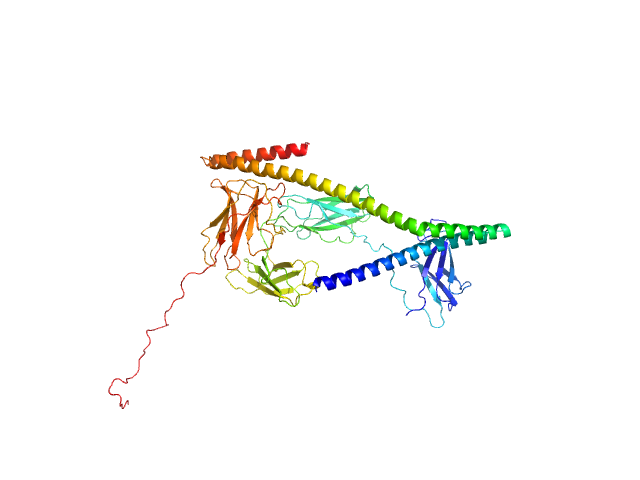
UniProt ID: P27951 (258-428) IgA FC receptor
UniProt ID: A0A0G2JM46 (24-443) Leukocyte immunoglobulin-like receptor subfamily B member 3 (P288R)
|
|
|
|
| Sample: |
IgA FC receptor monomer, 20 kDa Streptococcus agalactiae protein
Leukocyte immunoglobulin-like receptor subfamily B member 3 (P288R) monomer, 47 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Nov 17
|
Bacterial targeting of the neutrophil inhibitory receptor LILRB3 to evade antibody immunity
Ying Zhang
|
| RgGuinier |
3.9 |
nm |
| Dmax |
13.5 |
nm |
| VolumePorod |
202 |
nm3 |
|
|
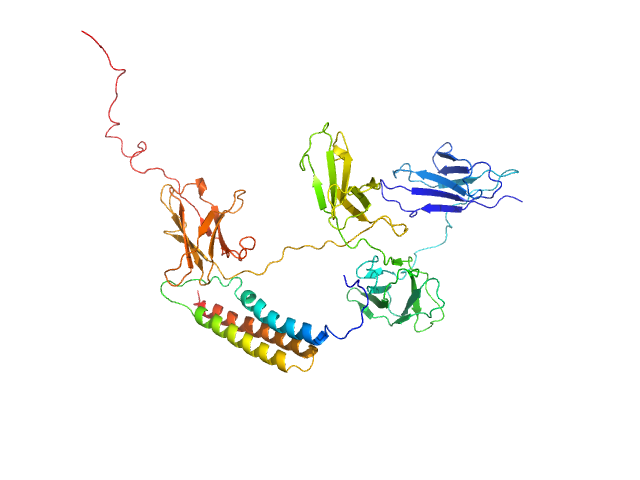
UniProt ID: A0A0G2JM46 (24-443) Leukocyte immunoglobulin-like receptor subfamily B member 3 (P288R)
UniProt ID: P27951 (725-826) IgA FC receptor (N770H)
|
|
|
|
| Sample: |
Leukocyte immunoglobulin-like receptor subfamily B member 3 (P288R) monomer, 47 kDa Homo sapiens protein
IgA FC receptor (N770H) monomer, 12 kDa Streptococcus agalactiae protein
|
| Buffer: |
20 mM HEPES, 300 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2023 Nov 17
|
Bacterial targeting of the neutrophil inhibitory receptor LILRB3 to evade antibody immunity
Ying Zhang
|
| RgGuinier |
3.9 |
nm |
| Dmax |
14.1 |
nm |
| VolumePorod |
175 |
nm3 |
|
|
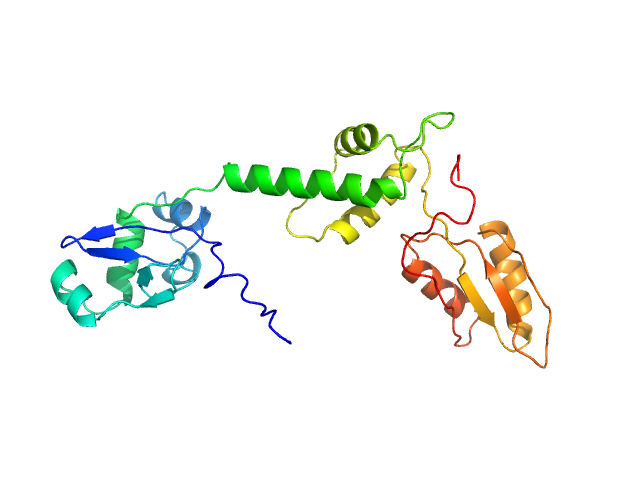
UniProt ID: Q9Y3A5 (1-250) Ribosome maturation protein SBDS
|
|
|
|
| Sample: |
Ribosome maturation protein SBDS monomer, 29 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Apr 6
|
Structural Implications of Missense Point Mutations in Shwachman–Bodian–Diamond Syndrome Protein (SBDS): A Combined SAXS/MD Investigation
ACS Omega (2025)
Mattiotti G, Nanna V, Giulini M, Alberga D, Mangiatordi G, Sánchez-Puig N, Saviano M, Tubiana L, Potestio R, Lattanzi G, Siliqi D
|
| RgGuinier |
3.0 |
nm |
| Dmax |
11.4 |
nm |
| VolumePorod |
50 |
nm3 |
|
|
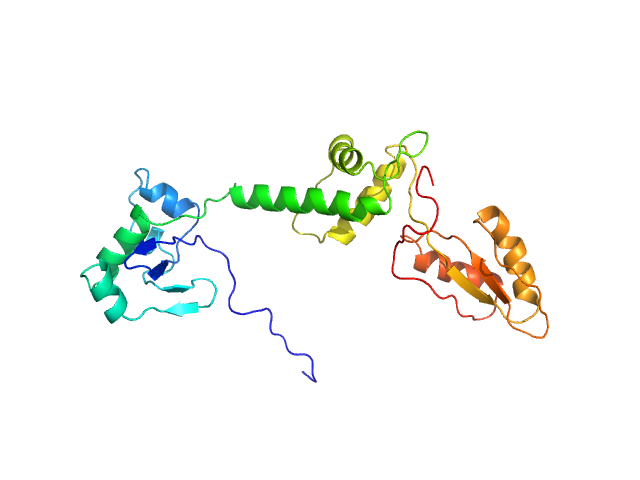
UniProt ID: Q9Y3A5 (1-250) Ribosome maturation protein SBDS (R19Q)
|
|
|
|
| Sample: |
