UniProt ID: P24300 (None-None) Xylose isomerase
|
|
|

|
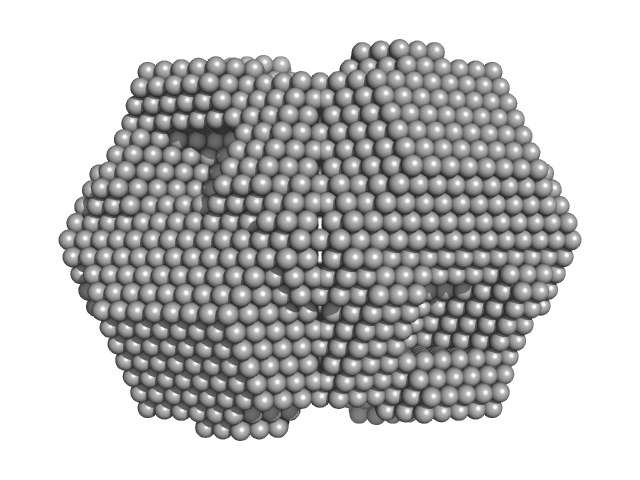
| Sample: |
Xylose isomerase tetramer, 172 kDa Streptomyces rubiginosus protein
|
| Buffer: |
PBS, 50% Glycerol, 0.076 M NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at EMBL X33, DORIS III, DESY on 2012 Apr 24
|
Standard proteins
Erica Valentini
|
| RgGuinier |
3.4 |
nm |
| Dmax |
9.7 |
nm |
| VolumePorod |
293 |
nm3 |
|
|
UniProt ID: P24300 (1-388) Xylose isomerase
|
|
|

|
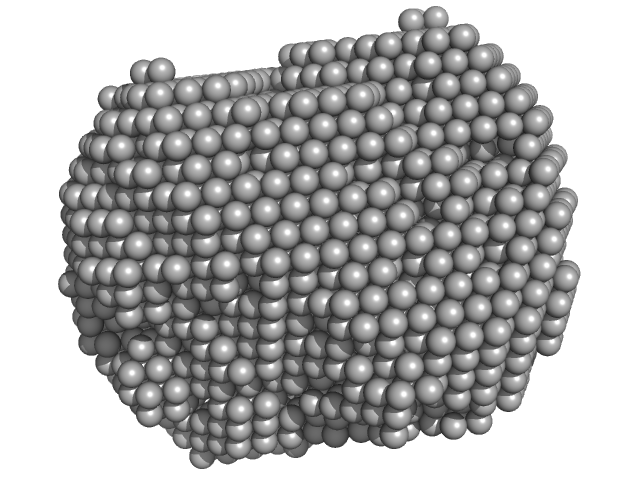
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
25 mM MOPS, 250 mM NaCl, 50 mM KCl, 2 mM TCEP, 0.1% NaN3, pH: 7.5 |
| Experiment: |
SAXS
data collected at SAXS/WAXS, Australian Synchrotron on 2017 Mar 9
|
2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update.
Acta Crystallogr D Struct Biol 73(Pt 9):710-728 (2017)
Trewhella J, Duff AP, Durand D, Gabel F, Guss JM, Hendrickson WA, Hura GL, Jacques DA, Kirby NM, Kwan AH, Pérez J, Pollack L, Ryan TM, Sali A, Schneidman-Duhovny D, Schwede T, Svergun DI, Sugiyama M, Tainer JA, Vachette P, Westbrook J, Whitten AE
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
229 |
nm3 |
|
|
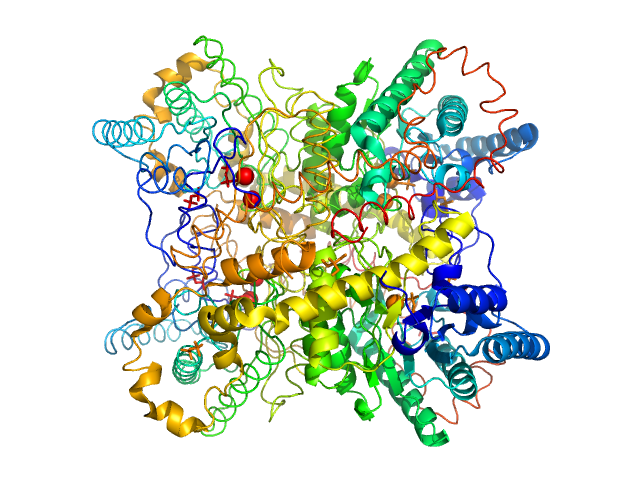
UniProt ID: P24300 (None-None) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 172 kDa Streptomyces rubiginosus protein
|
| Buffer: |
100 mM tris pH 8.0, 100 mM NaCl, 1 mM MgCl2, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Oct 17
|
WAXS benchmark on standard proteins
Maxim Petoukhov
|
|
|
UniProt ID: P24300 (None-None) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
100 mM HEPES, 1 mM MgCl2, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Dec 21
|
A high flux setup for millisecond-scale small-angle X-ray scattering studies on macromolecular solutions
Clement Blanchet
|
|
|
UniProt ID: P24300 (1-388) Xylose isomerase
|
|
|

