|
|
|
|

|
| Sample: |
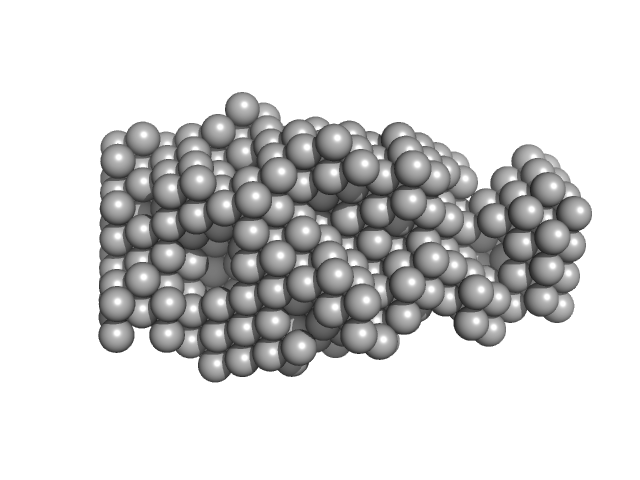
Thyroglobulin dimer, 610 kDa Homo sapiens protein
|
| Buffer: |
phosphate buffered saline, 150mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2020 Jul 17
|
Thyroglobulin - human and bovine SAXS/WAXS data
James Byrnes
|
| RgGuinier |
7.7 |
nm |
| Dmax |
26.3 |
nm |
| VolumePorod |
1800 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
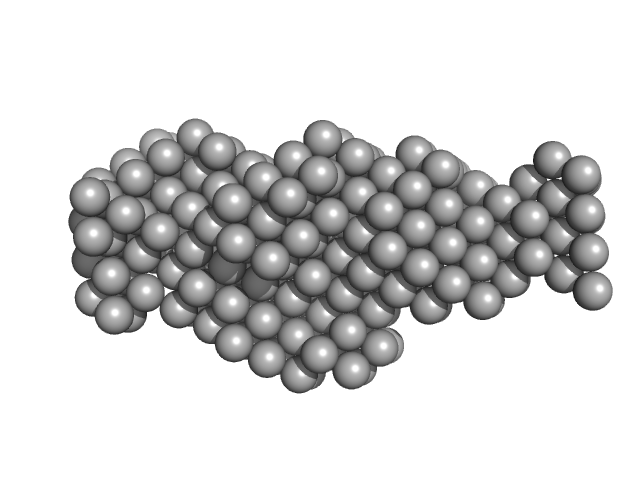
Thyroglobulin dimer, 606 kDa Bos taurus protein
|
| Buffer: |
phosphate buffered saline, 150mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2020 Jul 17
|
Thyroglobulin - human and bovine SAXS/WAXS data
James Byrnes
|
| RgGuinier |
8.1 |
nm |
| Dmax |
29.8 |
nm |
| VolumePorod |
1950 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
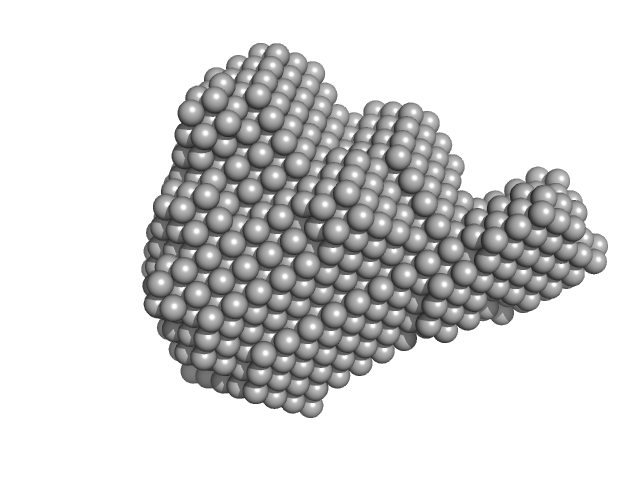
Aprataxin and PNK-like factor (acidic domain) dimer, 15 kDa Homo sapiens protein
Histone H2A dimer, 26 kDa Drosophila melanogaster protein
Histone H2B dimer, 27 kDa Drosophila melanogaster protein
Histone H3 dimer, 30 kDa Drosophila melanogaster protein
Histone H4 dimer, 23 kDa Drosophila melanogaster protein
|
| Buffer: |
25 mM NaPi, 300 mM NaCl, 3% v/v glycerol, 1 mM DTT,, pH: 7 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2018 Jul 10
|
Chaperoning of the histone octamer by the acidic domain of DNA repair factor APLF.
Sci Adv 8(30):eabo0517 (2022)
Corbeski I, Guo X, Eckhardt BV, Fasci D, Wiegant W, Graewert MA, Vreeken K, Wienk H, Svergun DI, Heck AJR, van Attikum H, Boelens R, Sixma TK, Mattiroli F, van Ingen H
|
| RgGuinier |
3.7 |
nm |
| Dmax |
10.8 |
nm |
| VolumePorod |
260 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
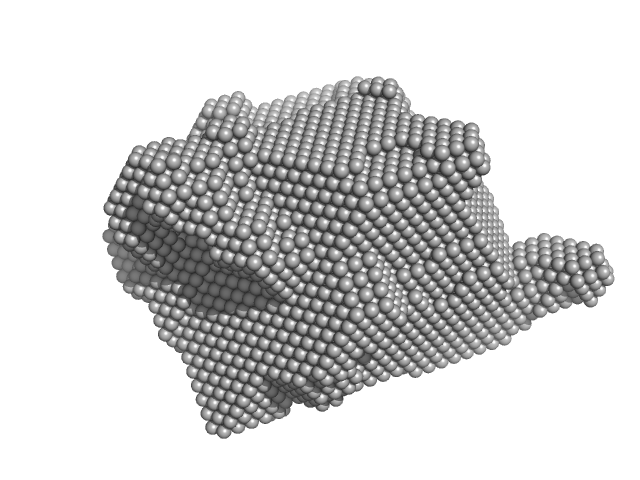
RuvB-like 1 hexamer, 311 kDa Homo sapiens protein
RuvB-like 2 hexamer, 311 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1% glycerol, 5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 21
|
Deciphering cellular and molecular determinants of human DPCD protein in complex with RUVBL1/RUVBL2 AAA-ATPases.
J Mol Biol :167760 (2022)
Dos Santos Morais R, Santo PE, Ley M, Schelcher C, Abel Y, Plassart L, Deslignière E, Chagot ME, Quinternet M, Paiva ACF, Hessmann S, Morellet N, M F Sousa P, Vandermoere F, Bertrand E, Charpentier B, Bandeiras TM, Plisson-Chastang C, Verheggen C, Cianférani S, Manival X
|
| RgGuinier |
6.2 |
nm |
| Dmax |
20.0 |
nm |
| VolumePorod |
1490 |
nm3 |
|
|
|
|
|
|

