAD kinase
|
|
|
|
| Sample: |
NAD kinase, 33 kDa Escherichia coli (strain … protein
|
| Buffer: |
50 mM HEPES, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Oct 2
|
Protein quaternary structures in solution are a mixture of multiple forms
Chemical Science 13(39):11680-11695 (2022)
Marciano S, Dey D, Listov D, Fleishman S, Sonn-Segev A, Mertens H, Busch F, Kim Y, Harvey S, Wysocki V, Schreiber G
|
| RgGuinier |
4.0 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
260 |
nm3 |
|
|
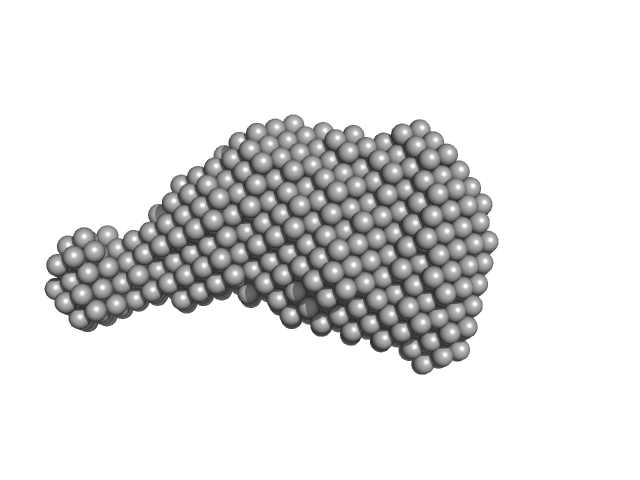
UniProt ID: E0IW31 (253-448) Accessory colonization factor
|
|
|

|
| Sample: |
Accessory colonization factor monomer, 23 kDa Escherichia coli (strain … protein
|
| Buffer: |
20 mM Tris, 200 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2021 Apr 21
|
Molecular and cellular insight into Escherichia coli SslE and its role during biofilm maturation
npj Biofilms and Microbiomes 8(1) (2022)
Corsini P, Wang S, Rehman S, Fenn K, Sagar A, Sirovica S, Cleaver L, Edwards-Gayle C, Mastroianni G, Dorgan B, Sewell L, Lynham S, Iuga D, Franks W, Jarvis J, Carpenter G, Curtis M, Bernadó P, Darbari V, Garnett J
|
| RgGuinier |
2.2 |
nm |
| Dmax |
7.7 |
nm |
| VolumePorod |
42 |
nm3 |
|
|
UniProt ID: P9WJD9 (7-278) ESX-1 secretion-associated protein EspB
|
|
|
|
| Sample: |
ESX-1 secretion-associated protein EspB monomer, 30 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM Tris-HCl, 300 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Sep 21
|
The crystal structure of the EspB-EspK virulence factor-chaperone complex suggests an additional type VII secretion mechanism in M. tuberculosis.
