|
|
|
|
|
| Sample: |
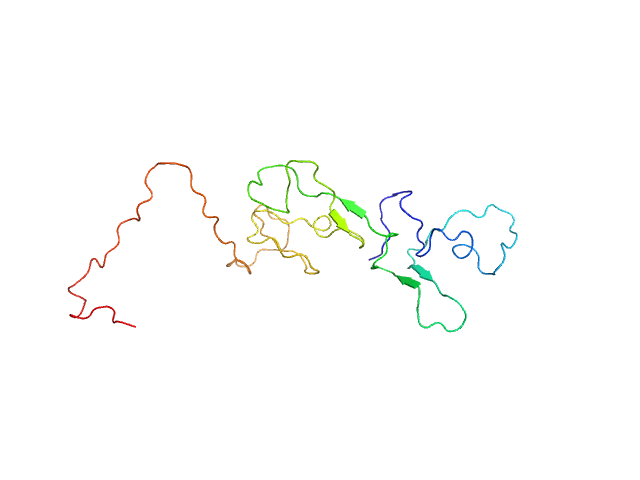
Estrogen receptor monomer, 20 kDa Homo sapiens protein
|
| Buffer: |
20 mM sodium phosphate, 50 mM NaCl, 0.05 mM TCEP, pH: 7.4
|
| Experiment: |
SAXS
data collected at 16-ID (LiX), National Synchrotron Light Source II (NSLS-II) on 2017 Jun 12
|
A Metastable Contact and Structural Disorder in the Estrogen Receptor Transactivation Domain.
Structure 27(2):229-240.e4 (2019)
...Xiao X, Du Z, Hsieh A, Ko S, Chen Y, Agrawal P, Zheng W, Shi W, Jiang W, Yang L, Chance MR, Surewicz WK, Buck M, Yang S
|
| RgGuinier |
3.0 |
nm |
| Dmax |
10.0 |
nm |
|