UniProt ID: Q88MI5 (1-91) Putative killer protein, Toxin GraT.
UniProt ID: Q88MI6 (1-99) antitoxin GraA, antidote of toxin GraT
UniProt ID: None (1-91) Putative killer protein, Toxin GraT
|
|
|
|
| Sample: |
Putative killer protein, Toxin GraT. monomer, 11 kDa Pseudomonas putida protein
Antitoxin GraA, antidote of toxin GraT dimer, 22 kDa Pseudomonas putida protein
Putative killer protein, Toxin GraT monomer, 11 kDa Pseudomonas putida protein
|
| Buffer: |
50 mM Tris 250 mM NaCl 2 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 Jun 29
|
A dual role in regulation and toxicity for the disordered N-terminus of the toxin GraT.
Nat Commun 10(1):972 (2019)
Talavera A, Tamman H, Ainelo A, Konijnenberg A, Hadži S, Sobott F, Garcia-Pino A, Hõrak R, Loris R
|
| RgGuinier |
2.3 |
nm |
| Dmax |
7.9 |
nm |
| VolumePorod |
63 |
nm3 |
|
|
UniProt ID: None (None-None) GraTA operator region
UniProt ID: Q88MI6 (1-99) antitoxin GraA, antidote of toxin GraT
UniProt ID: Q88MI6 (1-99) atitoxin GraA, antidote of GraT.
|
|
|
|
| Sample: |
GraTA operator region monomer, 20 kDa DNA
Antitoxin GraA, antidote of toxin GraT dimer, 22 kDa Pseudomonas putida protein
Atitoxin GraA, antidote of GraT. dimer, 23 kDa Pseudomonas putida protein
|
| Buffer: |
50 mM Tris 250 mM NaCl 2 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2017 Dec 17
|
A dual role in regulation and toxicity for the disordered N-terminus of the toxin GraT.
Nat Commun 10(1):972 (2019)
Talavera A, Tamman H, Ainelo A, Konijnenberg A, Hadži S, Sobott F, Garcia-Pino A, Hõrak R, Loris R
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
94 |
nm3 |
|
|
UniProt ID: A0A0H2XIU4 (100-269) Diadenylate cyclase
UniProt ID: Q2FEX1 (None-None) Phosphoglucosamine mutase
|
|
|

|
| Sample: |
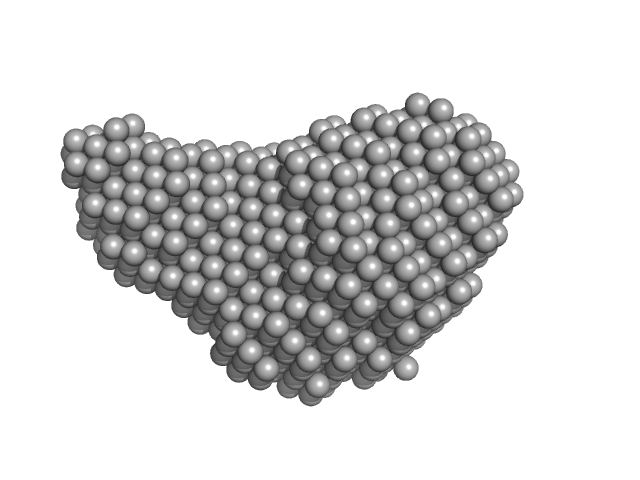
Diadenylate cyclase dimer, 39 kDa Staphylococcus aureus protein
Phosphoglucosamine mutase dimer, 99 kDa Staphylococcus aureus protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 7
|
Inhibition of the Staphylococcus aureus c-di-AMP cyclase DacA by direct interaction with the phosphoglucosamine mutase GlmM.
PLoS Pathog 15(1):e1007537 (2019)
Tosi T, Hoshiga F, Millership C, Singh R, Eldrid C, Patin D, Mengin-Lecreulx D, Thalassinos K, Freemont P, Gründling A
|
| RgGuinier |
3.9 |
nm |
| Dmax |
12.1 |
nm |
| VolumePorod |
204 |
nm3 |
|
|
UniProt ID: Q2FEX1 (None-None) Phosphoglucosamine mutase
|
|
|

|
| Sample: |
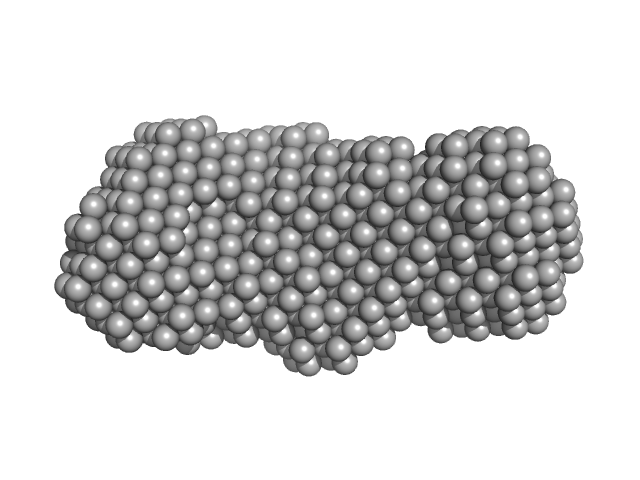
Phosphoglucosamine mutase dimer, 99 kDa Staphylococcus aureus protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 7
|
Inhibition of the Staphylococcus aureus c-di-AMP cyclase DacA by direct interaction with the phosphoglucosamine mutase GlmM.
