|
|
|
|
|
| Sample: |
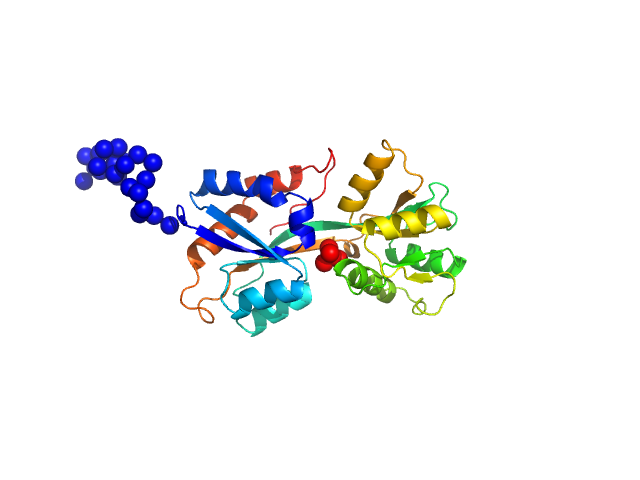
ABC transporter periplasmic substrate-binding protein monomer, 30 kDa Desulfovibrio alaskensis protein
|
| Buffer: |
5 mM Tris-HCl, pH: 7.6 |
| Experiment: |
SAXS
data collected at BM29, ESRF on 2014 Jun 21
|
Highly selective tungstate transporter protein TupA from Desulfovibrio alaskensis G20.
Sci Rep 7(1):5798 (2017)
Otrelo-Cardoso AR, Nair RR, Correia MAS, Cordeiro RSC, Panjkovich A, Svergun DI, Santos-Silva T, Rivas MG
|
| RgGuinier |
2.3 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
44 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
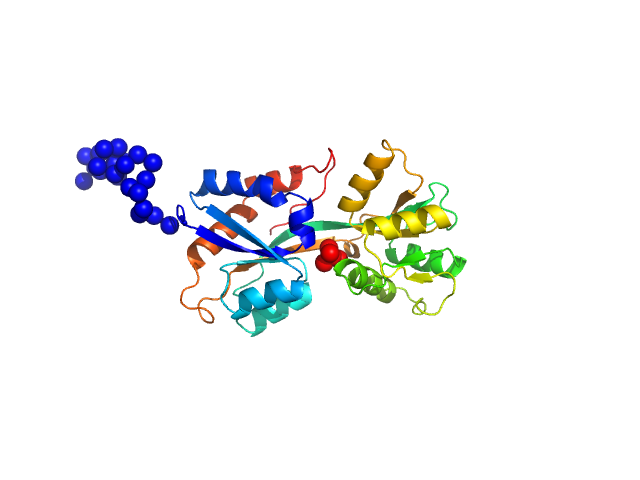
ABC transporter periplasmic substrate-binding protein monomer, 30 kDa Desulfovibrio alaskensis protein
|
| Buffer: |
5 mM Tris-HCl, pH: 7.6 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Dec 7
|
Highly selective tungstate transporter protein TupA from Desulfovibrio alaskensis G20.
Sci Rep 7(1):5798 (2017)
Otrelo-Cardoso AR, Nair RR, Correia MAS, Cordeiro RSC, Panjkovich A, Svergun DI, Santos-Silva T, Rivas MG
|
| RgGuinier |
2.3 |
nm |
| Dmax |
9.2 |
nm |
| VolumePorod |
46 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
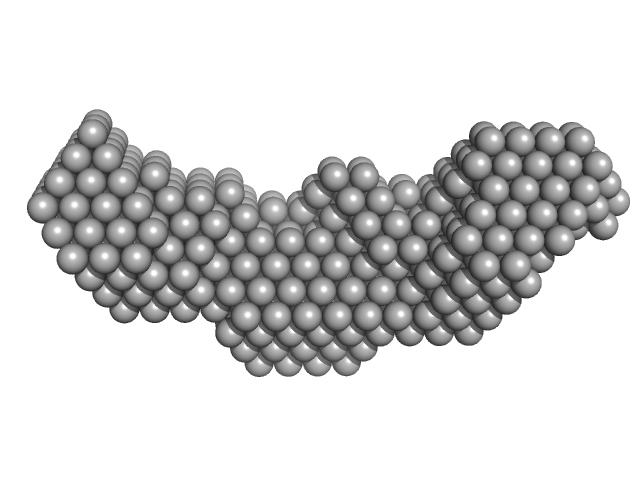
Homeobox protein CEH-14 monomer, 16 kDa Caenorhabditis elegans protein
CeLIM-7 monomer, 4 kDa Caenorhabditis elegans protein
|
| Buffer: |
50 mM Tris, 100 mM NaCl, 5 mM TCEP, pH: 8 |
| Experiment: |
SAXS
data collected at Anton Paar SAXSess, University of Sydney on 2009 Apr 7
|
Interactions between LHX3- and ISL1-family LIM-homeodomain transcription factors are conserved in Caenorhabditis elegans.