J Biol Chem :102761 (2022)
Gijsbers A, Eymery M, Gao Y, Menart I, Vinciauskaite V, Siliqi D, Peters PJ, McCarthy A, Ravelli RBG
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.8 |
nm |
| VolumePorod |
56 |
nm3 |
|
|
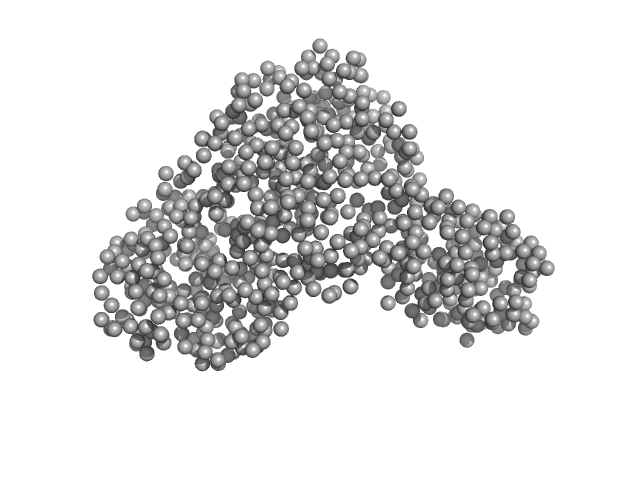
UniProt ID: P02787 (1-698) Serotransferrin
|
|
|

|
| Sample: |
Serotransferrin monomer, 77 kDa Homo sapiens protein
|
| Buffer: |
15 mM HEPES, 20 mM NaHCO3, 50 mM NaCl (Buffer APO-Tf-1), pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 7
|
X-ray Characterization of Conformational Changes of Human Apo- and Holo-Transferrin
International Journal of Molecular Sciences 22(24):13392 (2021)
Campos-Escamilla C, Siliqi D, Gonzalez-Ramirez L, Lopez-Sanchez C, Gavira J, Moreno A
|
| RgGuinier |
3.2 |
nm |
| Dmax |
13.1 |
nm |
| VolumePorod |
106 |
nm3 |
|
|
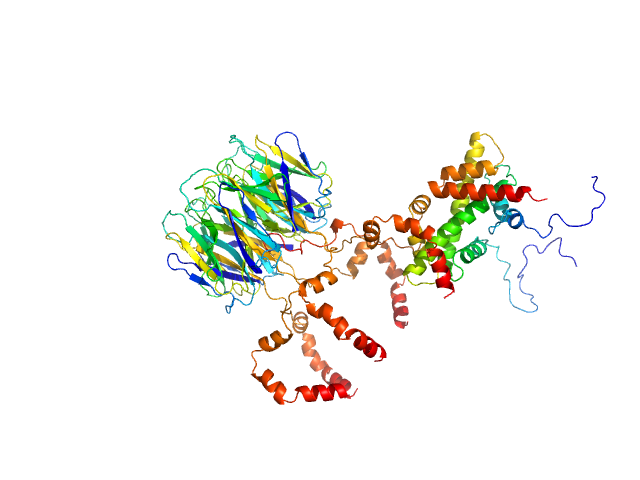
UniProt ID: F4J9Q6 (1-136) Peptidyl-prolyl cis-trans isomerase FKBP43
UniProt ID: P04908 (1-130) Histone H2A type 1-B/E
UniProt ID: P62807 (1-126) Histone H2B type 1-C/E/F/G/I
|
|
|
|
| Sample: |
Peptidyl-prolyl cis-trans isomerase FKBP43 pentamer, 81 kDa Arabidopsis thaliana protein
Histone H2A type 1-B/E monomer, 14 kDa Homo sapiens protein
Histone H2B type 1-C/E/F/G/I monomer, 14 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 300 mM NaCl, 1 mM β-mercaptoethanol, pH: 7.5 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Apr 28
|
The plant nucleoplasmin AtFKBP43 needs its extended arms for histone interaction.
Biochim Biophys Acta Gene Regul Mech 1865(7):194872 (2022)
Singh AK, Saharan K, Baral S, Vasudevan D
|
| RgGuinier |
4.1 |
nm |
| Dmax |
13.9 |
nm |
| VolumePorod |
194 |
nm3 |
|
|
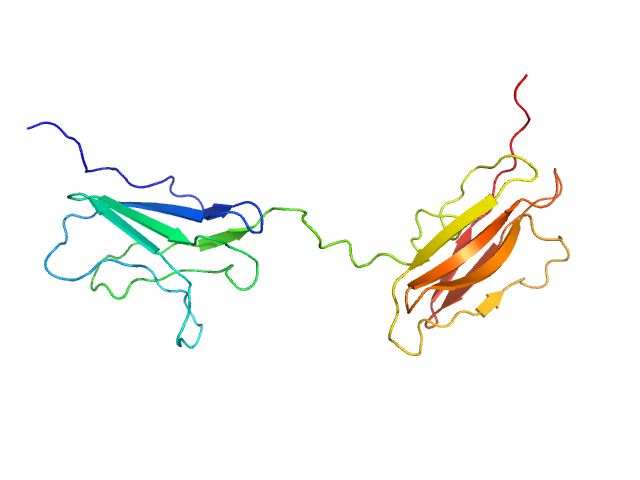
UniProt ID: Q5VST9 (1071-1254) Obscurin Ig domains 12/13 long
|
|
|
|
| Sample: |
Obscurin Ig domains 12/13 long monomer, 22 kDa Homo sapiens protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, 0.35 mM NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2021 Oct 20
|
The N-terminus of obscurin is flexible in solution.