|
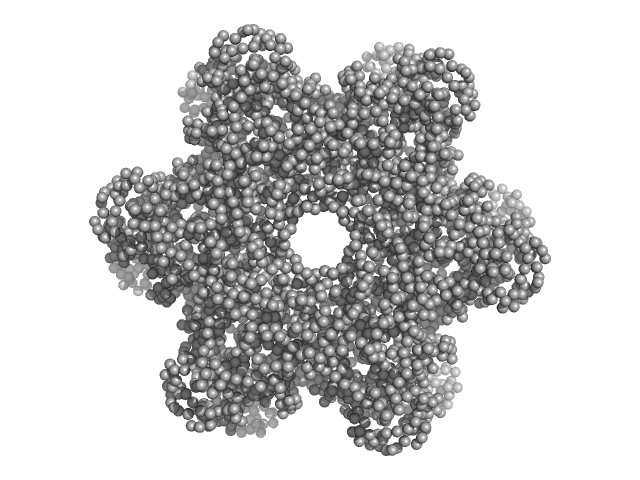
| Sample: |
Protein DPCD trimer, 71 kDa Homo sapiens protein
RuvB-like 1 trimer, 155 kDa Homo sapiens protein
RuvB-like 2 trimer, 156 kDa Homo sapiens protein
|
| Buffer: |
20 mM HEPES, 150 mM NaCl, 1% glycerol, 5 mM TCEP, pH: 7.5 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2019 Mar 21
|
Deciphering cellular and molecular determinants of human DPCD protein in complex with RUVBL1/RUVBL2 AAA-ATPases.
J Mol Biol :167760 (2022)
Dos Santos Morais R, Santo PE, Ley M, Schelcher C, Abel Y, Plassart L, Deslignière E, Chagot ME, Quinternet M, Paiva ACF, Hessmann S, Morellet N, M F Sousa P, Vandermoere F, Bertrand E, Charpentier B, Bandeiras TM, Plisson-Chastang C, Verheggen C, Cianférani S, Manival X
|
| RgGuinier |
5.4 |
nm |
| Dmax |
15.9 |
nm |
| VolumePorod |
856 |
nm3 |
|
|
|
|
|
|

|
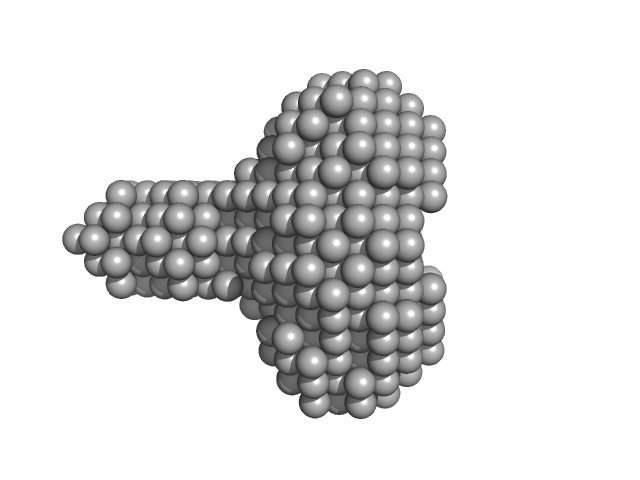
| Sample: |
Lysin [Streptococcus phage P7951] hexamer, 65 kDa Streptococcus phage P7951 protein
|
| Buffer: |
50 mM HEPES, 500 mM NaCl, and 1% glycerol, pH: 7 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2021 Nov 21
|
On the Occurrence and Multimerization of Two-Polypeptide Phage Endolysins Encoded in Single Genes.
Microbiol Spectr :e0103722 (2022)
Pinto D, Gonçalo R, Louro M, Silva MS, Hernandez G, Cordeiro TN, Cordeiro C, São-José C
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.1 |
nm |
| VolumePorod |
111 |
nm3 |
|
|
|
|
|
|