Sci Rep 7(1):4579 (2017)
Bhati M, Llamosas E, Jacques DA, Jeffries CM, Dastmalchi S, Ripin N, Nicholas HR, Matthews JM
|
| RgGuinier |
2.4 |
nm |
| Dmax |
8.9 |
nm |
| VolumePorod |
26 |
nm3 |
|
|
|
|
|
|
|
| Sample: |
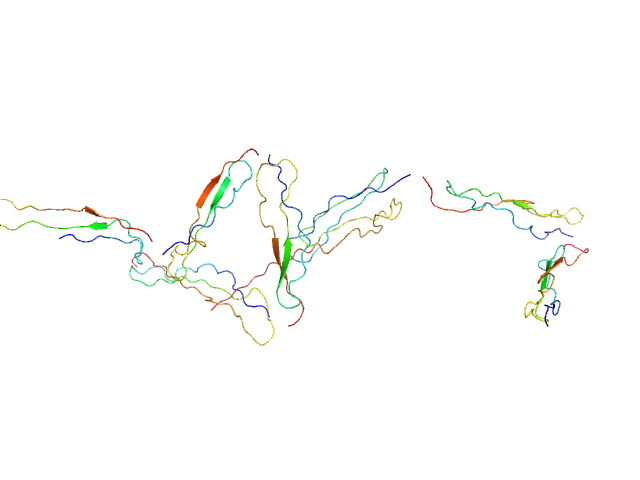
Human recombinant Properdin TSR 0-3 monomer, 25 kDa Homo sapiens protein
Human recombinant Properdin TSR 4-6 monomer, 24 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES 150 mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2015 Aug 16
|
Functional and structural insight into properdin control of complement alternative pathway amplification.
EMBO J 36(8):1084-1099 (2017)
Pedersen DV, Roumenina L, Jensen RK, Gadeberg TA, Marinozzi C, Picard C, Rybkine T, Thiel S, Sørensen UB, Stover C, Fremeaux-Bacchi V, Andersen GR
|
| RgGuinier |
5.1 |
nm |
| Dmax |
18.0 |
nm |
|
|
|
|
|
|
|
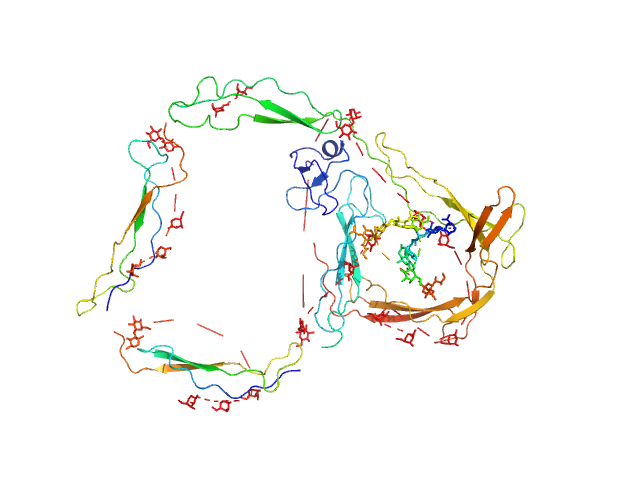
| Sample: |
E244K Human Properdin monomer, 49 kDa Homo sapiens protein
|
| Buffer: |
10 mM HEPES 150 mM NaCl, pH: 7.2 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2016 Jul 27
|
Functional and structural insight into properdin control of complement alternative pathway amplification.
