|
|
|
|
|
| Sample: |
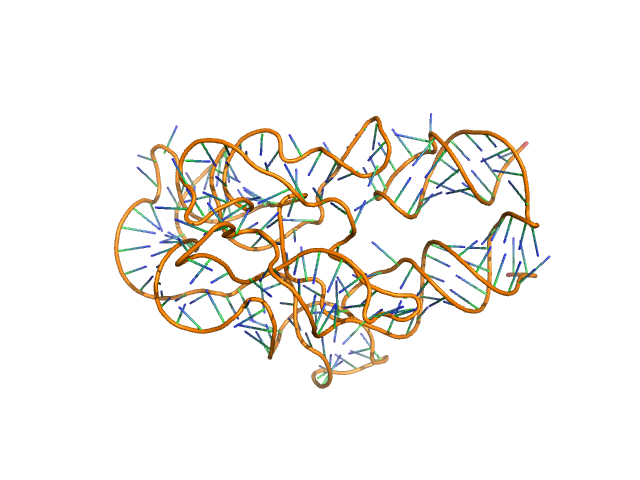
Thermoanearobacter tengcongensis (Tte) fecB riboswitch aptamer domain, 68 kDa marine metagenome RNA
|
| Buffer: |
50 mM MES pH 6.0, 10 mM KCl, 1 mM MgCl2, pH: 6 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Apr 18
|
Visualizing RNA conformational and architectural heterogeneity in solution.
Nat Commun 14(1):714 (2023)
Ding J, Lee YT, Bhandari Y, Schwieters CD, Fan L, Yu P, Tarosov SG, Stagno JR, Ma B, Nussinov R, Rein A, Zhang J, Wang YX
|
| RgGuinier |
5.9 |
nm |
| Dmax |
20.7 |
nm |
| VolumePorod |
206 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
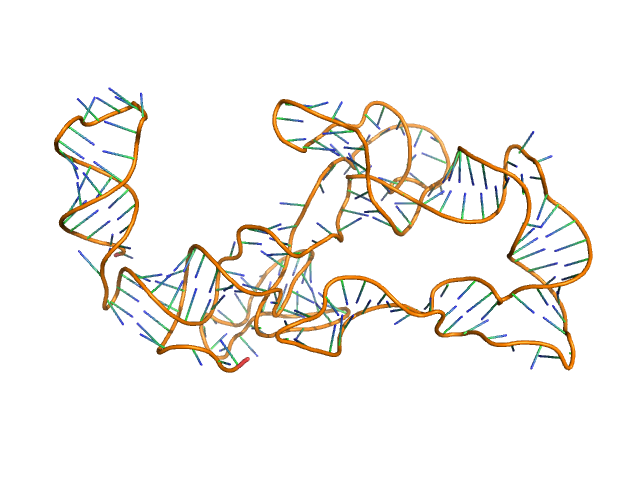
Thermoanearobacter tengcongensis (Tte) fecB riboswitch aptamer domain, 68 kDa marine metagenome RNA
|
| Buffer: |
50 mM MES pH 6.0, 10 mM KCl, 1 mM MgCl2, 4-18 μM coenzyme B12 ligand, pH: 6 |
| Experiment: |
SAXS
data collected at 12-ID-B SAXS/WAXS, Advanced Photon Source (APS), Argonne National Laboratory on 2021 Apr 18
|
Visualizing RNA conformational and architectural heterogeneity in solution.
Nat Commun 14(1):714 (2023)
Ding J, Lee YT, Bhandari Y, Schwieters CD, Fan L, Yu P, Tarosov SG, Stagno JR, Ma B, Nussinov R, Rein A, Zhang J, Wang YX
|
| RgGuinier |
5.7 |
nm |
| Dmax |
19.7 |
nm |
| VolumePorod |
230 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
P123 expressed protein dimer, 248 kDa Mycoplasma mobile (strain … protein
|
| Buffer: |
100 mM ammonium acetate, pH: 6.8 |
| Experiment: |
SAXS
data collected at BL-10C, Photon Factory (PF), High Energy Accelerator Research Organization (KEK) on 2019 Oct 25
|
Structure and Function of Gli123 Involved in Mycoplasma mobile Gliding.