PLoS Pathog 15(1):e1007537 (2019)
Tosi T, Hoshiga F, Millership C, Singh R, Eldrid C, Patin D, Mengin-Lecreulx D, Thalassinos K, Freemont P, Gründling A
|
| RgGuinier |
3.7 |
nm |
| Dmax |
12.5 |
nm |
| VolumePorod |
134 |
nm3 |
|
|
UniProt ID: A0A0H2XIU4 (100-269) Diadenylate cyclase
|
|
|

|
| Sample: |
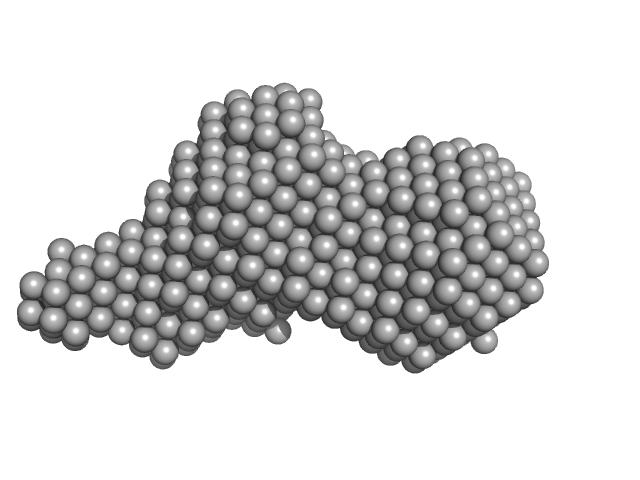
Diadenylate cyclase dimer, 39 kDa Staphylococcus aureus protein
|
| Buffer: |
30 mM Tris, 150 mM NaCl, pH: 7.5 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2018 May 7
|
Inhibition of the Staphylococcus aureus c-di-AMP cyclase DacA by direct interaction with the phosphoglucosamine mutase GlmM.
PLoS Pathog 15(1):e1007537 (2019)
Tosi T, Hoshiga F, Millership C, Singh R, Eldrid C, Patin D, Mengin-Lecreulx D, Thalassinos K, Freemont P, Gründling A
|
| RgGuinier |
2.6 |
nm |
| Dmax |
8.6 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
UniProt ID: P05096 (None-None) DNA primase
|
|
|
|
| Sample: |
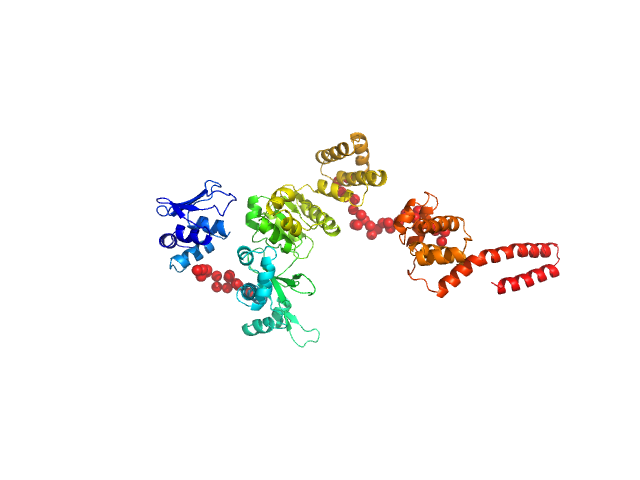
DNA primase monomer, 69 kDa Bacillus subtilis protein
|
| Buffer: |
25 mM Tris-HCl,100 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BL19U2, Shanghai Synchrotron Radiation Facility (SSRF) on 2018 Oct 1
|
Full length of DnaG primase from Bacillus subtilis
Zhongchuan Liu
|
| RgGuinier |
3.7 |
nm |
| Dmax |
14.5 |
nm |
| VolumePorod |
119 |
nm3 |
|
|
UniProt ID: A0A2B4EEI3 (1-210) Bacillus thuringiensis LexA repressor
|
|
|
|
| Sample: |
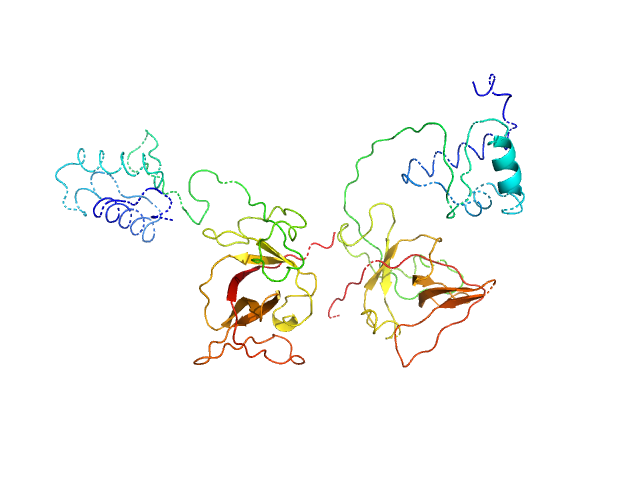
Bacillus thuringiensis LexA repressor dimer, 47 kDa Bacillus thuringiensis protein
|
| Buffer: |
20 mM Hepes, 300 mM NaCl, 10% glycerol,, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, University of British Columbia on 2017 Aug 25
|
Structural Insights into Bacteriophage GIL01 gp7 Inhibition of Host LexA Repressor.