EMBO J 36(8):1084-1099 (2017)
Pedersen DV, Roumenina L, Jensen RK, Gadeberg TA, Marinozzi C, Picard C, Rybkine T, Thiel S, Sørensen UB, Stover C, Fremeaux-Bacchi V, Andersen GR
|
| RgGuinier |
4.1 |
nm |
| Dmax |
15.0 |
nm |
|
|
|
|
|
|
|
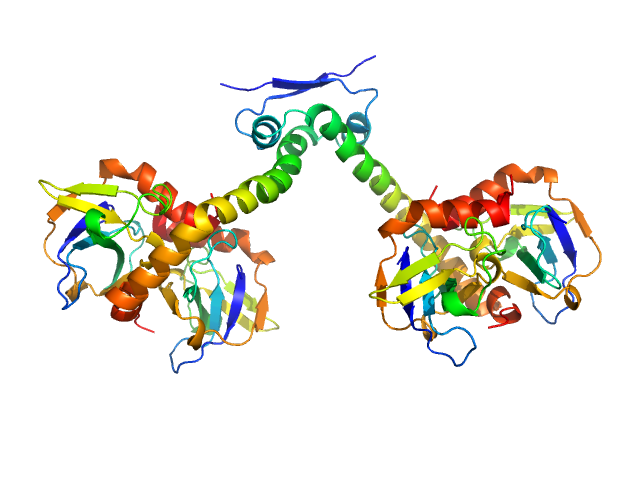
| Sample: |
Toxin CcdB dimer, 23 kDa Escherichia coli protein
Antitoxin CcdA dimer, 17 kDa Escherichia coli protein
Toxin CcdB dimer, 23 kDa Escherichia coli protein
|
| Buffer: |
10 mM Tris 50 mM NaCl, pH: 7.3 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2013 Jul 24
|
Molecular mechanism governing ratio-dependent transcription regulation in the ccdAB operon.
Nucleic Acids Res 45(6):2937-2950 (2017)
Vandervelde A, Drobnak I, Hadži S, Sterckx YG, Welte T, De Greve H, Charlier D, Efremov R, Loris R, Lah J
|
| RgGuinier |
3.4 |
nm |
| Dmax |
12.0 |
nm |
| VolumePorod |
92 |
nm3 |
|
|
|
|
|
|

|
| Sample: |
Type-2 restriction enzyme AgeI monomer, 31 kDa Thalassobius gelatinovorus protein
|
| Buffer: |
10 mM Tris-HCl, pH 7.5, 150 mM NaCl, 5 mM CaCl₂, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Nov 14
|
Restriction endonuclease AgeI is a monomer which dimerizes to cleave DNA.
Nucleic Acids Res 45(6):3547-3558 (2017)
Tamulaitiene G, Jovaisaite V, Tamulaitis G, Songailiene I, Manakova E, Zaremba M, Grazulis S, Xu SY, Siksnys V
|
| RgGuinier |
2.2 |
nm |
| Dmax |
6.7 |
nm |
| VolumePorod |
53 |
nm3 |
|
|
|
|
|
|
|
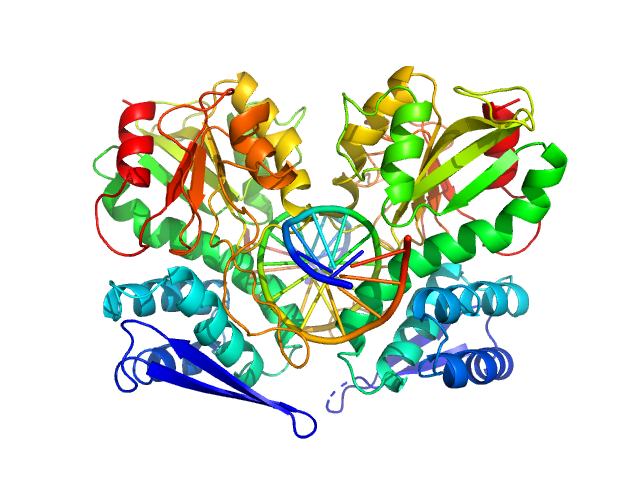
| Sample: |
Type-2 restriction enzyme AgeI dimer, 61 kDa Thalassobius gelatinovorus protein
Cognate DNA oligoduplex with 5'-T overhang dimer, 8 kDa DNA
|
| Buffer: |
10 mM Tris-HCl, pH 7.5, 150 mM NaCl, 5 mM CaCl₂, pH: 7.5 |
| Experiment: |
SAXS
data collected at EMBL P12, PETRA III on 2013 Nov 14
|
Restriction endonuclease AgeI is a monomer which dimerizes to cleave DNA.