Proteins (2022)
Mauriello GE, Moncure GE, Nowzari RA, Miller CJ, Wright NT
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.0 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
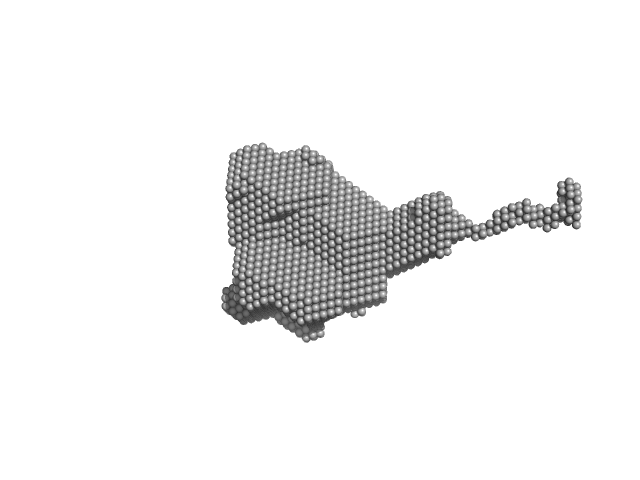
UniProt ID: P74102 (2-317) Orange carotenoid-binding protein
|
|
|

|
| Sample: |
Orange carotenoid-binding protein dimer, 70 kDa Synechocystis sp. (strain … protein
|
| Buffer: |
50 mM Tris, 150 mM NaCL, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 23
|
Oligomerization processes limit photoactivation and recovery of the orange carotenoid protein.
Biophys J 121(15):2849-2872 (2022)
Andreeva EA, Niziński S, Wilson A, Levantino M, De Zitter E, Munro R, Muzzopappa F, Thureau A, Zala N, Burdzinski G, Sliwa M, Kirilovsky D, Schirò G, Colletier JP
|
| RgGuinier |
2.8 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
89 |
nm3 |
|
|
UniProt ID: Q8U3D2 (5-770) Piwi domain-containing protein
|
|
|
|
| Sample: |
Piwi domain-containing protein monomer, 91 kDa Pyrococcus furiosus (strain … protein
|
| Buffer: |
20 mM Tris-HCl, 250 mM NaCl, 2 mM DTT, 6 M urea, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2021 Nov 17
|
Argonaute protein SAXS investigation
lirong zheng
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
148 |
nm3 |
|
|
UniProt ID: None (719-1706) Non-acylated variant of a genetic fusion comprising the C-terminal folding scaffold of RTX block V (residues 1562–1681 of CyaA) and the CyaA residues 719–1294
|
|
|
|
| Sample: |
Non-acylated variant of a genetic fusion comprising the C-terminal folding scaffold of RTX block V (residues 1562–1681 of CyaA) and the CyaA residues 719–1294 monomer, 78 kDa Bordetella pertussis protein
|
| Buffer: |
50 mM NaCl, 3 mM CaCl2, 20 mM Tris, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSpoint 2.0, Institute of Biotechnology, Czech Academy of Sciences/Centre of Molecular Structure on 2022 Feb 22
|
Acyl chains stabilize the acylated domain and determine the receptor-mediated interaction of the Bordetella adenylate cyclase toxin with cell membrane.
J Biol Chem :110392 (2025)
Espinosa-Vinals C, Stransky J, Osicka R, Osickova A, Jurnecka D, Sebo P, Bumba L
|
| RgGuinier |
4.2 |
nm |
| Dmax |
17.8 |
nm |
| VolumePorod |
133 |
nm3 |
|
|