J Bacteriol :e0034022 (2023)
Matsuike D, Tahara YO, Nonaka T, Wu HN, Hamaguchi T, Kudo H, Hayashi Y, Arai M, Miyata M
|
| RgGuinier |
8.2 |
nm |
| Dmax |
31.6 |
nm |
| VolumePorod |
436 |
nm3 |
|
|
|
|
|
|

|
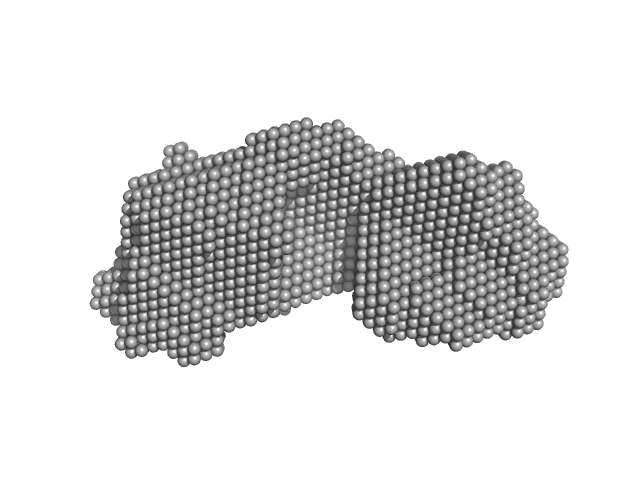
| Sample: |
SAVED domain-containing protein monomer, 58 kDa Sulfurihydrogenibium sp. (strain … protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Dec 14
|
Antiviral signalling by a cyclic nucleotide activated CRISPR protease.
Nature 614(7946):168-174 (2023)
Rouillon C, Schneberger N, Chi H, Blumenstock K, Da Vela S, Ackermann K, Moecking J, Peter MF, Boenigk W, Seifert R, Bode BE, Schmid-Burgk JL, Svergun D, Geyer M, White MF, Hagelueken G
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
80 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
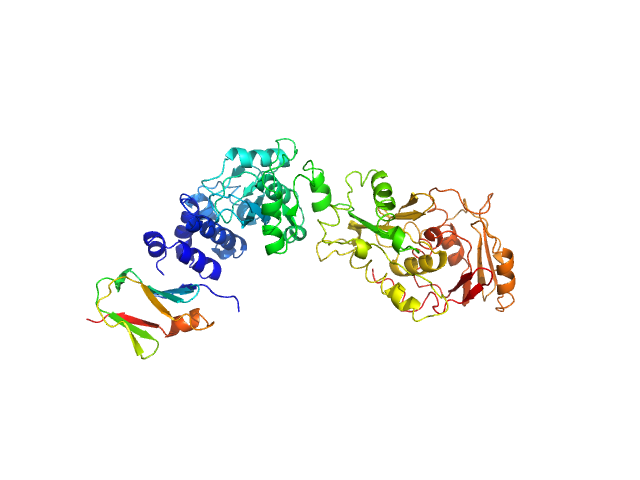
SAVED domain-containing protein monomer, 58 kDa Sulfurihydrogenibium sp. (strain … protein
Uncharacterized protein (putative MazF-like toxin) monomer, 9 kDa Sulfurihydrogenibium sp. (strain … protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2021 Dec 14
|
Antiviral signalling by a cyclic nucleotide activated CRISPR protease.
Nature 614(7946):168-174 (2023)
Rouillon C, Schneberger N, Chi H, Blumenstock K, Da Vela S, Ackermann K, Moecking J, Peter MF, Boenigk W, Seifert R, Bode BE, Schmid-Burgk JL, Svergun D, Geyer M, White MF, Hagelueken G
|
| RgGuinier |
3.4 |
nm |
| Dmax |
11.7 |
nm |
| VolumePorod |
93 |
nm3 |
|
|
|
|
|
|

|
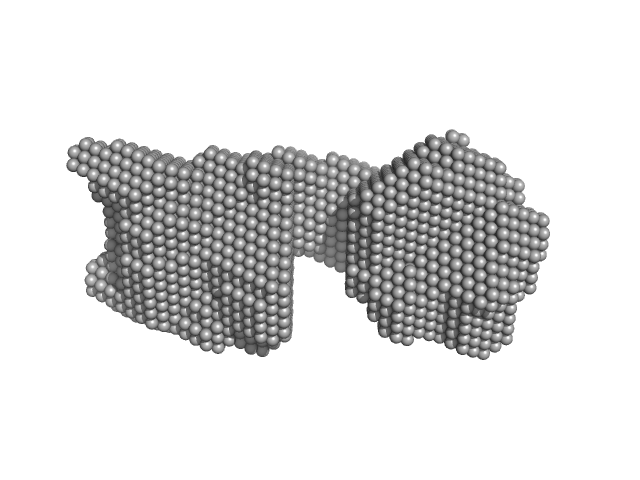
| Sample: |
SAVED domain-containing protein monomer, 58 kDa Sulfurihydrogenibium sp. (strain … protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Apr 8
|
Antiviral signalling by a cyclic nucleotide activated CRISPR protease.