Nucleic Acids Res 45(6):3547-3558 (2017)
Tamulaitiene G, Jovaisaite V, Tamulaitis G, Songailiene I, Manakova E, Zaremba M, Grazulis S, Xu SY, Siksnys V
|
| RgGuinier |
2.4 |
nm |
| Dmax |
7.4 |
nm |
| VolumePorod |
69 |
nm3 |
|
|
|
|
|
|
|
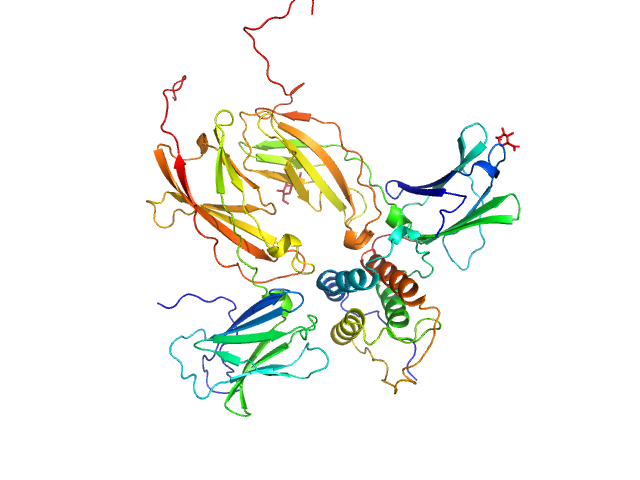
| Sample: |
Thymic stromal lymphopoietin monomer, 15 kDa Homo sapiens protein
Interleukin-7 receptor subunit alpha monomer, 26 kDa Homo sapiens protein
Cytokine receptor-like factor 2 monomer, 24 kDa Homo sapiens protein
|
| Buffer: |
10 mM Hepes, 150 mM NaCl, pH: 7.4 |
| Experiment: |
SAXS
data collected at SWING, SOLEIL on 2016 Apr 8
|
Structure and antagonism of the receptor complex mediated by human TSLP in allergy and asthma.
Nat Commun 8:14937 (2017)
Verstraete K, Peelman F, Braun H, Lopez J, Van Rompaey D, Dansercoer A, Vandenberghe I, Pauwels K, Tavernier J, Lambrecht BN, Hammad H, De Winter H, Beyaert R, Lippens G, Savvides SN
|
| RgGuinier |
3.1 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
98 |
nm3 |
|
|
|
|
|
|
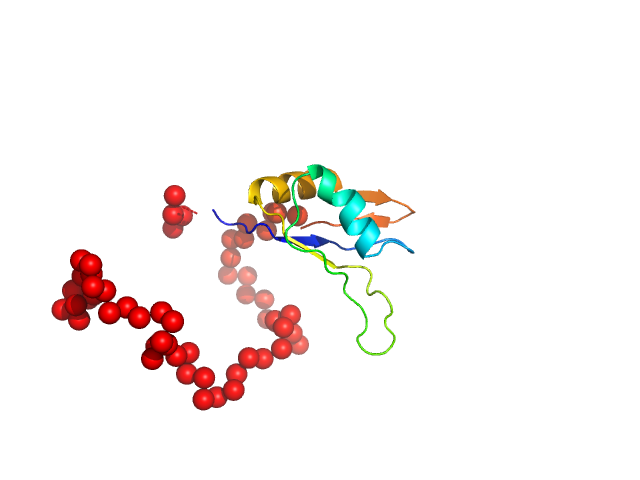
|
| Sample: |
Ribosome biogenesis protein 15 monomer, 17 kDa Saccharomyces cerevisiae protein
|
| Buffer: |
25 mM HEPES, 500 mM NaCl, 2 mM DTT, pH: 7.5 |
| Experiment: |
SAXS
data collected at 12.3.1 (SIBYLS), Advanced Light Source (ALS) on 2013 Apr 25
|
Structural analysis reveals the flexible C-terminus of Nop15 undergoes rearrangement to recognize a pre-ribosomal RNA folding intermediate.
Nucleic Acids Res 45(5):2829-2837 (2017)
Zhang J, Gonzalez LE, Hall TMT
|
| RgGuinier |
2.4 |
nm |
| Dmax |
10.3 |
nm |
| VolumePorod |
38 |
nm3 |
|
|