|
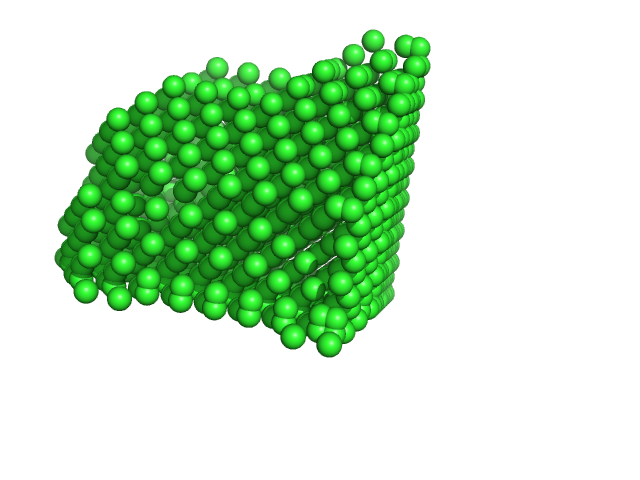
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
ConsensusBuffer_50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
3.3 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
243 |
nm3 |
|
|
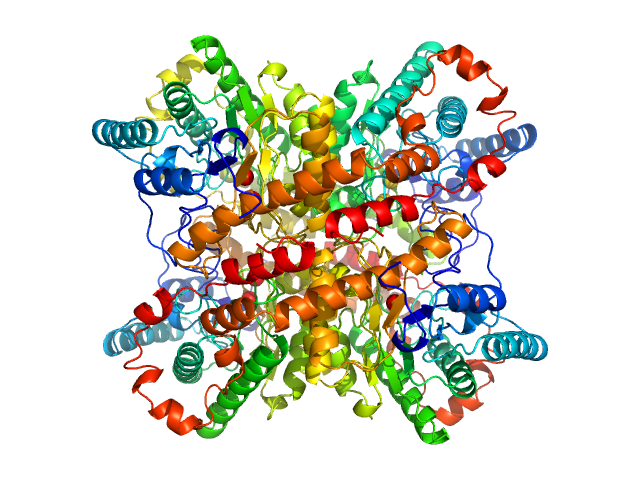
UniProt ID: P24300 (1-388) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 6
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
3.1 |
nm |
| Dmax |
9.5 |
nm |
|
|
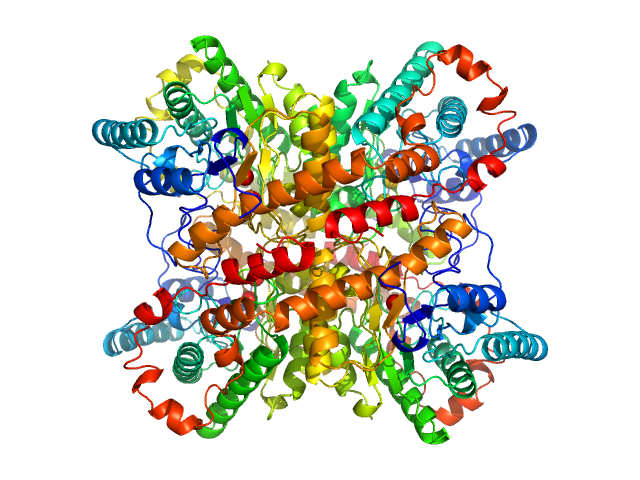
UniProt ID: P24300 (1-388) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 1 mM MgCl2, pH: 7.5 |
| Experiment: |
SANS
data collected at (Consensus SAS), Multi-facility, Multiple countries on 2022 Jun 24
|
A round-robin approach provides a detailed assessment of biomolecular small-angle scattering data reproducibility and yields consensus curves for benchmarking
Acta Crystallographica Section D Structural Biology 78(11) (2022)
Trewhella J, Vachette P, Bierma J, Blanchet C, Brookes E, Chakravarthy S, Chatzimagas L, Cleveland T, Cowieson N, Crossett B, Duff A, Franke D, Gabel F, Gillilan R, Graewert M, Grishaev A, Guss J, Hammel M, Hopkins J, Huang Q, Hub J, Hura G, Irving T, Jeffries C, Jeong C, Kirby N, Krueger S, Martel A, Matsui T, Li N, Pérez J, Porcar L, Prangé T, Rajkovic I, Rocco M, Rosenberg D, Ryan T, Seifert S, Sekiguchi H, Svergun D, Teixeira S, Thureau A, Weiss T, Whitten A, Wood K, Zuo X
|
| RgGuinier |
3.3 |
nm |
| Dmax |
9.7 |
nm |
|
|
UniProt ID: P24300 (1-380) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2020 Mar 7
|
Inline small‐angle X‐ray scattering‐coupled chromatography under extreme hydrostatic pressure
Protein Science 31(12) (2022)
Miller R, Cummings C, Huang Q, Ando N, Gillilan R
|
| RgGuinier |
3.4 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
253 |
nm3 |
|
|
UniProt ID: P24300 (1-380) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
25 mM HEPES, 150 mM NaCl, and 3% v/v glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at ID7A1 BioSAXS / HP-Bio Beamline, Cornell High Energy Synchrotron Source (CHESS) on 2020 Mar 7
|
Inline small‐angle X‐ray scattering‐coupled chromatography under extreme hydrostatic pressure
Protein Science 31(12) (2022)
Miller R, Cummings C, Huang Q, Ando N, Gillilan R
|
| RgGuinier |
3.5 |
nm |
| Dmax |
10.6 |
nm |
| VolumePorod |
267 |
nm3 |
|
|
UniProt ID: P24300 (1-388) Xylose isomerase
|
|
|
|
| Sample: |
Xylose isomerase tetramer, 173 kDa Streptomyces rubiginosus protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 1 mM MgCl2, 1 % v/v glycerol, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2019 Sep 15
|
Small-Angle X-ray Scattering data (benchmarking/consensus): EMBL-P12 SAXS beam line, DESY
Clement Blanchet,
Melissa Graewert,
Cy M Jeffries,
Dmitri Svergun
|
| RgGuinier |
3.2 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
228 |
nm3 |
|
|