Structure 27(7):1094-1102.e4 (2019)
Caveney NA, Pavlin A, Caballero G, Bahun M, Hodnik V, de Castro L, Fornelos N, Butala M, Strynadka NCJ
|
| RgGuinier |
3.7 |
nm |
| VolumePorod |
110 |
nm3 |
|
|
UniProt ID: A0A2B4EEI3 (1-210) Bacillus thuringiensis LexA repressor
UniProt ID: Q7WSG2 (1-50) Bacteriophage pGIL01 gp7
|
|
|
|
| Sample: |
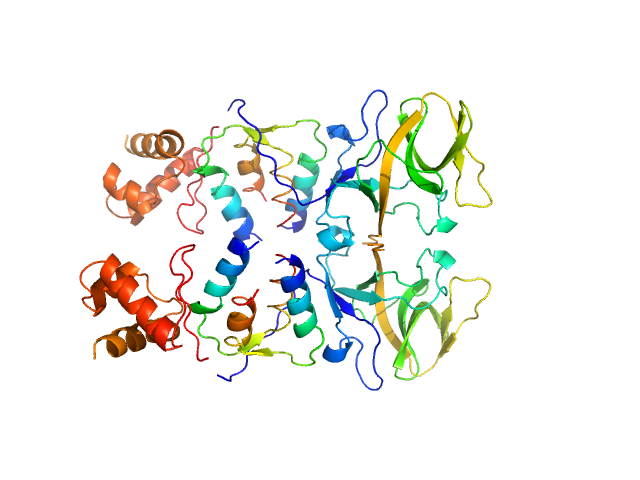
Bacillus thuringiensis LexA repressor dimer, 47 kDa Bacillus thuringiensis protein
Bacteriophage pGIL01 gp7 tetramer, 24 kDa Bacteriophage pGIL01 protein
|
| Buffer: |
20 mM Hepes, 300 mM NaCl, 10% glycerol,, pH: 8 |
| Experiment: |
SAXS
data collected at Rigaku BioSAXS-2000, University of British Columbia on 2017 Aug 25
|
Structural Insights into Bacteriophage GIL01 gp7 Inhibition of Host LexA Repressor.
Structure 27(7):1094-1102.e4 (2019)
Caveney NA, Pavlin A, Caballero G, Bahun M, Hodnik V, de Castro L, Fornelos N, Butala M, Strynadka NCJ
|
|
|
UniProt ID: P32455 (1-592) farnesylated human Guanylate-binding protein 1
|
|
|
|
| Sample: |
Farnesylated human Guanylate-binding protein 1 monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 5 mM MgCl2, 150 mM NaCl, pH: 7.9 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Apr 30
|
Farnesylation of human guanylate binding protein 1 as safety mechanism preventing structural rearrangements and uninduced dimerization.
FEBS J (2019)
Lorenz C, Ince S, Zhang T, Cousin A, Batra-Safferling R, Nagel-Steger L, Herrmann C, Stadler AM
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
102 |
nm3 |
|
|
UniProt ID: P32455 (1-592) human Guanylate-binding protein 1
|
|
|
|
| Sample: |
Human Guanylate-binding protein 1 monomer, 69 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris-HCl, 5 mM MgCl2, 150 mM NaCl, pH: 7.9 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2017 Apr 30
|
Farnesylation of human guanylate binding protein 1 as safety mechanism preventing structural rearrangements and uninduced dimerization.
FEBS J (2019)
Lorenz C, Ince S, Zhang T, Cousin A, Batra-Safferling R, Nagel-Steger L, Herrmann C, Stadler AM
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.4 |
nm |
| VolumePorod |
106 |
nm3 |
|
|