Nature 614(7946):168-174 (2023)
Rouillon C, Schneberger N, Chi H, Blumenstock K, Da Vela S, Ackermann K, Moecking J, Peter MF, Boenigk W, Seifert R, Bode BE, Schmid-Burgk JL, Svergun D, Geyer M, White MF, Hagelueken G
|
| RgGuinier |
3.2 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
82 |
nm3 |
|
|
|
|
|
|

|
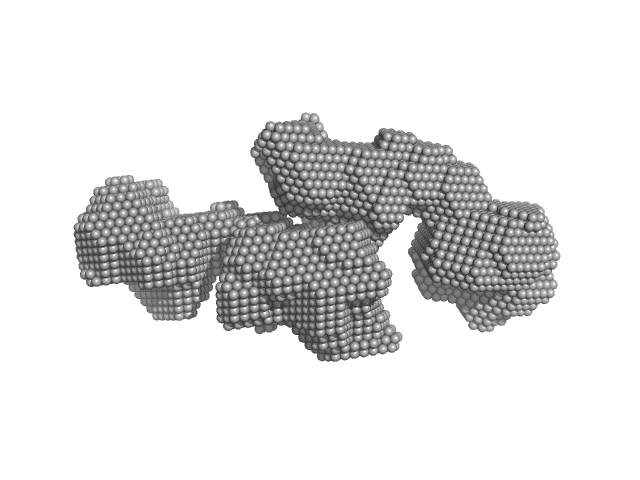
| Sample: |
SAVED domain-containing protein monomer, 58 kDa Sulfurihydrogenibium sp. (strain … protein
|
| Buffer: |
20 mM Tris, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Apr 8
|
Antiviral signalling by a cyclic nucleotide activated CRISPR protease.
Nature 614(7946):168-174 (2023)
Rouillon C, Schneberger N, Chi H, Blumenstock K, Da Vela S, Ackermann K, Moecking J, Peter MF, Boenigk W, Seifert R, Bode BE, Schmid-Burgk JL, Svergun D, Geyer M, White MF, Hagelueken G
|
| RgGuinier |
3.8 |
nm |
| Dmax |
13.0 |
nm |
| VolumePorod |
116 |
nm3 |
|
|
|
|
|
|

|
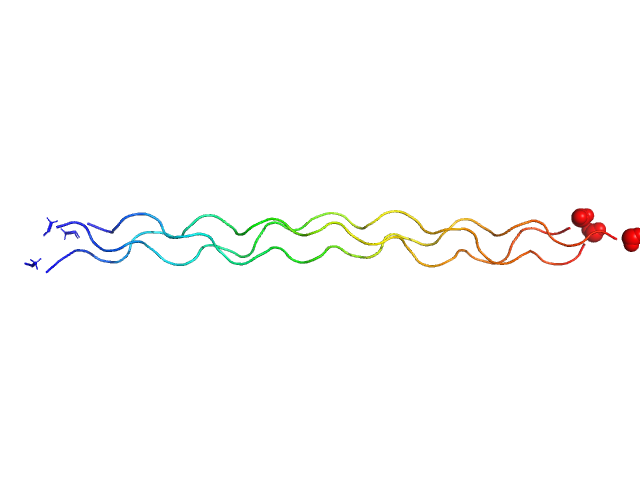
| Sample: |
Ac-(POG)4-POA-(POG)5-NH2 trimer, 12 kDa protein
|
| Buffer: |
20 mM L-histidine, 138 mM NaCl, 2.7 mM KCL,, pH: 6 |
| Experiment: |
SAXS
data collected at B21, Diamond Light Source on 2019 Feb 1
|
A solution structure analysis reveals a bent collagen triple helix in the complement activation recognition molecule mannan-binding lectin.