Ribosome maturation protein SBDS (R19Q) monomer, 29 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Apr 6
|
Structural Implications of Missense Point Mutations in Shwachman–Bodian–Diamond Syndrome Protein (SBDS): A Combined SAXS/MD Investigation
ACS Omega (2025)
Mattiotti G, Nanna V, Giulini M, Alberga D, Mangiatordi G, Sánchez-Puig N, Saviano M, Tubiana L, Potestio R, Lattanzi G, Siliqi D
|
| RgGuinier |
2.9 |
nm |
| Dmax |
12.2 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
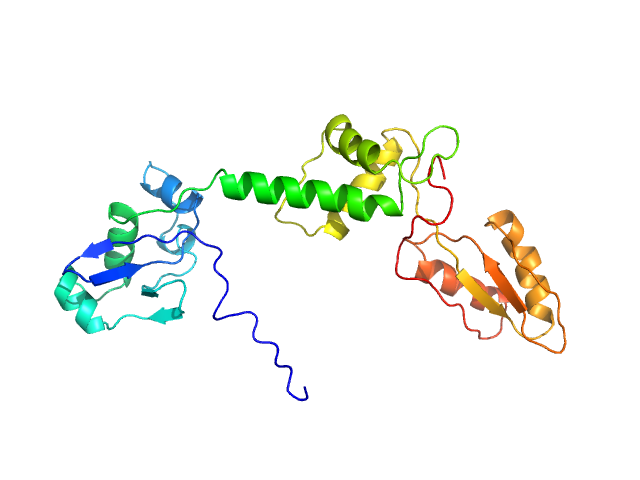
UniProt ID: Q9Y3A5 (1-250) Ribosome maturation protein SBDS (I167T)
|
|
|
|
| Sample: |
Ribosome maturation protein SBDS (I167T) monomer, 29 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Apr 6
|
Structural Implications of Missense Point Mutations in Shwachman–Bodian–Diamond Syndrome Protein (SBDS): A Combined SAXS/MD Investigation
ACS Omega (2025)
Mattiotti G, Nanna V, Giulini M, Alberga D, Mangiatordi G, Sánchez-Puig N, Saviano M, Tubiana L, Potestio R, Lattanzi G, Siliqi D
|
| RgGuinier |
2.9 |
nm |
| Dmax |
10.5 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
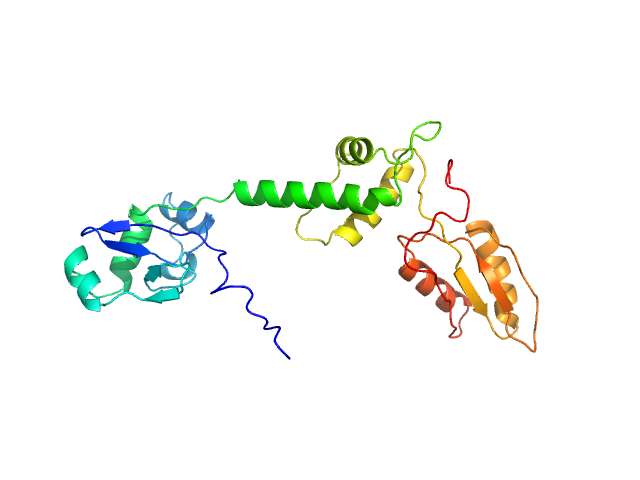
UniProt ID: Q9Y3A5 (1-250) Ribosome maturation protein SBDS (R175W)
|
|
|
|
| Sample: |
Ribosome maturation protein SBDS (R175W) monomer, 29 kDa Homo sapiens protein
|
| Buffer: |
50 mM Tris pH 8.0, 10% glycerol, 300 mM NaCl, 5 mM MgCl2., pH: |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2022 Apr 6
|
Structural Implications of Missense Point Mutations in Shwachman–Bodian–Diamond Syndrome Protein (SBDS): A Combined SAXS/MD Investigation
ACS Omega (2025)
Mattiotti G, Nanna V, Giulini M, Alberga D, Mangiatordi G, Sánchez-Puig N, Saviano M, Tubiana L, Potestio R, Lattanzi G, Siliqi D
|
| RgGuinier |
3.1 |
nm |
| Dmax |
11.6 |
nm |
| VolumePorod |
48 |
nm3 |
|
|
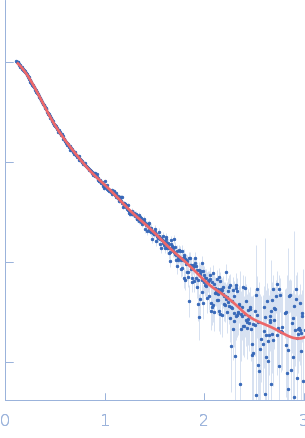
UniProt ID: Q9Y371 (1-365) Endophilin-B1
|
|
|
|
| Sample: |