|
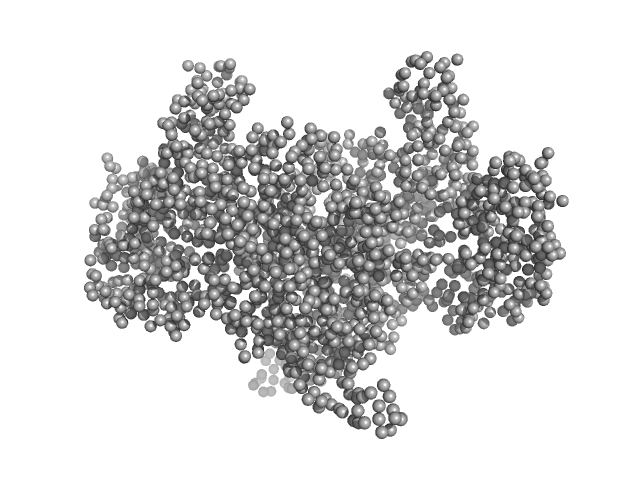
| Sample: |
Aconitate hydratase B, 96 kDa Acinetobacter baumannii protein
|
| Buffer: |
100 mM NaCl, 20 mM Tris, 1 mM DTT, 3% v/v glycerol, pH: 8 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2020 Dec 3
|
Recombinant AcnB, NrdR and RibD of Acinetobacter baumannii and their potential interaction with DNA adenine methyltransferase AamA.
Protein Expr Purif :106134 (2022)
Weber K, Doellinger J, Jeffries CM, Wilharm G
|
| RgGuinier |
4.0 |
nm |
| Dmax |
14.0 |
nm |
| VolumePorod |
230 |
nm3 |
|
|
|
|
|
|

|
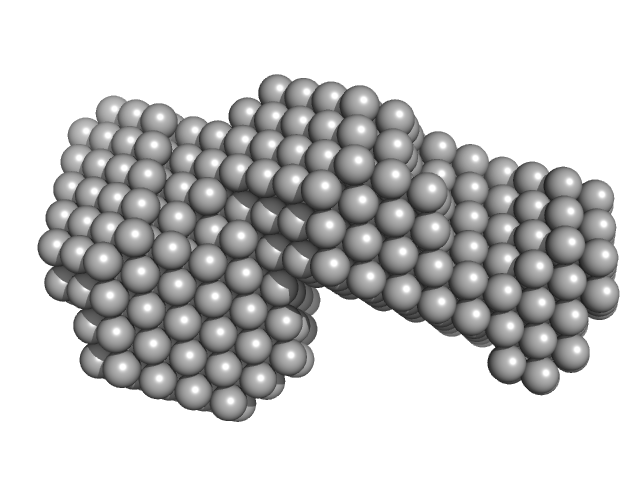
| Sample: |
ESX-5 secretion system protein EccA5 monomer, 31 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
20 mM Tris, 150 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2018 Mar 9
|
Structural and Biophysical characterization of N-terminal domain of EccA5 from Mycobacterium tuberculosis.
Jyoti Vishwakarma
|
| RgGuinier |
2.6 |
nm |
| Dmax |
7.8 |
nm |
| VolumePorod |
57 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Polyketide synthase Pks13 monomer, 188 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2013 Mar 1
|
Solution structure of the type I polyketide synthase Pks13 from Mycobacterium tuberculosis.
BMC Biol 20(1):147 (2022)
Bon C, Cabantous S, Julien S, Guillet V, Chalut C, Rima J, Brison Y, Malaga W, Sanchez-Dafun A, Gavalda S, Quémard A, Marcoux J, Waldo GS, Guilhot C, Mourey L
|
| RgGuinier |
7.3 |
nm |
| Dmax |
24.8 |
nm |
| VolumePorod |
480 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
Polyketide synthase Pks13 dimer, 376 kDa Mycobacterium tuberculosis (strain … protein
|
| Buffer: |
50 mM Tris-HCl, 50 mM NaCl, pH: 8 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2011 Nov 23
|
Solution structure of the type I polyketide synthase Pks13 from Mycobacterium tuberculosis.
BMC Biol 20(1):147 (2022)
Bon C, Cabantous S, Julien S, Guillet V, Chalut C, Rima J, Brison Y, Malaga W, Sanchez-Dafun A, Gavalda S, Quémard A, Marcoux J, Waldo GS, Guilhot C, Mourey L
|
| RgGuinier |
10.3 |
nm |
| Dmax |
36.0 |
nm |
| VolumePorod |
2300 |
nm3 |
|
|