J Biol Chem 299(2):102799 (2023)
Iqbal H, Fung KW, Gor J, Bishop AC, Makhatadze GI, Brodsky B, Perkins SJ
|
| RgGuinier |
2.3 |
nm |
| Dmax |
11.0 |
nm |
| VolumePorod |
10 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
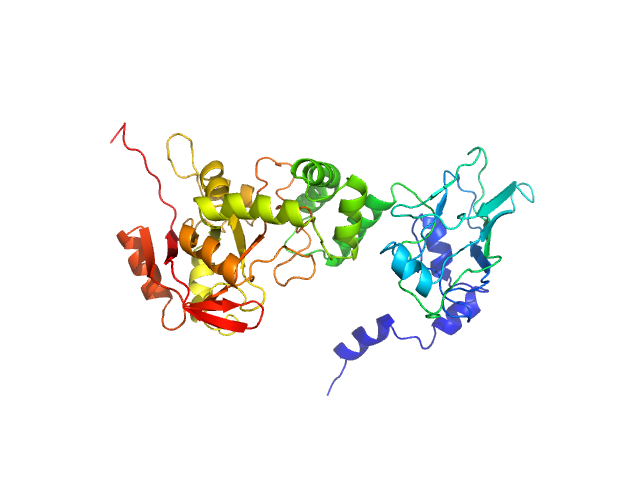
NAD glycohydrolase monomer, 47 kDa Streptococcus pyogenes M1 … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Oct 16
|
Structural basis underlying the synergism of NADase and SLO during group A Streptococcus infection.
Commun Biol 6(1):124 (2023)
Tsai WJ, Lai YH, Shi YA, Hammel M, Duff AP, Whitten AE, Wilde KL, Wu CM, Knott R, Jeng US, Kang CY, Hsu CY, Wu JL, Tsai PJ, Chiang-Ni C, Wu JJ, Lin YS, Liu CC, Senda T, Wang S
|
| RgGuinier |
3.0 |
nm |
| Dmax |
103.0 |
nm |
| VolumePorod |
66 |
nm3 |
|
|
|
|
|
|
|
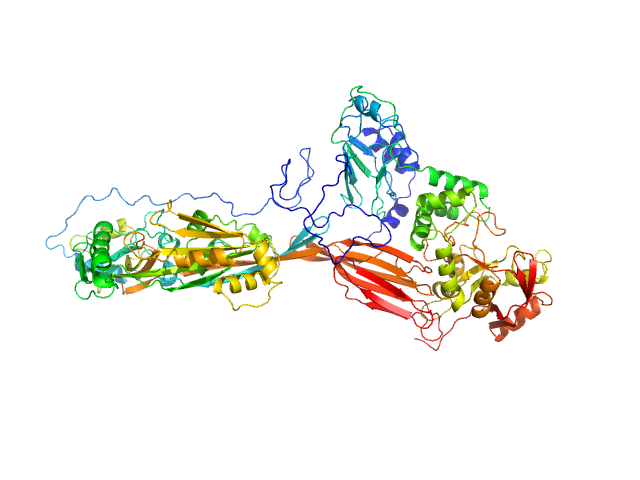
| Sample: |
NAD glycohydrolase monomer, 47 kDa Streptococcus pyogenes M1 … protein
Streptolysin O (T66M) monomer, 63 kDa Streptococcus pyogenes serotype … protein
|
| Buffer: |
phosphate buffered saline, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2018 Oct 16
|
Structural basis underlying the synergism of NADase and SLO during group A Streptococcus infection.
Commun Biol 6(1):124 (2023)
Tsai WJ, Lai YH, Shi YA, Hammel M, Duff AP, Whitten AE, Wilde KL, Wu CM, Knott R, Jeng US, Kang CY, Hsu CY, Wu JL, Tsai PJ, Chiang-Ni C, Wu JJ, Lin YS, Liu CC, Senda T, Wang S
|
| RgGuinier |
4.8 |
nm |
| Dmax |
18.4 |
nm |
| VolumePorod |
125 |
nm3 |
|
|