Endophilin-B1 dimer, 82 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.5 mM DTT, pH: 8.1 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2024 Mar 10
|
Peripheral membrane protein endophilin B1 probes, perturbs and permeabilizes lipid bilayers
Communications Biology 8(1) (2025)
Thorlacius A, Rulev M, Sundberg O, Sundborger-Lunna A
|
| RgGuinier |
5.1 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
151 |
nm3 |
|
|
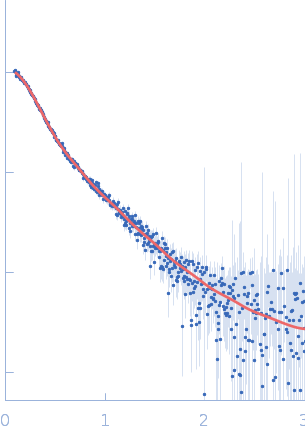
UniProt ID: Q9Y371 (1-365) Endophilin-B1
|
|
|
|
| Sample: |
Endophilin-B1 dimer, 82 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.5 mM DTT, pH: 8.1 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2024 Mar 10
|
Peripheral membrane protein endophilin B1 probes, perturbs and permeabilizes lipid bilayers
Communications Biology 8(1) (2025)
Thorlacius A, Rulev M, Sundberg O, Sundborger-Lunna A
|
| RgGuinier |
5.0 |
nm |
| Dmax |
16.0 |
nm |
| VolumePorod |
152 |
nm3 |
|
|
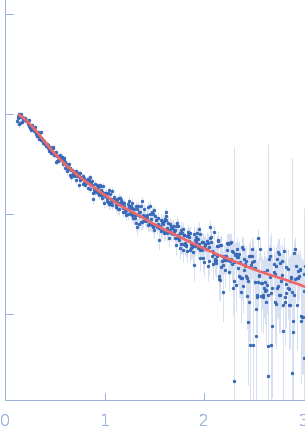
UniProt ID: Q9Y371 (1-365) Endophilin-B1
|
|
|
|
| Sample: |
Endophilin-B1 monomer, 41 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.5 mM DTT, pH: 8.1 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2024 Mar 10
|
Peripheral membrane protein endophilin B1 probes, perturbs and permeabilizes lipid bilayers
Communications Biology 8(1) (2025)
Thorlacius A, Rulev M, Sundberg O, Sundborger-Lunna A
|
| RgGuinier |
3.7 |
nm |
| Dmax |
15.3 |
nm |
| VolumePorod |
47 |
nm3 |
|
|
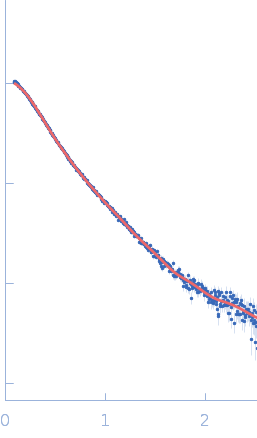
UniProt ID: Q9Y371 (1-306) Endophilin-B1 (Δ307-360)
|
|
|
|
| Sample: |
Endophilin-B1 (Δ307-360) dimer, 68 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1 mM TCEP, 0.5 mM DTT, pH: 8.1 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2024 Mar 10
|
Peripheral membrane protein endophilin B1 probes, perturbs and permeabilizes lipid bilayers
Communications Biology 8(1) (2025)
Thorlacius A, Rulev M, Sundberg O, Sundborger-Lunna A
|
| RgGuinier |
4.6 |
nm |
| Dmax |
15.2 |
nm |
| VolumePorod |
149 |
nm3 |
|
|